Tuberculosis-associated IFN-I induces Siglec-1 on tunneling nanotubes and favors HIV-1 spread in macrophages
Figures
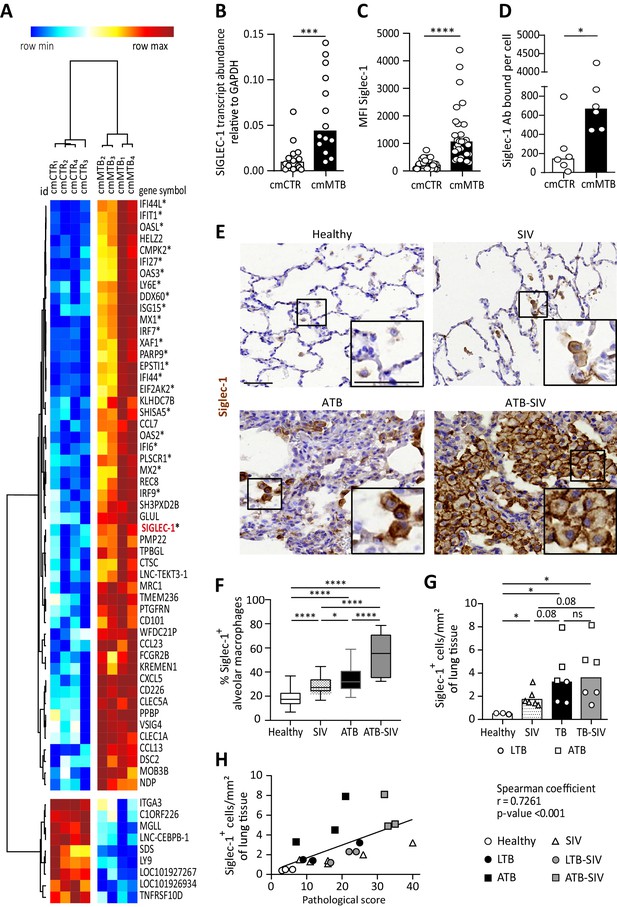
Tuberculosis-associated microenvironments induce Siglec-1 expression in macrophages.
(A–D) For 3 days, human monocytes were differentiated into macrophages with cmCTR (white) or cmMTB (black) supernatants. (A) Heatmap from a transcriptomic analysis (GEO submission GSE139511) illustrating the top 60 differentially expressed genes (DEGs) between cmCTR- or cmMTB-cells. Selection of the top DEGs was performed using an adjusted p-value ≤ 0.05, a fold change of at least 2, and a minimal expression of 6 in a log2 scale. Hierarchical clustering was performed using the complete linkage method and the Pearson correlation metric with Morpheus (Broad Institute). Interferon-stimulated genes (ISG) are labelled with an asterisk and Siglec-1 is indicated in red. (B–D) Validation of Siglec-1 expression in cmMTB-treated macrophages. Vertical scatter plots showing the relative abundance to mRNA (B), median fluorescent intensity (MFI) (C), and mean number of Siglec-1 antibody binding sites per cell (D). Each circle represents a single donor and histograms median values. (E) Representative immunohistochemical images of Siglec-1 staining (brown) in lung biopsies of healthy, SIV infected (SIV), active TB (ATB), and co-infected (ATB-SIV) non-human primates (NHP). Scale bar, 100 µm. Insets are 2x zoom. (F) Vertical Box and Whisker plot indicating the distribution of the percentage of Siglec-1+ alveolar macrophages in lung biopsies from the indicated NHP groups. Quantification analysis from n = 800 alveolar macrophages grouped from three independent animals per NHP group. (G) Vertical scatter plots displaying the number of cells that are positive for Siglec-1 per mm² of lung biopsies from the indicated NHP groups. Each symbol represents a single animal per NHP group. (H) Correlation between Siglec-1+ cells per mm² of lung tissue and the pathological score for healthy (white circle), SIV+ (white triangles), latent (black circle) or active (black square) TB, and SIV+ with latent (grey circle) or active (grey square) TB. Each symbol represents a single animal per NHP group. Mean value is represented as a black line. Statistical analyses: Two-tailed, Wilcoxon signed-rank test (B–D), Mann-Whitney unpaired test (F–G), Spearman correlation (H). *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001. ns: not significant. See Figure 1—source data 1.
-
Figure 1—source data 1
Raw data and statistical analyses supporting Siglec-1 expression in human and non-human primate macrophages exposed to TB-associated microenvironment.
- https://cdn.elifesciences.org/articles/52535/elife-52535-fig1-data1-v2.xlsx
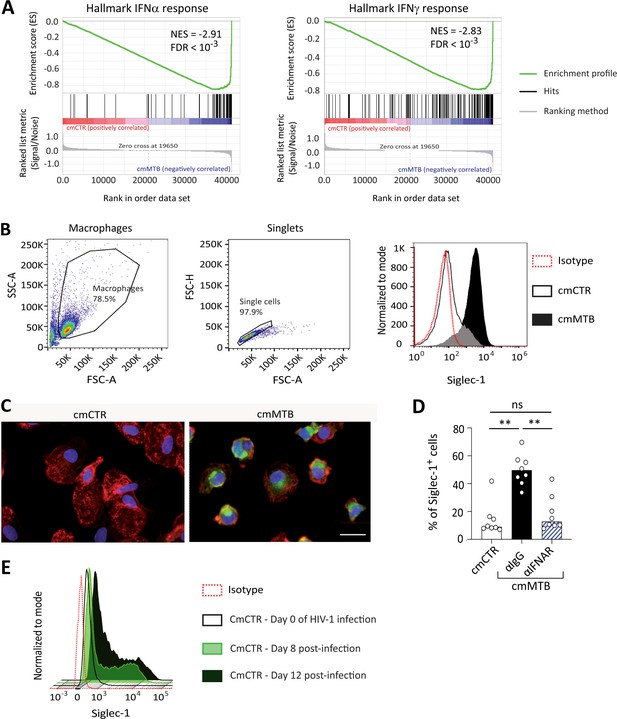
Tuberculosis-associated microenvironments increase Siglec-1 expression in human macrophages.
(A–D) For 3 days, human monocytes were differentiated into macrophages with cmCTR (white) or cmMTB (black) supernatants. (A) (Left) Gene set enrichment plot of the interferon alpha (IFNα) response (hallmark collection of MSigDB). This plot shows the distribution of the barcode between macrophages exposed to cmCTR (red) versus cmMTB (blue) supernatants. Each bar of the barcode corresponds to a signature gene of the gene set. The skewing to the right indicates enrichment in macrophages exposed to cmMTB versus cmCTR supernatant of genes up-regulated in response to IFNα. (Right) Gene set enrichment plot of the IFNγ response (hallmark collection of MSigDB). (B) Flow cytometry gating strategy to assess Siglec-1 cell-surface expression in human macrophages exposed to cmCTR (white) and cmMTB (black). (Left) Based on size (FSC-A) and granularity (SSC-A), a gate was created to separate human macrophages from cell debris and dying cells. Macrophages were then subjected through a second gate based FSC Area and Height Scaling (FSC-A and FSC-H) to separate singlets from doublets. (Right) Based on the singlet gate, the histogram plot illustrates Siglec-1 expression that is higher in cmMTB- (black) than in cmCTR-treated (white) macrophages. (C) Representative immunofluorescence of Siglec-1 intracellular staining (green), actin (red) and nuclei (blue) after 3 days of monocyte conditioning with cmMTB. Scale bar, 10 μm. (D) Vertical scatter plot showing the quantification of Siglec-1 intracellular staining in cmCTR- or cmMTB-treated cells in the presence of an IFNAR-2 blocking (α-IFNAR) or control (α-IgG) antibodies during cell conditioning. Each circle represents a single donor and histograms median value. (E) Median fluorescence intensity (MFI) of Siglec-1 cell-surface expression in human macrophages exposed to cmCTR and infected with HIV-1 assessed by flow cytometry. The histogram plot illustrates Siglec-1 expression that increases within days post-HIV-1 infection of cmCTR-macrophages. Statistical analyses: Two-tailed, Wilcoxon signed-rank test (D). *p<0.05, **p<0.01, ns: not significant.
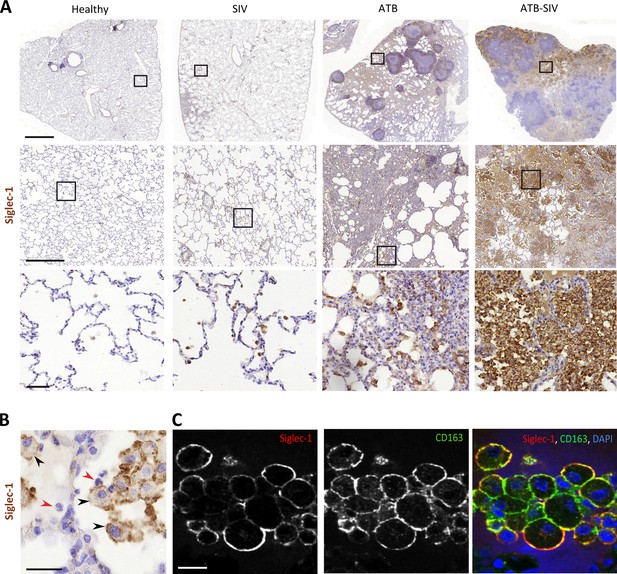
Tuberculosis-associated microenvironments increase Siglec-1 expression in non-human primate alveolar macrophages.
(A) Accumulation of Siglec-1+ alveolar macrophages in the lung of co-infected non-human primates (NHP). Representative immunohistochemical images of Siglec-1 staining (brown) in lung biopsies of healthy, SIV-infected (SIV), active tuberculosis (ATB) and co-infected with SIV (ATB-SIV) NHP. Scale bars from top to bottom: 2 mm, 500 µm and 50 µm. (B–C) Siglec-1+ cells display the alveolar macrophage morphology. (B) Representative immunohistochemistry image from lung biopsy of an ATB-SIV NHP stained for Siglec-1 (brown). Siglec-1+ cells display a cell morphology with a single nucleus and large cytoplasm reminiscent of macrophage (black arrowhead); Siglec-1- cells display a different nucleus morphology and small cytoplasm reminiscent of neutrophils (red arrowhead). Scale bar, 20 µm. (C) Representative immunofluorescence images of alveolar macrophages found in lung biopsy of a representative ATB-SIV NHP stained for Siglec-1 (red), CD163 (green) and nuclei (DAPI, blue). Scale bar, 20 µm.
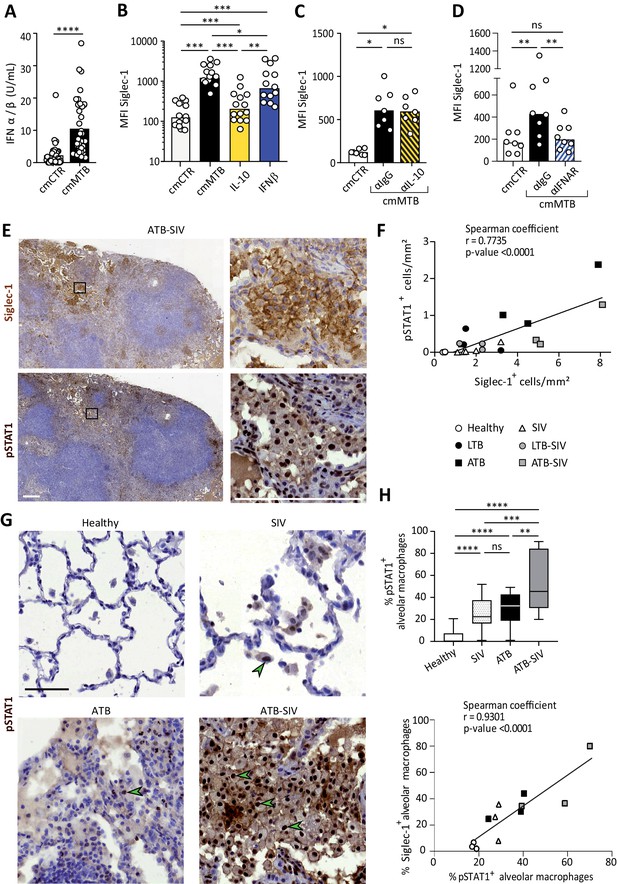
Siglec-1 expression is dependent on Mtb-induced type I IFN signaling.
(A) Vertical scatter plot showing the relative abundance of IFN-I in cmCTR (white) and cmMTB (black) media, as measured indirectly after 24 hr exposure to the HEK-Blue IFN-α/β reporter cell line yielding reporter activity in units (U) per mL. (B–D) Vertical scatter plots displaying the median fluorescent intensity (MFI) of Siglec-1 cell-surface expression after three days of monocyte differentiation into macrophages either with cmMTB (black) or cmCTR (white), the indicated recombinant cytokines (B), the presence of an IL-10 depletion (α-IL-10) or a control (α-IgG) antibodies (C), or the presence of an IFNAR-2 blocking (α-IFNAR) or control (α-IgG) antibodies (D). (E) Representative serial immunohistochemical images of lung biopsies of a co-infected (ATB-SIV) NHP stained for Siglec-1 (brown, top) and pSTAT1 (brown, bottom). Scale bar, 250 µm. Insets are 10x zooms. (F) Correlation of the percentage of cells positive for Siglec-1 and pSTAT1, as measured per mm2 of lung tissue from the indicated NHP groups. Mean value is represented as a black line. (G) Representative immunohistochemical images of lung biopsies from the indicated NHP group stained for pSTAT1 (brown). Arrowheads show pSTAT1-positive nuclei. Scale bar, 500 µm. (H) Upper panel: Vertical Box and Whisker plot illustrating the percentage of pSTAT1+ alveolar macrophages in lung biopsies from the indicated NHP groups. Quantification analysis from n = 600 alveolar macrophages grouped from three independent animals per NHP group. Lower panel: Correlation of the percentage of alveolar macrophages positive for Siglec-1 and pSTAT1, from the indicated NHP groups. Mean value is represented as a black line. (A–D) Each circle within vertical scatter plots represents a single donor and histograms median value. Statistical analyses: Two-tailed, Wilcoxon signed-rank test (A–D), Spearman correlation (F, H lower panel), and Mann-Whitney unpaired test (H, upper panel). *p<0.05, **p<0.01, ***p<0.001, ****p<0.0001. ns: not significant. See Figure 2—source data 1.
-
Figure 2—source data 1
Raw data and statistical analyses supporting that IFN-I induced by M. tuberculosis is responsible for Siglec-1 expression in human and non-human primate macrophages.
- https://cdn.elifesciences.org/articles/52535/elife-52535-fig2-data1-v2.xlsx
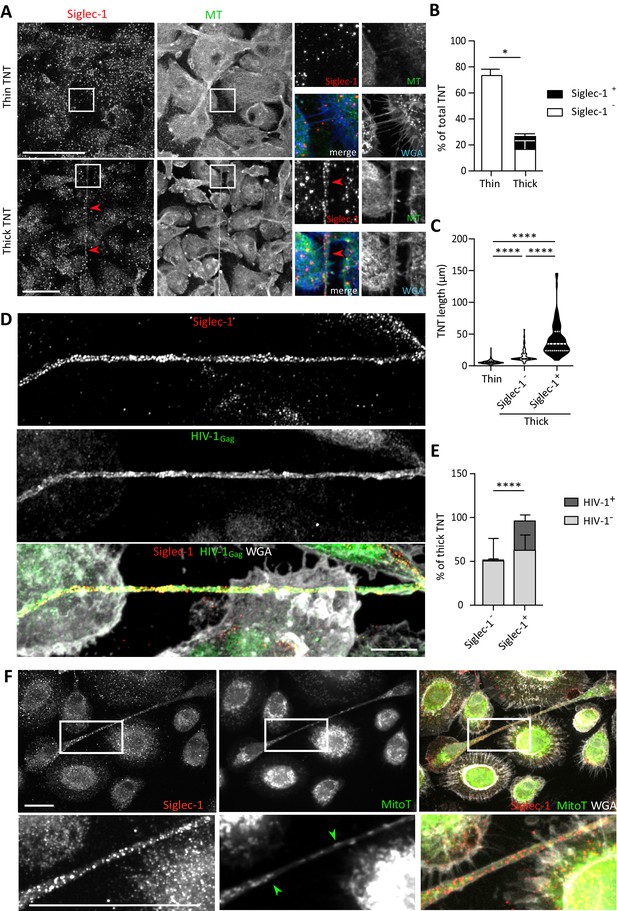
Siglec-1 localization on thick TNT is associated with their length and HIV-1/mitochondria cargo.
(A–F) Human monocytes were differentiated into macrophages with cmMTB for 3 days, and then infected with HIV-1-ADA strain (unless indicated otherwise) and fixed 3 days post-infection. (A) Representative immunofluorescence images of cmMTB-treated macrophages infected with HIV-1-ADA, and stained for extracellular Siglec-1 (red), intracellular tubulin (MT, green) and Wheat Germ Agglutinin (WGA, blue). Inserts are 3x zooms. Red arrowheads show Siglec-1 localization on TNT. Scale bar, 20 µm. (B) Vertical bar plot showing the semi-automatic quantification of Siglec-1+ TNT (black) and Siglec-1- TNT (white) in thick (WGA+, MT+) and thin (WGA+, MT-) TNT. 400 TNT were analyzed from two independent donors. (C) Siglec-1+ TNT exhibit a larger length index. Violin plots displaying the semi-automatic quantification of TNT length (in μm) for thin (WGA+, MT-), and thick TNT (WGA+, MT+) expressing Siglec-1 or not. 400 TNT were analyzed per condition from two independent donors. (D) Representative immunofluorescence images of cmMTB-treated macrophages 3 day post-infection with HIV-1-NLAD8-VSVG strain, and stained for extracellular Siglec-1 (red), intracellular HIV-1Gag (green) and WGA (grey). Scale bar, 10 µm. (E) Vertical bar plots indicating the quantification of presence (dark grey) or absence (light grey) of HIV-1Gag in thick TNT (WGA+, MT+) expressing Siglec-1 or not. 120 TNT in at least 1000 cells were analyzed from four independent donors. (F) Representative immunofluorescence images of cmMTB-treated macrophages infected with HIV-1-ADA loaded with MitoTracker (MitoT, green), and stained for extracellular Siglec-1 (red) and WGA (grey). Inserts are 3x zooms. Green arrowheads show mitochondria inside TNT. Scale bar, 10 µm. Statistical analyses: Two-way ANOVA comparing the presence of Siglec-1 in thin and thick TNT (B), and two-tailed Mann-Whitney unpaired test comparing TNT length (C) and the presence of HIV-1 in TNT (E). *p<0.05, ****p<0.0001. See Figure 3—source data 1.
-
Figure 3—source data 1
Raw data and statistical analyses supporting Siglec-1 expression on thick TNT and its correlation with TNT length.
- https://cdn.elifesciences.org/articles/52535/elife-52535-fig3-data1-v2.xlsx
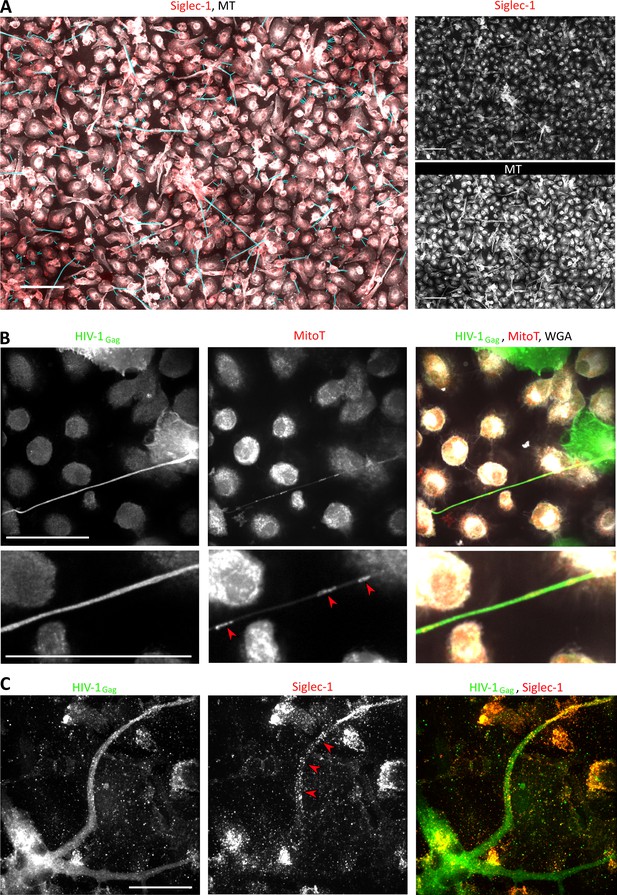
Siglec-1 localizes specifically on thick tunneling nanotubes that contain HIV-1Gag and mitochondria.
(A–C) Human monocytes were differentiated into macrophages with cmMTB for 3 days, infected with HIV-1-ADA strain (unless indicated otherwise) and then fixed at day 3 (A–B) or 14 (C) post-infection. (A) Representative immunofluorescence images used for semi-automatic quantification of TNT in cmMTB-treated macrophages infected with HIV-1. Cells were stained for extracellular Siglec-1 (red), intracellular tubulin (MT, grey) and Wheat Germ Agglutinin (WGA, not shown). Blue lines show all TNT considered. Thick (WGA+, MT+) and thin (WGA+, MT-) TNT were assessed for Siglec-1 positivity by applying a threshold and measured in length. Scale bar, 20 µm. (B) Representative immunofluorescence images of cmMTB-treated macrophages infected with HIV-1-NLAD8-VSVG, loaded with Mitotracker (MitoT, red) and stained for intracellular HIV-1Gag (green) and WGA (grey). Red arrowheads show mitochondria inside HIV-1Gag-containing TNT. Inserts are 2x zoom. Scale bar, 20 µm. (C) Representative immunofluorescence images of cmMTB-treated macrophages infected with HIV-1 and kept in culture until day 14. Cells were fixed and stained for intracellular HIV-1Gag (green), extracellular Siglec-1 (red) and WGA (grey). Red arrowheads show Siglec-1 on HIV-1Gag-containing TNT emanating from an infected multinucleated giant cell (MGC). Scale bar, 20 µm.
Related to Figure 3A.
Z-stack of confocal microscopy images, showing Siglec-1 (red), microtubules (MT, green) and F-actin (grey) of day 6 HIV-1-infected macrophages, treated with cmMTB. Siglec-1 localizes on the thick MT+ F-actin+ TNT but not on thin MT- F-actin+ TNT.
Related to Figure 3D.
3D reconstitution of confocal images, showing Siglec-1 (red), HIV-1Gag (green) and WGA (grey) of day 6 HIV-1-infected macrophages, treated with cmMTB.
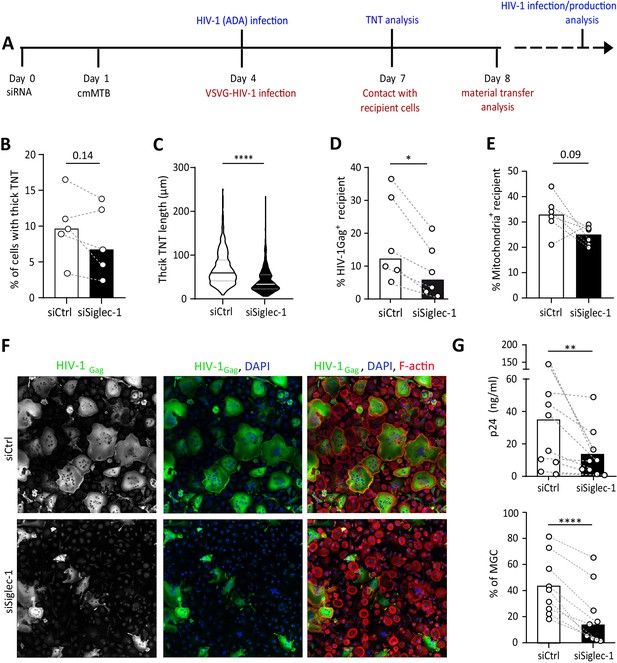
The exacerbation of HIV-1 infection and spread in macrophages treated with cmMTB requires Siglec-1.
(A) Experimental design. Monocytes from healthy subjects were transfected with siRNA targeting of Siglec-1 (siSiglec-1, black) or not (siCtrl, white). A day after, monocytes were differentiated into macrophages with cmMTB for 3 days. Cells were then infected with HIV-1-ADA (blue protocol) to measure the formation (B) and length (C) of TNT at day 7, or assess HIV-1 production and multinucleated giant cell (MGC) formation at day 14 (F–G). In parallel, cells were either infected with HIV-NLAD8-VSVG or labelled with mitoTracker to measure the transfer (red protocol) of HIV-1 (D) or mitochondria (E), respectively. (B) Before-and-after plots showing the percentage of cells forming thick TNT (F-actin+, WGA+, MT+). (C) Violin plots displaying the semi-automatic quantification of TNT length (in μm) for thick (WGA+, MT+) TNT; 300 TNT were analyzed per condition from two independent donors. (D–E) Before-and-after plots indicating the percentage of HIV-1Gag+ cells (D) or MitoTracker+ cells (E) among CellTracker+ cells after 24 hr co-culture. (F) Representative immunofluorescence images of siRNA transfected cells treated with cmMTB, 14 days post-HIV-1 infection. Cells were stained for intracellular HIV-1Gag (green), F-actin (red) and DAPI (blue). Scale bar, 500 µm. (G) Vertical scatter plots showing HIV-1-p24 concentration in cell supernatants (upper panel) and percentage of MGC (lower panel) at day 14 post-HIV-1 infection in cells represented in F (siSiglec-1, black; siCtrl, white). (B, D, E and G) Each circle represents a single donor and histograms median value. Statistical analyses: Paired t-test (B, G lower panel) or two-tailed, Wilcoxon signed-rank test (C-E, G upper panel). *p<0.05, **p<0.01, ****p<0.0001. See Figure 4—source data 1.
-
Figure 4—source data 1
Raw data and statistical analyses supporting a role for Siglec-1 in TNT length, HIV-1 and mitochondrial cell-to-cell trasfer, and exacerbation of HIV-1 infection.
- https://cdn.elifesciences.org/articles/52535/elife-52535-fig4-data1-v2.xlsx
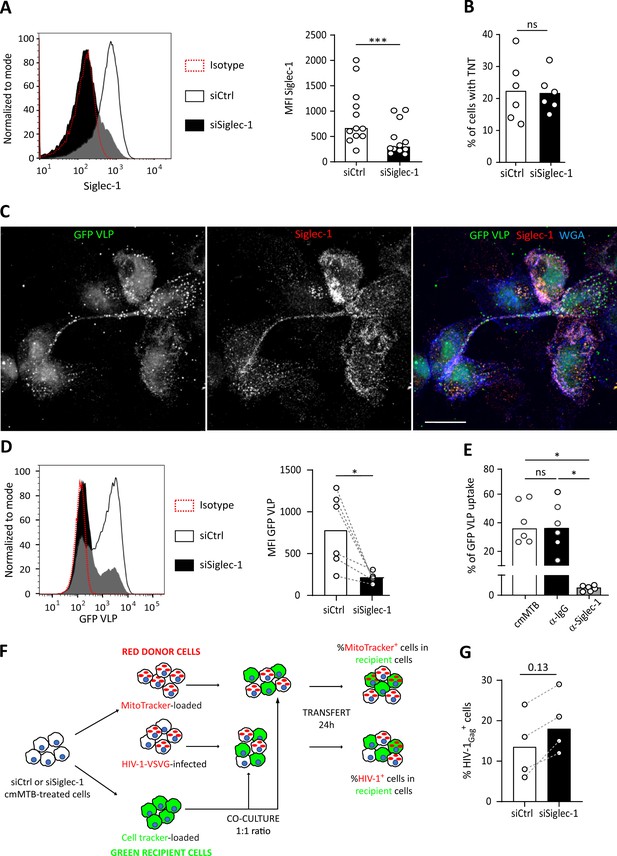
Siglec-1 is required for the capture and transfer of HIV-1 in cmMTB-treated macrophages.
(A–D, F–G) Monocytes from healthy subjects were transfected with siRNA targeting of Siglec-1 (siSiglec-1, black) or not (siCtrl, white). A day after, monocytes were differentiated into macrophages with cmMTB for 3 days. (A) Representative histogram (left) and vertical scatter plot showing the median fluorescent intensity (MFI) (right) of Siglec-1 expression on the indicated cell populations. (B) Vertical scatter plot indicating the percentage of cells forming TNT. (C–E) Inhibition of Siglec-1 reduces binding of HIV-1-Gag−eGFP virus like particles (GFP-VLP). (C) Representative immunofluorescence images of cmMTB-treated cells incubated with GFP-VLP (green) for 3.5 hr. Cells were fixed and stained for extracellular Siglec-1 (red) and Wheat Germ Agglutinin (WGA, blue). Scale bar, 500 µm. (D) Representative histogram (left) and vertical scatter plot showing the median fluorescent intensity (MFI) (right) displaying of GFP-VLP binding in the indicated cell populations. (E) Vertical scatter plot showing the percentage of GFP-VLP binding in cmMTB treated cells pre-incubated with specific anti-Siglec-1 (α-Siglec-1, grey), anti-Isotype control antibodies (α-IgG, black) or mock (white). (F) Schematics of the experimental procedure for material (HIV-1 and mitochondria) transfer experiments. (G) Vertical scatter plot showing the percentage of HIV-1Gag+ cells at the time of co-culture experiment in the indicated cells. Statistical analyses: Two-tailed, Wilcoxon matched-pairs signed rank test (A, B, D-E, G). *p<0.05, ***p<0.001. ns: not significant.
-
Figure 4—figure supplement 1—source data 1
Supplemental raw data and statistical analyses supporting the functional role of Siglec-1 in human macrophages using an siRNA-mediated gene silencing approach.
- https://cdn.elifesciences.org/articles/52535/elife-52535-fig4-figsupp1-data1-v2.xlsx
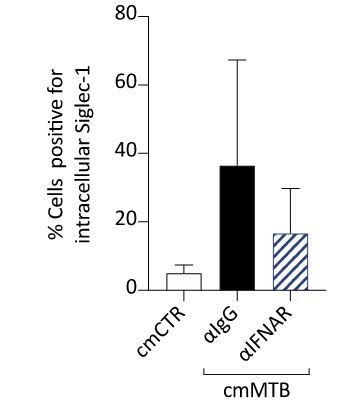
Immunofluorescence analysis of cmMTB-treated cells positive for Siglec-1.
For 3 days, monocytes were differentiated into macrophages either with cmCTR (white) or with cmMTB in the presence of an IFNAR-2 blocking (α-IFNAR, stripped) or control (α-IgG, black) antibodies. Immunofluorescence analyses were performed and the percentage of cells positive for intracellular Siglec-1 were quantified. n=4 donors.
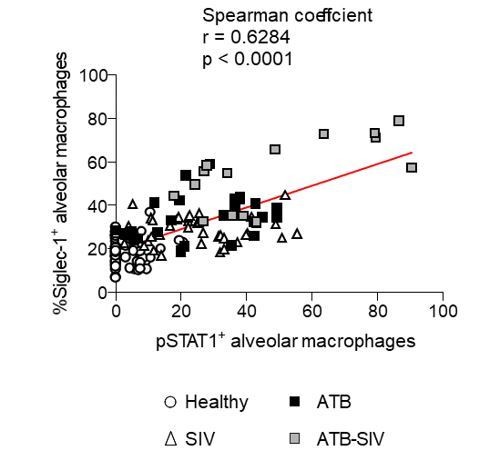
Correlation of alveolar macrophages that are positive for either Siglec-1 or phospho-STAT1 in NHP (Related to Figure 2H, bottom).
Each symbol represents an image field containing the lung alveoli that was obtained from the three independent animals per NHP group. Mean value is represented as a red line.
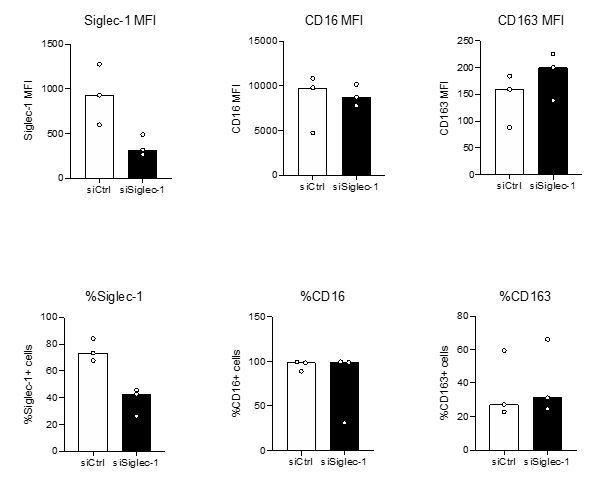
Inactivation of Siglec-1 by siRNA-mediated gene silencing does not affect the expression of M(IL-10) markers in cmMTB-treated cells (Related to Figure 4—figure supplement 1A).
Monocytes from healthy subjects were transfected with siRNA targeting of Siglec-1 (siSiglec-1, black) or not (siCtrl, white). A day after, monocytes were differentiated into macrophages with cmMTB for 3 days. Vertical scatter plots showing the median fluorescent intensity (MFI, upper panels) of Siglec-1, CD16 and CD163, and the percentage of cells expressing Siglec-1, CD16 and CD163 (lower panels). Each circle represents a single donor and histograms median values. n=3 donors.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| M. tuberculosis | H37Rv | Derived from E.R. Baldwin's human-lung isolate H37 by W. Steenken New York, United States, 1934 | (ATCC 25618) | |
| HIV-1 | ADA | Gift from Dr. S Benichou Institut Cochin, Paris, France | N/A | |
| HIV-1 | NLAD8 | Gift from Dr. S Benichou Institut Cochin, Paris, France | N/A | |
| HIV-1 | ADA Gag-iGFP-VSVG | This paper | This paper | |
| Buffy Coat | Leukocytes | Etablissement Français du Sang, Toulouse, France | N/A | |
| Lung biopsies from rhesus macaques | Histological slides | Tulane National Primate Research Center | N/A | |
| Cell line (human HeLa JC.53) | TZM-bl | NIH AIDS Reagent Program | Cat# 8129 | Cultured in: DMEM, 90%; FBS, 10%; 100 units of Penicillin and 0.1 mg/mL of Streptomycin |
| Cell line (human) | HEK-293T | NIH AIDS Reagent Program | Cat# 3318 | Cultured in: DMEM, 10% FCS |
| Cell line (human) | HEK-Blue IFN-α/β Cells | Invivogen | Cat# hkb-ifnab | Cultured in: DMEM, 4.5 g/l glucose, 2 mM L-glutamine, 10% (v/v) heat-inactivated fetal bovine serum, 100 U/ml penicillin, 100 µg/mL streptomycin, 100 µg/mL Normocin |
| Transfected construct (human) | siRNA to Siglec-1 (SMART-Pool) | Horizon Discovery | Cat# L-017521-01-0020 | (200 nM) |
| Transfected construct (human) | siRNA scramble (SMART Pool) | Horizon Discovery | Cat# D-001810-10-50 | (200 nM) |
| Antibody | Mouse monoclonal anti‑human Siglec-1 (clone 7-293) | Biolegend | Cat# 346008 RRID:AB_11147948 | FACS (1 µg/mL) |
| Antibody | Mouse monoclonal anti‑human CD16 (clone 3G8) | Biolegend | Cat# 302019 and 302018; RRID:AB_492974 andAB_314218 | FACS (1 µg/mL) |
| Antibody | Mouse monoclonal anti‑human CD163 (clone GHI/61) | Biolegend | Cat# 333608 RRID:AB_2228986 | FACS (1 µg/mL) |
| Antibody | Mouse monoclonal anti‑human MerTK (clone 590H11G1E3) | Biolegend | Cat# 367607 RRID:AB_2566400 | FACS (1 µg/mL) |
| Antibody | Rabbit monoclonal anti‑human STAT1 (clone 42H3) | Cell Signaling Technology | Cat# 9175 RRID:AB_2197984 | WB (1:100) |
| Antibody | Rabbit anti‑human actin (a.a. 20‑33) | Sigma‑Aldrich | Cat# A5060 RRID:AB_476738 | WB (1:100) |
| Antibody | Rabbit polyclonal anti-a-tubulin | Abcam | Cat# ab18251 RRID:AB_2210057 | IF (5 µg/mL) |
| Antibody | Mouse monoclonal anti-Siglec-1 (clone hsn 7D2) | Novus Biologicals | Cat# NB 600-534 RRID:AB_526814 | IF (10 µg/mL) IHC (1:200) |
| Antibody | Mouse monoclonal anti-Gag RD1 (clone KC57) | NIH AIDS Reagent program | Cat# 13449 | IF (1:200) |
| Antibody | Mouse monoclonal anti-HIV-1 p24 (clone 183-H12-5C) | NIH AIDS Reagent Program | Cat# 3537 | ELISA (2.5 µg/mL) |
| Antibody | Human polyclonal anti-HIV Immune Globulin (HIVIG) | NIH AIDS Reagent Program | Cat# 3957 | ELISA (6.25 µg/mL) |
| Antibody | Polyclonal goat anti-human IgG | Sigma-Aldrich | Cat# A0170 | ELISA (1:10000) |
| Antibody | Mouse monoclonal anti‑human CD163 (clone 10D6) | Leica/Novocastra | Cat# NCL-L-CD163 RRID:AB_2756375 | IHC (1:100) |
| Antibody | Anti-pSTAT1 | Cell Signaling Technology | Cat# 9167 RRID:AB_561284 | WB (1:100) |
| Antibody | Mouse monoclonal anti-IFNAR2 (clone MMHAR-2) | Thermo Fisher Scientific | Cat# 213851 RRID:AB_223508 | Blocking (20 µg/mL) FACS (1 µg/mL) |
| Antibody | Mouse IgG2a isotype control | Thermo Fisher Scientific | Cat# 02-6200 RRID:AB_2532943 | Blocking (20 µg/mL) IF (0.6 µg/mL) |
| Antibody | Polyclonal F(ab)2 goat anti‑rabbit IgG, AlexaFluor 555 | Thermo Fisher Scientific | Cat# A-21430 RRID:AB_2535851 | IF (2 µg/mL) |
| Antibody | Polyclonal F(ab)2 goat anti‑mouse IgG, AlexaFluor 488 | Thermo Fisher Scientific | Cat# A-10684 RRID:AB_2534064 | IF (2 µg/mL) |
| Antibody | Plyclonal F(ab)2 goat anti‑mouse IgG, AlexaFluor 555 | Cell Signaling Technology | Cat# 4409 RRID:AB_1904022 | IF (2 µg/mL) |
| Antibody | Polyclonal goat anti‑rabbit IgG, HRP | Thermo Fisher Scientific | Cat# 32460 RRID:AB_1185567 | WB (1:10000) |
| Antibody | Polyclonal goat anti‑mouse IgG, HRP | Thermo Fisher Scientific | Cat# 31430 RRID:AB_228307 | WB (1:10000) |
| Cytokine (recombinant, human) | M-CSF | Peprotech | Cat# 300‑25 | (20 ng/mL) |
| Cytokine (recombinant, human) | IFNb | Peprotech | Cat# 300-02BC | 10 and 100 U/mL |
| Cytokine (recombinant, human) | IL-10 | Peprotech | Cat# 200-10 | 10 ng/mL |
| Monocyte isolation | Mouse anti‑human CD14 microbeads | Miltenyi Biotec | Cat# 130‑050‑201 | |
| Monocyte isolation | LS magnetic columns | Miltenyi Biotec | Cat# 130‑042‑401 | |
| Western blot | Amersham ECL Prisme Western Blotting Detection Reagent | GE Healthcare | Cat# RPN2232 | |
| Western blot | SuperSignal WestPico Chemiluminescent Substrate | Thermo Scientific | Cat# 34080 | |
| ELISA | IL‑10 ELISA set | BD Bioscience | Cat# 555157 | |
| Cell culture | Trypsin EDTA 0.05% | Thermo Fisher Scientific | Cat# 25200072 | |
| Cell culture | Accutase | Sigma-Aldrich | Cat# A-6964 | |
| Probe | Phalloidin AlexaFluor 488 | Thermo Fisher Scientific | Cat# A12379 | (33 mM) |
| Probe | Phalloidin Alexa Fluor 647 | Thermo Fisher Scientific | Cat# A22287 | (33 mM) |
| Probe | DAPI | Sigma Aldrich | Cat# D9542 | (500 ng/mL) |
| Probe | CellTracker Green CMFDA Dye | Thermo Fisher Scientific | Cat# C7025 | (500 ng/mL) |
| Probe | MitoTracker Deep Red FM | Invitrogen | Cat# M22426 | (500 ng/mL) |
| IF | Fluorescence Mounting Medium | Agilent Technologies | Cat# S302380‑2 | |
| IF | Antibody diluent, Background reducing | DAKO, Agilent Technologies | Cat# S302283-2 | |
| Software | ImageJ | ImageJ | http://www.imagej.nih.gov/ij | |
| Software | Prism (v8.0.0) | GraphPad | http://www.graphpad.com | |
| Software | Photoshop CS3 | Adobe | http://www.adobe.com | |
| Software | Adobe Illustrator CS5 | Adobe | https://www.adobe.com/fr/products/illustrator.html | |
| Software | Huygens Professional Version 16.10 | Scientific Volume Imaging | https://svi.nl/HuygensProfessional | |
| Software | FACS DIVA | BD Bioscience | http://www.bdbiosciences.com/ | |
| Software | FlowJo_v10 | FlowJo | https://www.flowjo.com/ | |
| Software | FCS Express V3 | DeNovo Software | http://www.denovosoftware.com | |
| Software | Image Lab | Bio‑Rad Laboratories | http://www.bio‑rad.com | |
| Software | Pannoramic Viewer | 3DHISTECH | https://www.3dhistech.com/pannoramic_viewer |
Additional files
-
Supplementary file 1
Clinical data of NHPs (Table S1) and histopathological scoring of lung lesions in NHPs (Table S2).
- https://cdn.elifesciences.org/articles/52535/elife-52535-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/52535/elife-52535-transrepform-v2.docx





