Evolution of a plant gene cluster in Solanaceae and emergence of metabolic diversity
Figures
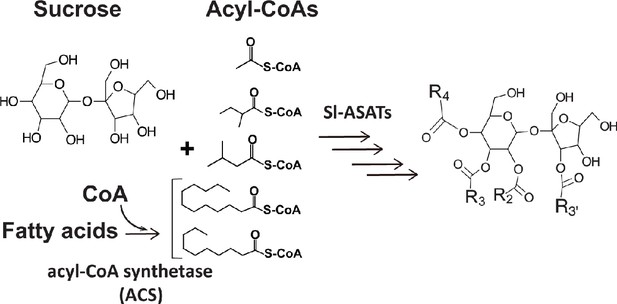
Primary metabolites are biosynthetic precursors of tomato trichome acylsugars.
In cultivated tomatoes, the trichome acylsucroses are synthesized by four Sl-ASATs using the primary metabolites – sucrose and different types of acyl-CoAs – as substrates. In this study we provide evidence that medium chain fatty acids are converted to acyl-CoAs by an acyl-CoA synthetase for medium chain acylsugar biosynthesis.
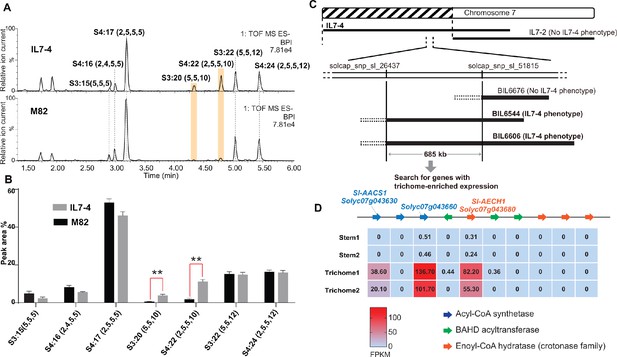
Mapping of a genetic locus related to acylsugar variations in tomato interspecific introgression lines.
(A) Electrospray ionization negative (ESI-) mode, base-peak intensity (BPI) LC/MS chromatogram of trichome metabolites from cultivated tomato S. lycopersicum M82 and introgression line IL7-4. The orange bars highlight two acylsugars that have higher abundance in IL7-4 than in M82. For the acylsucrose nomenclature, ‘S’ refers to a sucrose backbone, ‘3:22’ means three acyl chains with twenty-two carbons in total. The length of each acyl chain is shown in the parentheses. (B) Peak area percentage of seven major trichome acylsugars in M82 and IL7-4. The sum of the peak area percentage of each acylsugar is equal to 100% in each sample. The data is shown for three plants ± SEM. **p<0.01, Welch two-sample t test. Figure 2—source data 1 includes values for the analysis. (C) Mapping the genetic locus contributing to the IL7-4 acylsugar phenotype using selected backcross inbred lines (BILs) that have recombination break points within the introgression region of IL7-4. (D) Narrowing down candidate genes in the locus using trichome/stem RNA-seq datasets generated from previous study (Ning et al., 2015). A region with duplicated genes of three types – acyl-CoA synthetase (ACS), BAHD acyltransferase, and enoyl-CoA hydratase (ECH) – is shown. The red-blue color gradient provides a visual marker to rank the expression levels represented by Fragments Per Kilobase of transcript per Million mapped reads (FPKM). Coexpression analysis of tomato ACS, ECH, and BAHD acyltransferase family genes is shown in Figure 2—figure supplement 1.
-
Figure 2—source data 1
Data used to make Figure 2B.
Peak area percentage of seven major trichome acylsugars in M82 and IL7-4.
- https://cdn.elifesciences.org/articles/56717/elife-56717-fig2-data1-v2.xlsx
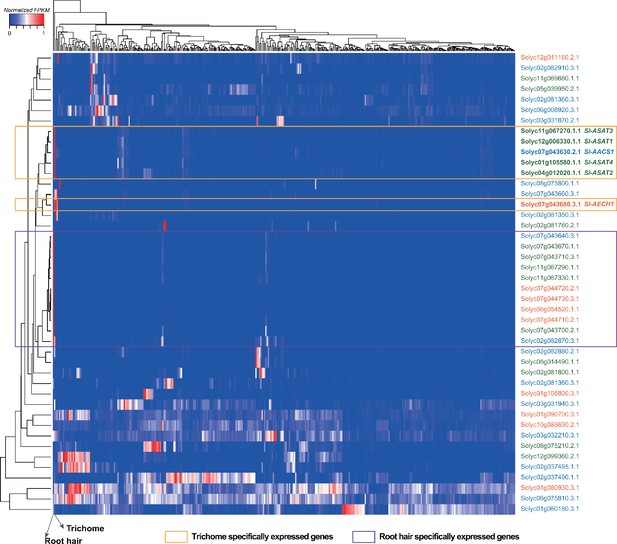
Expression profiles of tomato ACS, ECH, and BAHD acyltransferase family genes used for phylogenetic analysis in this study.
Each column represents one transcriptomic profiling dataset generated by RNA-seq analysis using samples from the cultivated tomato (S. lycopersicum). A total of 372 RNA-seq datasets were used for the analysis as described in the previous study (Moore et al., 2020). The normalized FPKM value of each gene across all the 372 datasets was illustrated by the color scale. The maximum FPKM value was set to 1 (red), while the minimum FPKM value was 0 (blue). The datasets generated using tomato root hairs (first column) or trichomes (second column) were indicated by arrows. Genes (y-axis) and expression datasets (x-axis) were both grouped using hierarchical clustering. Genes involved in acylsugar biosynthesis, such as Sl-ASATs, Sl-AACS1, and Sl-AECH1, were clustered together, which are highlighted by the orange box. Hierarchical clustering also revealed another group of genes that are root hair-specific as pointed out by the purple box. The gene ID was colored based on which gene family it belongs. Blue: ACS genes; red: ECH genes; green: BAHD acyltransferase genes.
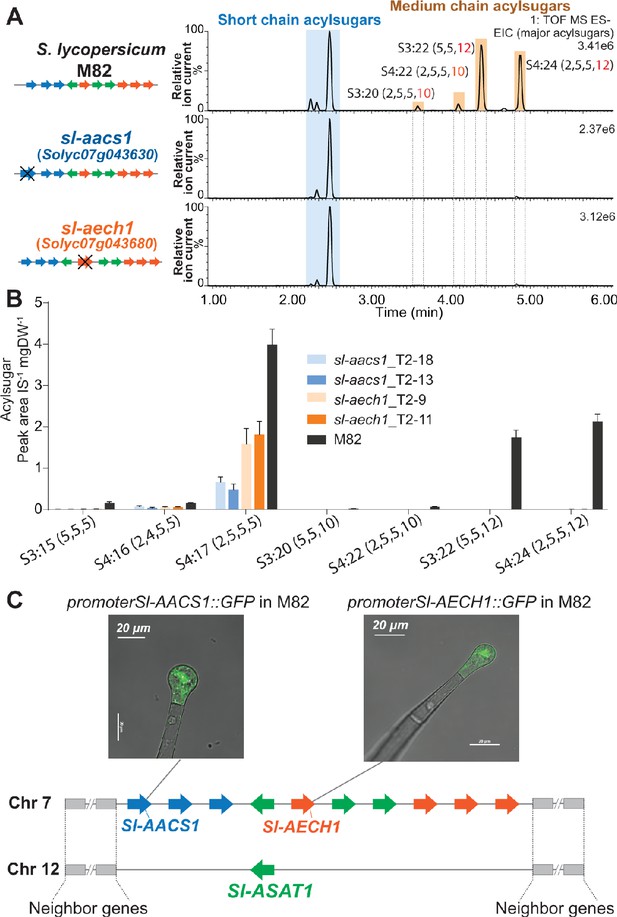
CRISPR/Cas9-mediated gene knockout of tomato Sl-AACS1 or Sl-AECH1 eliminates detectable medium chain containing acylsugars.
(A) Combined LC/MS extracted ion chromatograms of trichome metabolites from CRISPR mutants sl-aacs1 and sl-aech1. The medium chain acylsugars that are not detected in the two mutants are denoted by pairs of vertical dotted lines. Figure 3—figure supplement 1 describes the design of the gRNAs and details of the gene edits. (B) Quantification of seven major trichome acylsugars in sl-aacs1 and sl-aech1 mutants. Two independent T2 generation transgenic lines for each mutant were used for analysis. The peak area/internal standard (IS) normalized by leaf dry weight (DW) is shown from six plants ± SEM. Figure 3—source data 1 includes values for the analysis. (C) Confocal fluorescence images showing that GFP fluorescence driven by Sl-AACS1 or Sl-AECH1 is located in the tip cells of type I/IV trichomes. Their tissue specific expressions are similar to Sl-ASAT1 (Fan et al., 2016), which locates in a chromosome 12 region that is syntenic to the locus containing Sl-AACS1 and Sl-AECH1. Figure 3—figure supplement 2 provides the detailed information of the syntenic region. Sl-AACS1, Sl-AECH1, and Sl-ASAT1 are the only gene models with demonstrated functions in acylsugar biosynthesis.
-
Figure 3—source data 1
Data used to make Figure 3B.
Quantification of seven major trichome acylsugars in the CRISPR mutants sl-aacs1 and sl-aech1, as well as the parent M82.
- https://cdn.elifesciences.org/articles/56717/elife-56717-fig3-data1-v2.xlsx
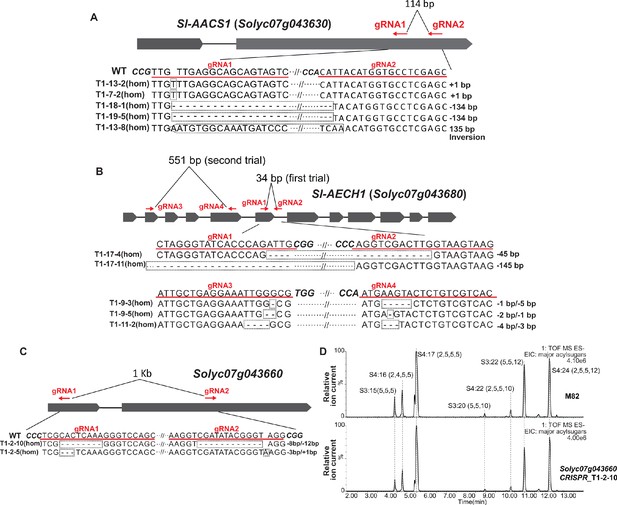
CRISPR-Cas9-mediated gene knockouts in cultivated tomato S. lycopersicum.
The gRNAs targeting Sl-AACS1 (A), Sl-AECH1(B), and Solyc07g043660 (C) in cultivated tomato are highlighted with red lines and text. Two pairs of gRNAs were designed that target Sl-AECH1, which were followed by two trials of plant transformation. DNA sequences of the self-crossed T1 generation transgenic lines carrying homozygous gene edits are shown beneath the gene model. Dotted rectangle boxes highlight edited sequences. (D) Electrospray ionization negative (ESI-) mode, LC/MS extracted ion chromatograms of seven major trichome acylsugars of the CRISPR mutant solyc07g043660 and the M82 parent.
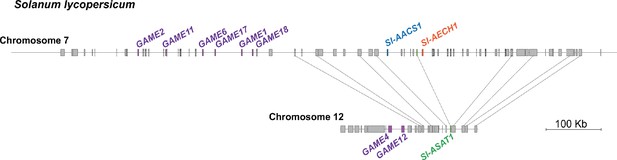
The syntenic region of cultivated tomato S. lycopersicum chromosome 7 and 12 harboring the acylsugar and steroidal glycoalkaloid gene clusters.
The gene models are represented by rectangles, with pseudogenes labeled with dotted lines. The lines linking the gene models of the two chromosomes denote putative orthologous genes in the synteny. GAME genes involved in glycoalkaloid metabolism were previously reported (Itkin et al., 2013).
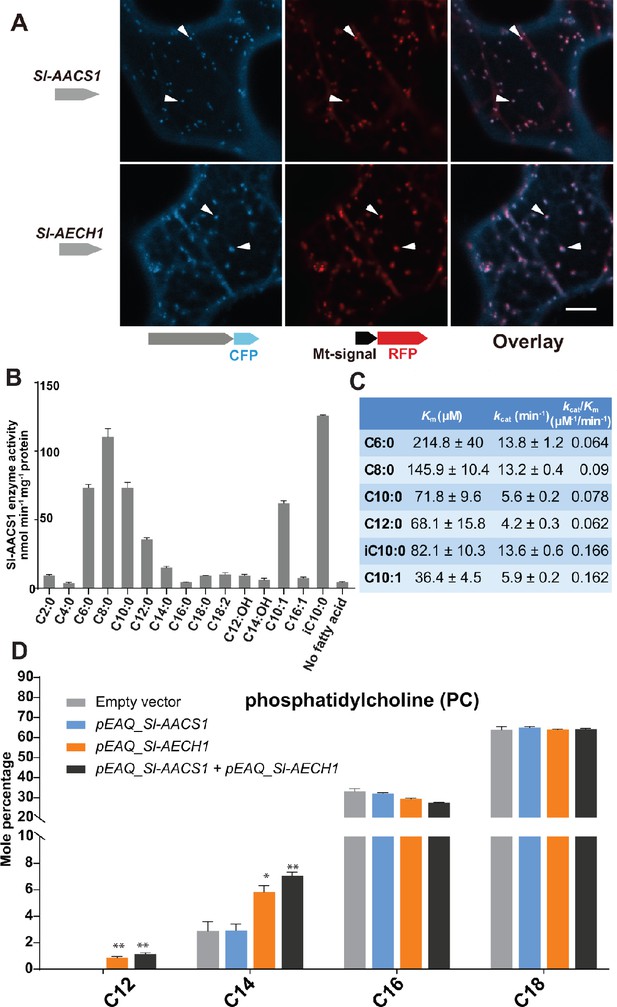
Functional analysis of Sl-AACS1 and Sl-AECH1 in N. benthamiana and recombinant Sl-AACS1 enzyme analysis.
(A) Confocal images of co-expression analysis in tobacco leaf epidermal cells using C-terminal CFP-tagged either Sl-AACS1 or Sl-AECH1 and the mitochondrial marker MT-RFP. Arrowheads point to mitochondria that are indicated by MT-RFP fluorescent signals. Scale bar equals 10 μm. Figure 4—figure supplement 1B describes that the expressed YFP-recombinant proteins were not co-localized with the peroxisomal marker RFP-PTS (B) Aliphatic fatty acids of different chain lengths were used as the substrates to test Sl-AACS1 acyl-CoA synthetase activity. Mean amount of acyl-CoAs generated (nmol min−1 mg−1 proteins) was used to represent enzyme activities. The results are from three measurements ± SEM. Figure 4—source data 1 includes values for the measurements. (C) Enzyme activity of Sl-AACS1 for six fatty acid substrates. (D) Identification of membrane lipid phosphatidylcholine (PC), which contains medium acyl chains, following transient expression of Sl-AECH1 in N. benthamiana leaves. The results from expressing Sl-AACS1 and co-expressing both Sl-AECH1 and Sl-AACS1 are also shown. Mole percentage (Mol %) of the acyl chains from membrane lipids with carbon number 12, 14, 16, and 18 are shown for three biological replicates ± SEM. *p<0.05, **p<0.01. Welch two-sample t test was performed comparing with the empty vector control. Figure 4—source data 2 includes values for the lipid analysis. Acyl groups of the same chain lengths with saturated and unsaturated bonds were combined in the calculation. Figure 4—figure supplement 2 shows that the putative Sl-AECH1 orthologs from S. pennellii and S. quitoense generated medium chain lipids in the infiltrated leaves.
-
Figure 4—source data 1
Data used to make Figure 4B.
Aliphatic fatty acids of different chain lengths were used as the substrates to test Sl-AACS1 acyl-CoA synthetase activity.
- https://cdn.elifesciences.org/articles/56717/elife-56717-fig4-data1-v2.xlsx
-
Figure 4—source data 2
Data used to make Figure 4D and Figure 4—figure supplement 1C.
N. benthamiana leaf membrane lipid acyl chain composition.
- https://cdn.elifesciences.org/articles/56717/elife-56717-fig4-data2-v2.xlsx
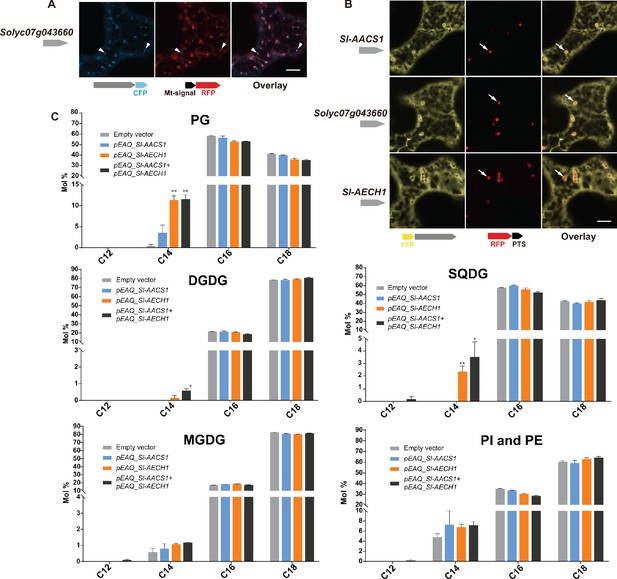
Characterization of cluster genes using leaf transient expression: protein subcellular targeting and impacts on lipid metabolism.
(A) Confocal images of co-expression analysis in N. tabacum leaf epidermal cells using C-terminal CFP-tagged Solyc07g043660 and the mitochondrial marker MT-RFP. Arrowheads point to mitochondria that are indicated by MT-RFP fluorescent signals. White bar equals 10 μm. (B) Confocal images of co-expression analysis in N. tabacum leaf epidermal cells using N-terminal YFP-tagged Sl-AACS1, Sl-AECH1, or Solyc07g043660, and the peroxisomal marker RFP-PTS. Arrowheads point to peroxisomes that are indicated by RFP-PTS fluorescent signals. Bar equals 10 μm. (C) N. benthamiana leaf membrane lipid acyl chain composition. The results from infiltrating Sl-AACS1 or Sl-AECH1 individually, and infiltrating Sl-AECH1 and Sl-AACS1 together are shown. The lipid abbreviations are: phosphatidylglycerol (PG), sulfoquinovosyl diacylglycerol (SQDG), digalactosyldiacylglycerol (DGDG), monogalactosyldiacylglycerol (MGDG), phosphtatidylinositols (PI), and phosphtatidylethanolamine (PE). Mole percentage (Mol %) of the acyl chains from membrane lipids with carbon number 12, 14, 16, and 18 are shown for three biological replicates ± SEM. *p<0.05, **p<0.01. Welch two-sample t test was performed comparing each experimental with the empty vector control. PI and PE lipids were adjacent on the TLC plates and were pooled for analysis. Acyl groups of the same chain lengths with saturated and unsaturated bonds were combined in the calculation.
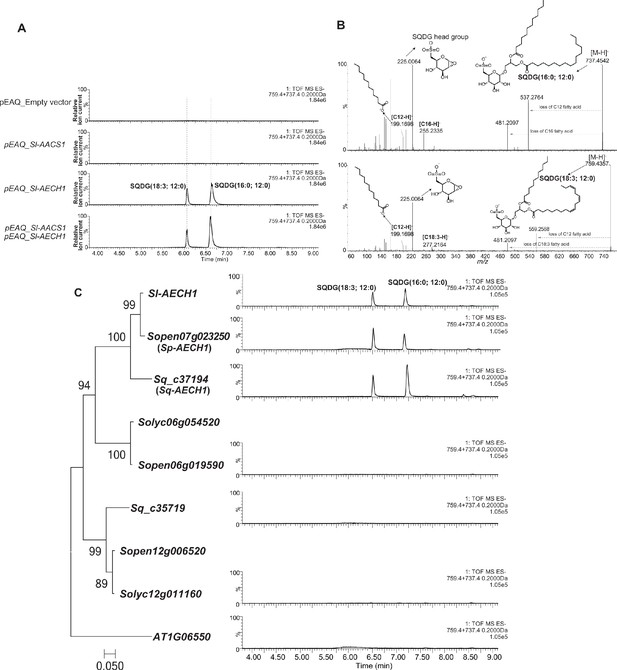
The closest homologs of Sl-AECH1 from other Solanum species generate medium chain lipids when transiently expressed in N. benthamiana.
(A) ESI- mode, LC/MS extracted ion chromatograms of two SQDGs with C12 as peaks diagnostic of lipids containing medium chain fatty acids. (B) Mass spectra of two SQDGs contain C12 acyl chain. Fragmentation of SQDG (16:0, 12:0) and SQDG (18:3, 12:0) in ESI- mode revealed the fragment ion C12 fatty acid (m/z: 199.17) and the SQDG head group (m/z: 225.0). (C) Among the Sl-AECH1 homologs tested, Sopen07g023250 (Sp-AECH1) and Sq_c37194 (Sq-AECH1), from S. pennellii and S. quitoense respectively, generated medium chain SQDG in the infiltrated leaves. The close homologs of Sl-AECH1 in S. lycopersicum, S. pennellii, and S. quitoense, which also have trichome expressions, were used to build the phylogenetic tree. The nucleotide sequences were aligned with MEGA7 (www.megasoftware.net) using the default MUSCLE algorithm. The T92+G maximum likelihood model was selected for phylogenetic tree construction from 24 different nucleotide substitution models based on the lowest Bayesian Information Criterion. The bootstrap values were obtained with 1000 replicates. The closest Sl-AECH1 Arabidopsis homolog AT1G06550 serves as an outgroup.
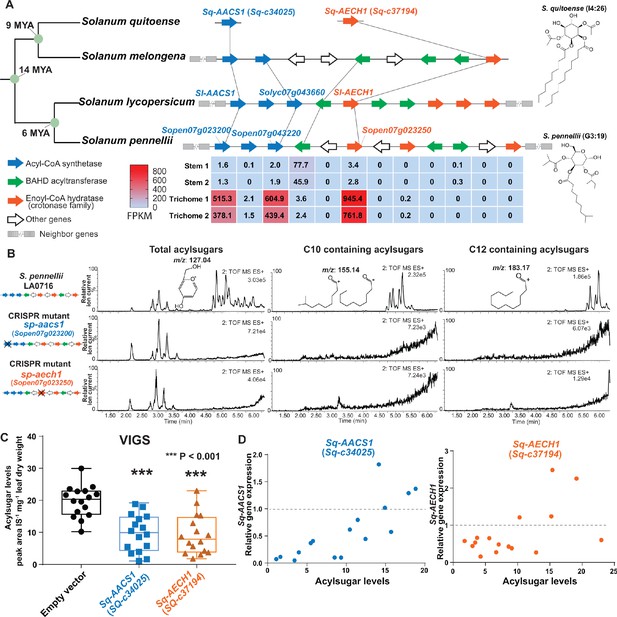
AACS1 and AECH1 are evolutionarily conserved in Solanum plants.
(A) A conserved syntenic genomic region containing AACS1 and AECH1 was found in three selected Solanum species. Nodes representing estimated dates since the last common ancestors (Särkinen et al., 2013) shown on the left. The closest homologs of AACS1 and AECH1 in Solanum quitoense are shown without genomic context because the genes were identified from RNA-seq and genome sequences are not available. The lines connect genes representing putative orthologs across the four species. The trichome/stem RNA-seq data of two biological S. pennellii replicates are summarized (Supplementary file 2) for genes in the syntenic region. The red-blue color gradient provides a visual marker to rank the expression levels in FPKM. Structures of representative medium chain acylsugars from S. quitoense (acylinositol, I4:26) (Hurney, 2018) and S. pennellii (acylglucose, G3:19) (Leong et al., 2019) are on the right. Figure 5—figure supplement 1 shows that stable Sp-AACS1 transformation of the M82 CRISPR mutant sl-aacs1 restores C12 containing acylsugars (B) CRISPR/Cas9-mediated gene knockout of Sp-AACS1 or Sp-AECH1 in S. pennellii produce no detectable medium chain containing acylsugars. The ESI+ mode LC/MS extracted ion chromatograms of trichome metabolites are shown for each mutant. The m/z 127.01 (left panel) corresponds to the glucopyranose ring fragment that both acylsucroses and acylglucoses generate under high collision energy positive-ion mode. The m/z 155.14 (center panel) and 183.17 (right panel) correspond to the acylium ions from acylsugars with chain length of C10 and C12, respectively. Figure 5—figure supplement 2A–C describes the design of the gRNAs and the detailed information of gene edits. (C) Silencing Sq-AACS1 (Sq-c34025) or Sq-AECH1 (Sq-c37194) in S. quitoense using VIGS leads to reduction of total acylsugars. The peak area/internal standard (IS) normalized by leaf dry weight was shown from sixteen plants ± SEM. ***p<0.001, Welch two-sample t test. Figure 5—figure supplement 2E and F describes the VIGS experimental design and the representative LC/MS extracted ion chromatograms of S. quitoense major acylsugars. (D) Reduced gene expression of Sq-AACS1 or Sq-AECH1 correlates with decreased acylsugar levels in S. quitoense. The qRT-PCR gene expression data are plotted with acylsugar levels of the same leaf as described in Figure 5—figure supplement 2E. Figure 5—source data 1 includes raw data for the S. quitoense VIGS experiments.
-
Figure 5—source data 1
Data used to make Figure 5C and D.
Silencing Sq-AACS1 or Sq-AECH1 in S. quitoense using VIGS leads to reduction of total acylsugars.
- https://cdn.elifesciences.org/articles/56717/elife-56717-fig5-data1-v2.xlsx
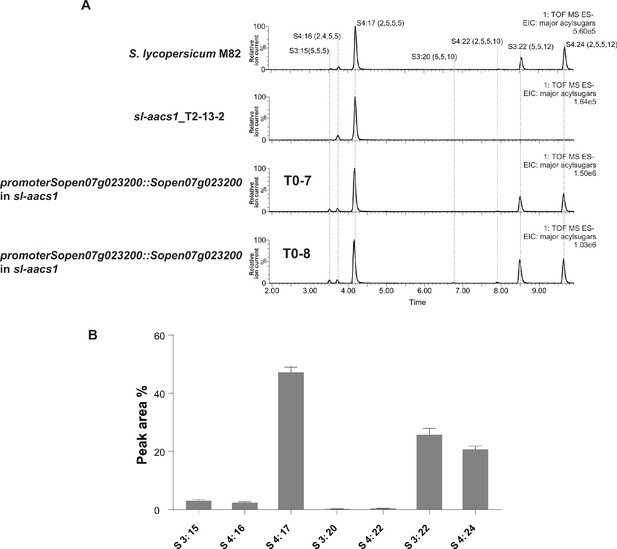
Stable Sp-AACS1 transformation of the M82 CRISPR mutant sl-aacs1 restores C12 containing acylsugars.
(A) ESI- mode, LC/MS extracted ion chromatograms are shown for seven major acylsugar peaks extracted from trichome of S. lycopersicum M82, sl-aacs1, and two independent T0 generation suppressed sl-aacs1 transgenic lines expressing Sp-AACS1 under its own promoter. (B) Peak area percentage of seven major trichome acylsugars of the suppression transgenic plants. The sum of the peak area percentage of each acylsugar equals to 100%. The results of acylsugar peak area percentage were calculated from six independent T0 transgenic lines ± SEM.
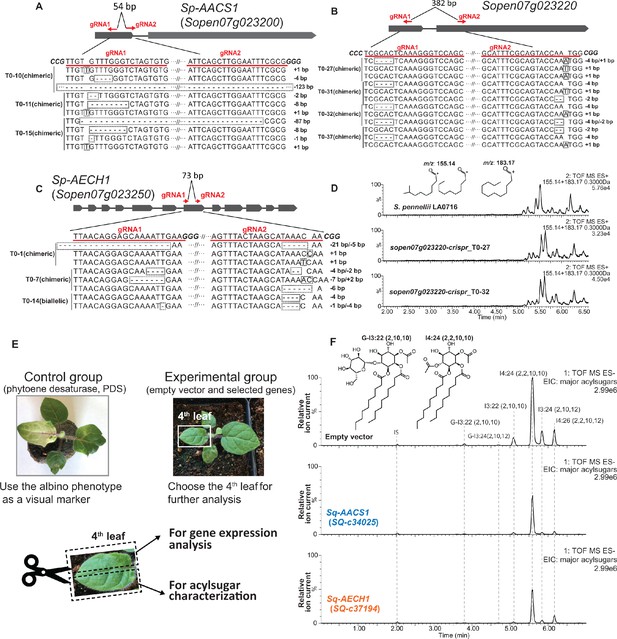
Functional analysis of AACS1 and AECH1 in S. pennellii and S. quitoense via CRISPR-Cas9 system and VIGS, respectively.
The design of gRNAs targeting wild tomato S. pennellii LA0716 Sp-AACS1 (A), Sopen07g023250 (B), and Sp-AECH1 (C) is highlighted with red lines and text. The transgenic T0 generation carrying chimeric or biallelic gene edits are shown beneath the gene model. DNA sequence of the gene edits was obtained through Sanger sequencing of cloned plant DNA fragments. The gene edits are highlighted with dotted rectangular boxes. (D) ESI+ mode, LC/MS extracted ion chromatograms shown for C10 (m/z: 155.14) and C12 (m/z: 183.17) fatty acid ions corresponding to medium chain trichome acylsugars extracted from the CRISPR mutants sopen07g023220 and the S. pennellii LA0716 parent. (E) Experimental design of VIGS in S. quitoense. A control group silencing the PDS genes was performed in parallel with the experimental groups. The onset of the albino phenotype of the control group was used as a visual marker to determine the harvest time and leaf selection in the experimental groups. The fourth true leaves were harvested and cut in half for gene expression analysis and acylsugar quantification, respectively. (F) ESI- mode, LC/MS extracted ion chromatograms of six major acylsugars of S. quitoense for the experimental group. The three LC/MS chromatograms show representative acylsugar profiles of the empty vector control plants and the VIGS plants targeting Sq-AACS1 and Sq-AECH1.
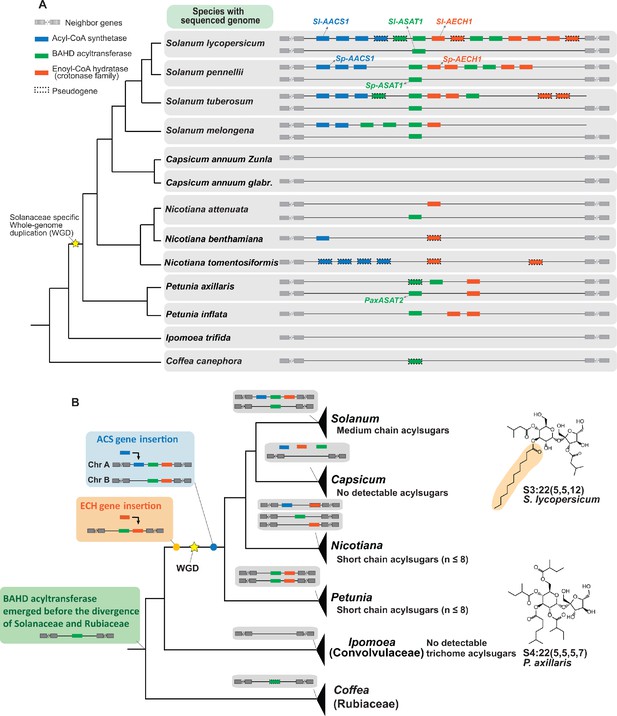
Evolution of the acylsugar gene cluster is associated with acylsugar acyl chain diversity across the Solanaceae family.
(A) The acylsugar gene cluster syntenic regions of 11 Solanaceae species and two outgroup species Ipomea trifida (Convolvulaceae) and Coffea canephora (Rubiaceae). This is a simplified version adapted from Figure 6—figure supplement 1. Only genes from the three families – ACS (blue), BAHD acyltransferase (green), and ECH (orange) – are shown. For information about the syntenic region size in each species refer to Figure 6—figure supplement 1 and Supplementary file 4. (B) The evolutionary history of the acylsugar gene cluster and its relation to the acylsugar phenotypic diversity. The evolution of BAHD acyltransferase genes is inferred based on Figure 6—figure supplement 2 and Figure 6—figure supplement 5. ECH genes based on Figure 6—figure supplement 3. ACS genes based on Figure 6—figure supplement 4 and Figure 6—figure supplement 6. The temporal order for the emergence for the three types of genes are shown in the colored boxes on the left: green box (BAHD acyltransferase), orange box (ECH), blue box (ACS). The yellow star represents the Solanaceae-specific WGD. Structures of representative short and medium chain acylsugars were shown on the right. Figure 6—figure supplement 8 describes distribution of acylsugar acyl chains with different lengths in species across the Solanaceae family.

Syntenic regions containing the acylsugar gene cluster.
The species name and chromosome/scaffold identifier are indicated with S. lycopersicum in blue font. Rectangle: protein-coding gene (solid line) or pseudogene (dotted line) colored according to the type of genes. Line connecting two genes: putative orthologous genes. Numbers underneath chromosomes: chromosome coordinates in million bases (Mb). The gene ID and location information used to generate the synteny figure is provided in Supplementary file 4.
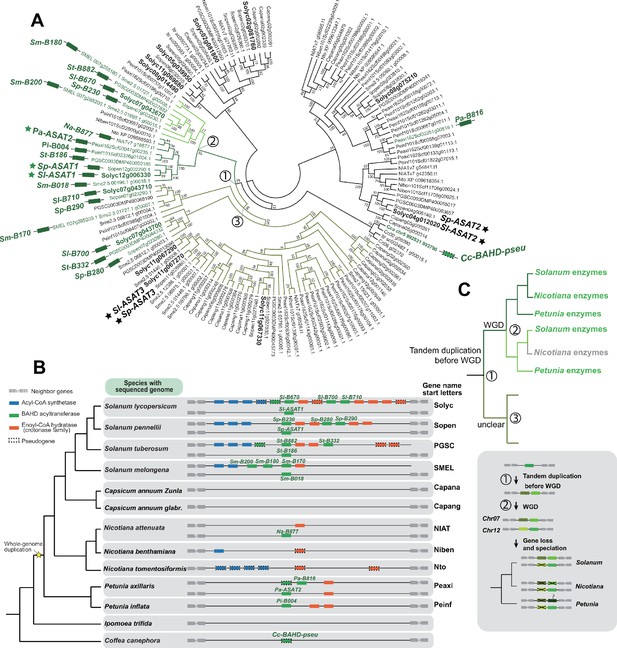
Analysis of the evolutionary history of the BAHD acyltransferases in the syntenic regions in different Solanaceae species.
(A) Phylogenetic tree of BAHD acyltransferases homologous to Solyc07g043670. Genes colored with green and labeled with green rectangles are from the syntenic regions of Solanaceae species shown in panel (B). Genes marked with stars have been biochemically tested involved in acylsugar biosynthesis in previous studies. (B) The acylsugar gene cluster syntenic regions of 11 Solanaceae species and two outgroup species Ipomea trifida (Convolvulaceae) and Coffea canephora (Rubiaceae). (C) Reconciled evolutionary history of BAHD acyltransferases based on panel (A) and (B). The colors of the branches correspond to different lineages shown in panel (A). Grey branch means enzymes from that lineage could not be found through BLAST and may have been lost. Numbers in the circle indicate the nodes in the phylogenetic tree as shown in panel A. The inferred evolutionary events were shown next to the nodes. Grey box highlighted the evolutionary history of the genes in the syntenic region. Before the Solanaceae specific whole genome duplication (WGD) events, the BAHD acyltransferase gene was tandemly duplicated. The WGD events resulted in at least two genomic regions (Chr07 and Chr12), each containing two BAHD acyltransferase genes. Before the divergence of Solanum, Nicotiana, and Petunia species, one of the tandem copies in Chr12 region was lost, and only orthologs of Sl-ASAT1 was retained. The question mark next to Petunia denotes the inconsistence of the phylogenetic relationship and the chromosome location of the gene Pa-B816 with an unknown mechanism.
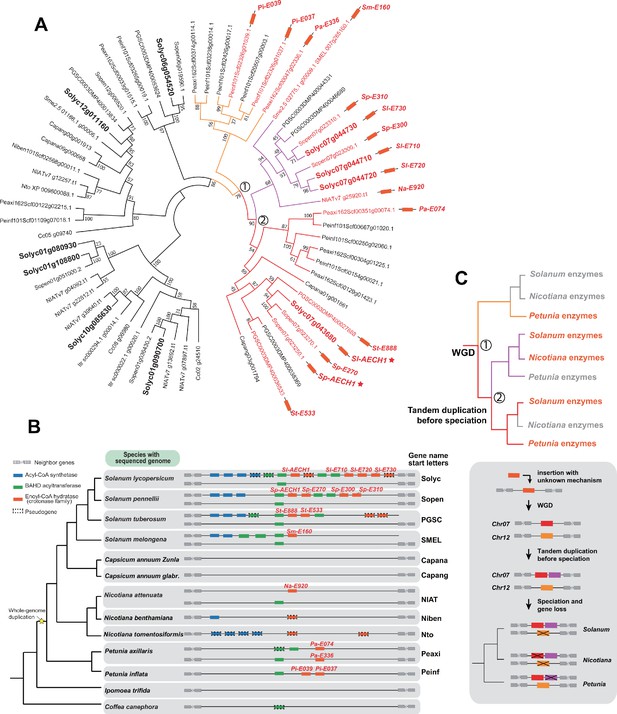
Analysis of the evolutionary history of the ECH genes in the syntenic regions in different Solanaceae species.
(A) Phylogenetic tree of ECH genes homologous to Sl-AECH1. Genes colored with orange and labeled with orange rectangles are from the syntenic regions of Solanaceae species shown in panel (B). Genes marked with stars have been tested involved in acylsugar biosynthesis. (B) The acylsugar gene cluster syntenic regions of 11 Solanaceae species and two outgroup species Ipomea trifida (Convolvulaceae) and Coffea canephora (Rubiaceae). (C) Reconciled evolutionary history of ECH enzyme based on panel (A) and (B). The colors of the branches correspond to different lineages shown in panel A. Grey branch means enzymes from that lineage could not be found through BLAST and may have been lost. Numbers in the circle indicate the nodes in the phylogenetic tree as shown in panel (A). The inferred evolutionary events were shown next to the nodes. Grey box highlighted the evolutionary history of the genes in the syntenic regions. Before the Solanaceae specific WGD events, an ECH was inserted into the syntenic region through unknown mechanism. After the WGD events, there was one ECH gene in each syntenic region on Chr07 and Chr12. Before the divergence of Solanum, Nicotiana, and Petunia species, the ECH gene on Chr07 had experienced a tandem duplication event, leading to two branches on the phylogenetic tree. During the speciation, the ECH gene on Chr12 was deleted from the genome in the most recent common ancestor of Nicotiana and Solanum after divergent from Petunia, while one of the tandem duplicates on Chr07 was lost in Petunia.
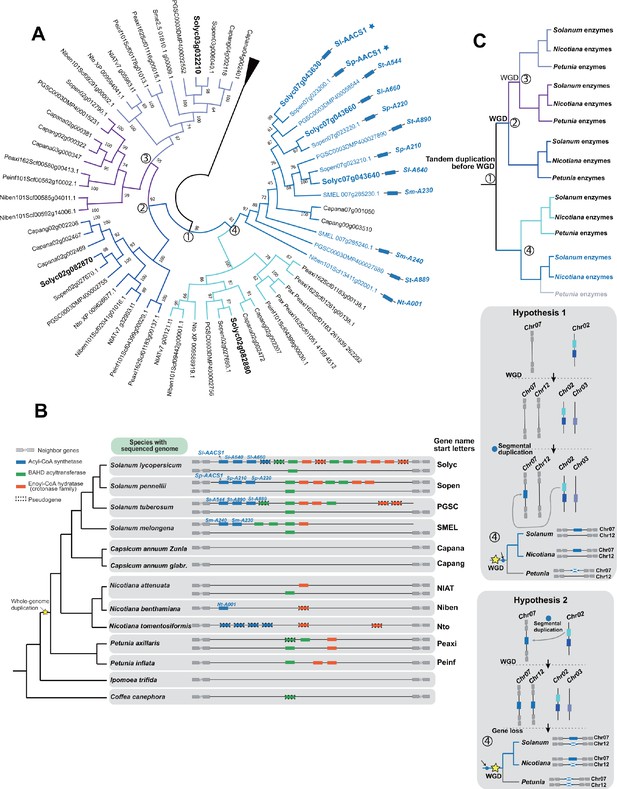
Analysis of the evolutionary history of the ACS genes in the syntenic regions in different Solanaceae species.
(A) Phylogenetic tree of ACS genes homologous to Sl-AACS1. Genes colored with blue and labeled with blue rectangles are from the syntenic regions of Solanaceae species shown in panel (B). Genes marked with stars have been tested involved in acylsugar biosynthesis. (B) The acylsugar gene cluster syntenic regions of 11 Solanaceae species and two outgroup species Ipomea trifida (Convolvulaceae) and Coffea canephora (Rubiaceae). (C) Reconciled evolutionary history of ACS enzyme based on panel (A) and (B). The colors of the branches correspond to different lineages shown in panel (A). Grey branch means enzymes from that lineage could not be found through BLAST and may have been lost. Numbers in the circle indicate the nodes in the phylogenetic tree as shown in panel A. The inferred evolutionary events were shown next to the nodes. A tandem duplication event happened before the Solanaceae specific WGD events, leading to two adjacent ACS genes on Chr02 (Solyc02g082880 and Solyc02g082870), which were placed on two independent lineages in the phylogenetic tree. Solyc02g082870 had gone through two rounds of WGD events, supported by the observation that Solyc02g082870 and Solyc03g032210 are located in corresponding syntenic blocks. Solyc02g082880 may have experienced the segmental duplication, resulting in the ACS gene on Chr07, which had experienced another two rounds of tandem duplication in the common ancestor of Solanum species (Sl-AACS1, Solyc07g043660, and Solyc07g043640). However, whether the segmental duplication event happened before or after the Solanaceae specific WGD events cannot be well resolved by the phylogenetic analysis. Two hypotheses were proposed as shown in the grey boxes. If the insertion happened before WGD, two independent gene loss events on chromosomes 7 and 12 should have happened in Petunia (Hypothesis 2). If the insertion happened after WGD, only one gene loss in Petunia was supposed to have happened (Hypothesis 1). Note that node 4 in (A) leads to two clades, one without any Petunia ACS homolog (darker blue) and the other with Petunia homologs (cyan). With regard to the timing of the duplication event leading to these two clades, it was likely before the split between the Petunia and the tomato/tobacco lineages where one Petunia loss event occurred (darker blue). If it was after the split, the presence of a Petunia gene would need to be explained by a gene gain through horizontal gene transfer or other means (cyan) - a far less likely scenario than a gene loss.
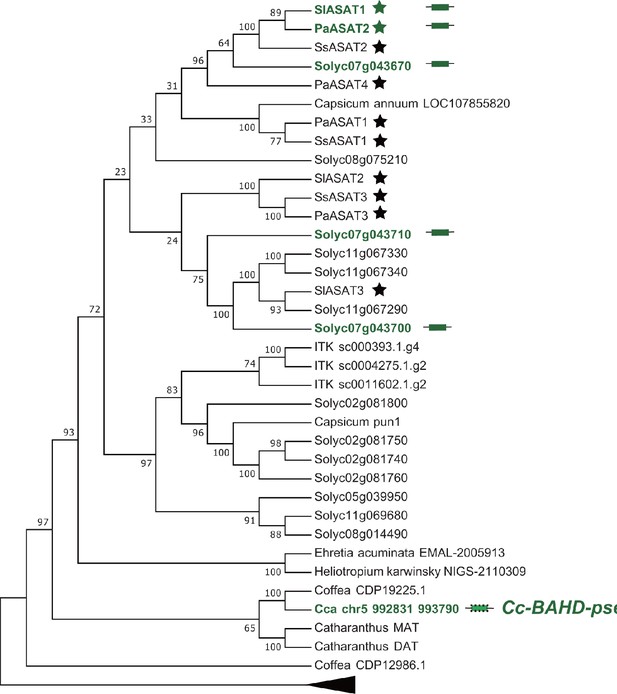
Phylogenetic analysis of the BAHD acyltransferase.
The BAHD acyltransferase pseudogene (Cc-BAHD-pseu) in the corresponding syntenic region (Figure 6) of Coffea canephora is one of the closest Coffea sequences sister to the ASAT clade. It indicates that the BAHD acyltransferase gene was the first to harbor in this syntenic region before the divergence between Solanaceae and Rubiaceae. The translated amino acid sequence of Cc-BAHD-pseu was aligned with sequences used in Figure 6A of a previous study (Moghe et al., 2017) using MUSCLE. The phylogenetic trees were built using the maximum likelihood method with 1000 bootstrap replicates. The tree was generated using RAxML/8.0.6 with the following parameters: -f a -x 12345 p 12345 -# 1000 m PROTGAMMAAUTO --auto-prot=bic, and was shown with the midpoint rooting. Genes colored with green are from the focused syntenic regions. Genes marked with stars have been biochemically tested involved in acylsugar biosynthesis in previous studies.
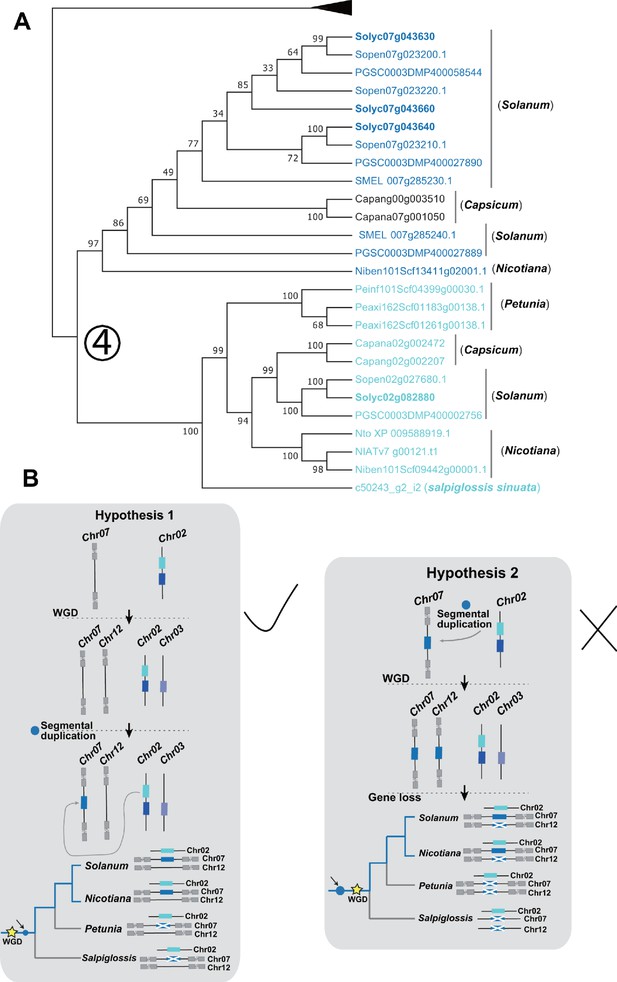
Additional evolutionary analysis of ACS genes in the syntenic regions to understand when the segmental duplication event happened.
(A) Sl-AACS1 homologs obtained from Salpiglossis sinuate trichome transcriptome dataset (Moghe et al., 2017) were added for additional phylogenetic analysis. Genes colored with blue are from the syntenic regions of Solanaceae species Only the lineage derived by the number (4) evolutionary event as depicted in Figure 6—figure supplement 5 was shown. (B) If the insertion happened before WGD, one gene loss on Solanum chromosome 12, as well as two independent gene losses on chromosomes 7 and 12 should have happened in Petunia and in Salpiglossis sinuate (Hypothesis 2). However, if the insertion happened after WGD, then only one gene loss event in Petunia and Salpiglossis was supposed to have happened (Hypothesis 1). The latter scenario is more likely due to the principle of parsimony.
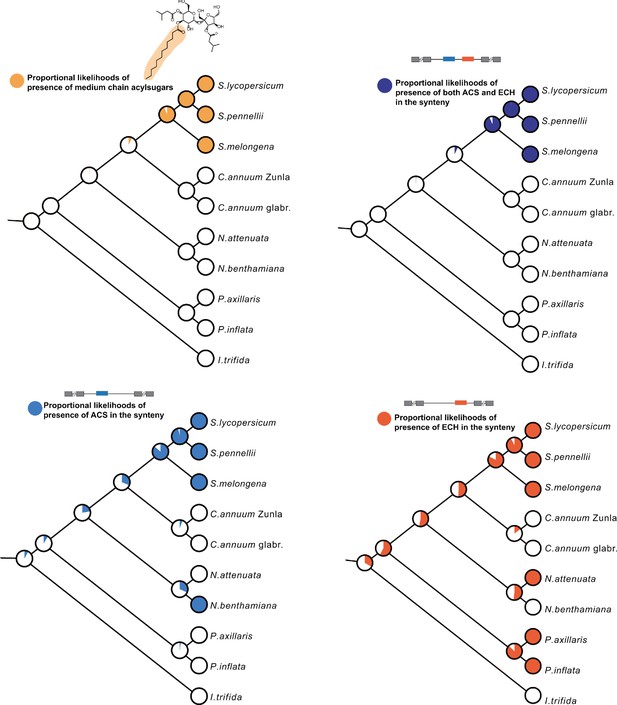
Ancestral trait state reconstruction analysis.
Four traits were inferred for their ancestral states using the maximum likelihood model Mk1 in Mesquite 3.6. They are the presence of medium chain acylsugars, presence of ACS in the synteny, presence of ECH in the synteny, and presence of both ACS and ECH in the synteny. The proportional likelihoods were shown in each diverging node in the ball charts.
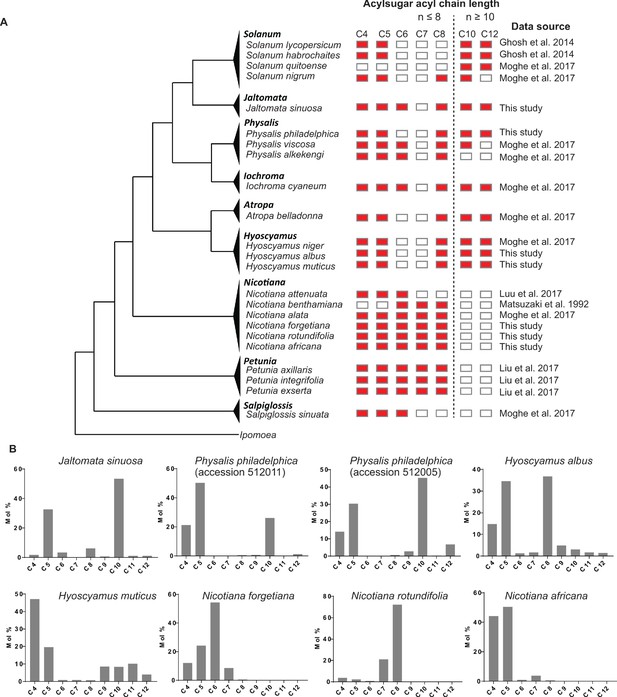
Phylogenetic distribution of acylsugar acyl chains with different lengths across the Solanaceae family.
(A) The collated results of acylsugar acyl chain distribution across different Solanaceae species. Red rectangles indicate detectable acyl chains in the acylsugars produced in the tested species and white rectangles indicate no detectable signals. The data source where the results are derived is listed on the right. (B) Results of acylsugar acyl chain characterization of selected Solanaceae species. Mole percentage (Mol %) of acylsugar acyl chains with different lengths were obtained from GC/MS analysis of fatty acid ethyl esters.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Gene (Solanum lycopersicum M82) | Sl-AACS1 | This paper | GeneBank: MT078737 | Characterized and named in the results |
| Gene (Solanum lycopersicum M82) | Sl-AECH1 | This paper | GeneBank: MT078736 | Characterized and named in the results |
| Gene (Solanum pennellii LA0716) | Sp-AACS1 | This paper | GeneBank: MT078735 | Characterized and named in the results |
| Gene (Solanum pennellii LA0716) | Sp-AECH1 | This paper | GeneBank: MT078734 | Characterized and named in the results |
| Gene (Solanum quitoense) | Sq-AACS1 | This paper | GeneBank: MT078732 | Characterized and named in the results |
| Gene (Solanum quitoense) | Sq-AECH1 | This paper | GeneBank: MT078731 | Characterized and named in the results |
| Gene (Solanum quitoense) | Sq_c35719 | This paper | GeneBank: MT078733 | Characterized and named in the results |
| Software, algorithm | Trimmomatic | http://www.usadellab.org/cms/index.php?page=trimmomatic | RRID:SCR_011848 | |
| Software, algorithm | TopHat | http://ccb.jhu.edu/software/tophat/index.shtml | RRID:SCR_013035 | |
| Software, algorithm | Cufflinks | http://cole-trapnell-lab.github.io/cufflinks/cuffmerge/ | RRID:SCR_014597 | |
| Software, algorithm | MCScanX-transposed | http://chibba.pgml.uga.edu/mcscan2/transposed/ | ||
| Software, algorithm | RAxML | https://github.com/stamatak/standard-RAxML | RRID:SCR_006086 | |
| Software, algorithm | Mesquite | https://www.mesquiteproject.org/ | RRID:SCR_017994 |
Additional files
-
Supplementary file 1
Co-expression analysis of tomato genes from ACS, ECH, and BAHD acyltransferase families used for phylogenetic analysis in this study.
The values of Pearson’s correlation coefficient of the expression profiles between any of the two genes were shown in the table. The coefficient values were generated using the FPKM values of these genes in the 372 RNA-seq samples as shown in Figure 2—figure supplement 1. The orange box highlights a group of co-expressed genes involved in acylsugar biosynthesis, such as Sl-ASATs, Sl-AACS1, and Sl-AECH1. The purple box points out another group of co-expressed genes that are root hair specific.
- https://cdn.elifesciences.org/articles/56717/elife-56717-supp1-v2.xlsx
-
Supplementary file 2
Gene expression levels of all analyzed transcripts in Solanum pennellii LA0716.
logFC: log2 fold change in stem trichomes versus shaved stems. logCPM: log (counts per million) in trichomes versus shaved stems. The F and Q-value test the significance of differential expression via a quasi- general linear model. The values noted in the sample columns represent the FPKM (Fragments Per Kilobase of transcript per Million mapped reads) analyzed via Cufflinks.
- https://cdn.elifesciences.org/articles/56717/elife-56717-supp2-v2.xlsx
-
Supplementary file 3
Synthesized gene fragments and primers used in this study.
- https://cdn.elifesciences.org/articles/56717/elife-56717-supp3-v2.xlsx
-
Supplementary file 4
The date used to generate the synteny figure shown in Figure 6—figure supplement 1.
- https://cdn.elifesciences.org/articles/56717/elife-56717-supp4-v2.zip
-
Supplementary file 5
The sequence alignment documents used to generate the phylogenetic trees for Figure 6—figure supplements 2, 3, 4, 5 and 6.
- https://cdn.elifesciences.org/articles/56717/elife-56717-supp5-v2.zip
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/56717/elife-56717-transrepform-v2.docx





