Rdh54/Tid1 inhibits Rad51-Rad54-mediated D-loop formation and limits D-loop length
Figures
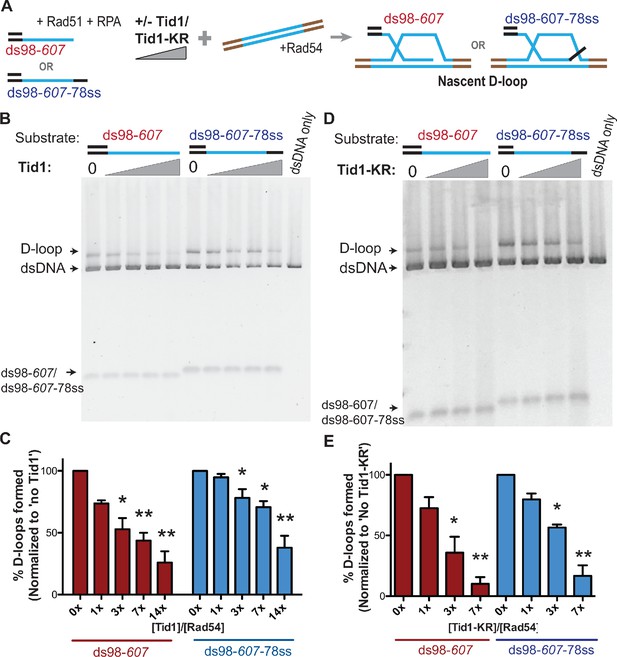
Tid1 inhibits D-loop formation in vitro in a concentration-dependent and an ATPase-independent manner.
(A) Reaction scheme for in vitro D-loop formation assays in presence of Tid1 or the ATPase-defective Tid1-K318R. Here and in all subsequent figures, unless otherwise stated, incubations were in the following order: ssDNA and Rad51 for 10 min, then RPA for 5 min, followed by Tid1/Tid1-KR for 5 min, finally a linearized dsDNA and Rad54 for 15 min (for details, see Materials and methods). Homology between the ssDNA and dsDNA is indicated in blue. (B, D) D-loop reactions performed as described in (A) with increasing concentrations of Tid1 or Tid1-KR, respectively. The gels were stained with SYBR gold. (C, E) Quantitation of the D-loops from the gels in (B) and (D) as the percentage of donor invaded by ssDNA, respectively. The D-loops are normalized to the amount formed in absence of Tid1/Tid1-KR for each paired reaction. Error bars indicate mean ± SD (n = 3). * indicates p-value<0.05, **<0.005, with a two-tailed t-test, in comparison to ‘No Tid1/Tid1-KR’ sample. Refer to Figure 1—figure supplement 1 for absolute D-loop values.
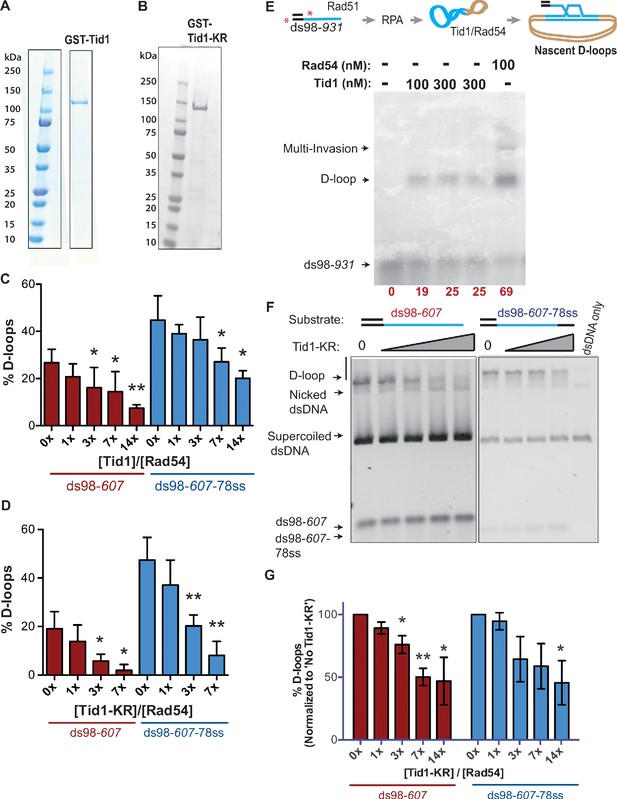
Tid1 by itself stimulates Rad51-mediated D-loops, while inhibits Rad54-mediated D-loops.
(A, B) Denville-Blue-stained SDS-PAGE gel showing purified GST-Tid1 and GST-Tid1-KR, respectively. (C, D) Absolute quantitation of D-loops from the gels in Figure 1B and D, without any normalization. Quantitation carried out as percentage of dsDNA donor invaded to form the D-loops. Error bars indicate mean ± SD (n = 3). (E) D-loop assay showing that Tid1 is able to enhance Rad51 DNA strand invasion activity, although less efficiently than Rad54. The D-loop assay was performed as described in the reaction schematic (for details, see Materials and methods). Incubations were in the following order: End-labeled ds98-931 substrate (end-labeling depicted as red stars) and Rad51 for 10 min, then RPA for 10 min, and then supercoiled dsDNA with Tid1 or Rad54 for 10 min. Quantification of D-loops as the percentage of radiolabeled ssDNA within a D-loop is shown in red at the bottom of the gel image. (F) SYBR-gold-stained gel of the D-loop reactions performed as per the reaction scheme in Figure 1A with increasing Tid1-KR titration, except in presence of supercoiled dsDNA instead of linear dsDNA. (G) Quantification of the D-loops from (F), normalized to the D-loops formed in absence of Tid1-KR for each paired reaction. Error bars indicate mean ± SD (n = 3). For all bar graphs (C, D, G), * indicates p-value<0.05, **<0.005 with a two-tailed t-test, in comparison to ‘No Tid1/Tid1-KR’ sample.
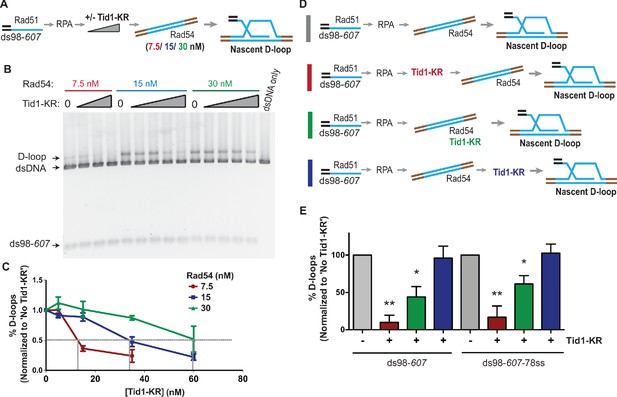
Tid1 competes with Rad54 activity and does not inhibit D-loops after they are formed.
(A) Reaction scheme depicting various Rad54 concentrations, along with Tid1-KR titration in an in vitro D-loop assay. (B) SYBR gold stain of gel showing D-loop reaction performed with varying Rad54 and Tid1-KR titrations as indicated in (A). (C) Quantitation of the D-loops from the gel with normalization to D-loops formed in absence of Tid1-K318R for each of the Rad54 concentrations Error bars indicate mean ± SD (n = 3). Gray lines are drawn to indicate 50% inhibition. (D) Different reaction schemes based on the timing of addition of Tid1-KR is indicated by different colored bars on the left. Gray bar indicates D-loop reaction performed in absence of Tid1-KR. Red bar indicates Tid1-KR added to the reaction 5 min before adding dsDNA and Rad54. Green bar indicates Tid1-KR added at the same time as dsDNA and Rad54, whereas blue bar indicates Tid1-KR added 10 min after dsDNA and Rad54. All these reactions were performed using both ds98-607 or ds98-607-78ss substrates. (E) Quantitation of D-loops formed as in (D) with normalization to the D-loop levels formed in absence of Tid1-KR for each paired reaction. The color of the bars in the graph correspond to the colored bars in (D) and represent the respective D-loop samples. Error bars indicate mean ± SD (n = 3). * indicates p-value<0.05, **<0.005, with a two-tailed t-test, in comparison to ‘No Tid1/Tid1-KR’ sample. Refer to Figure 2—figure supplement 1B for unnormalized D-loop values.
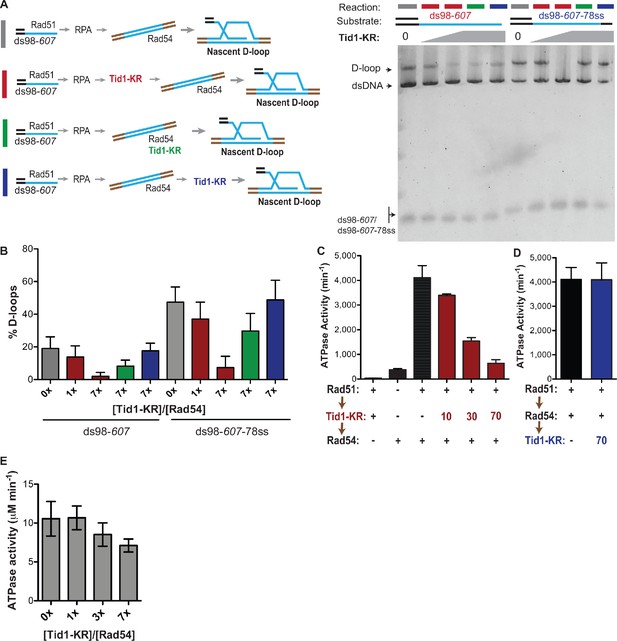
Tid1 competes with Rad54 to inhibit D-loops before they are formed and inhibits Rad54’s ATPase activity.
(A) On the left is the reaction schemes for different timings of Tid1-KR addition with respect to dsDNA and Rad54, each denoted by a colored bar (same as in Figure 2D). On the right is a SYBR-gold-stained gel of the D-loop reactions. The D-loop reactions were performed with either the ds98-607 or the ds98-607-78ss substrate. The concentration of Tid1-KR is as indicated. (B) Quantitation of D-loops from the gel in (A). (C) ATPase assay measuring rate of ATP hydrolysis with Tid1-KR added prior to Rad54. The order of additions followed the brown arrows on the left with 5 min incubations between each addition. All reactions were in presence of 3 kb dsDNA (6 µM nt) added before Rad51. ‘+’ indicates that 10 nM Rad54 or Tid1-KR was used, unless otherwise indicated (in nM) by a numeric value (for details, see Materials and methods). Mean ± SD (n = 2 or 3). (D) ATPase activity measured with Rad54 added prior to Tid1-KR. Again, the order of additions is as per the black arrows on the left, with 5 min incubations between each addition. All reactions were in presence of 3 kb dsDNA (6 µM nt) added before Rad51. ‘+’ indicates that 10 nM Rad54 or Tid1-KR was used, unless otherwise indicated (in nM) by a numeric value. Mean ± SD (n = 3). (E) ATPase activity of Rad54 measured with Tid1-KR titration on dsDNA, in absence of Rad51. 10 nM Rad54 was added to 3 kb dsDNA (6 µM nt) reaction containing the specified relative concentration of Tid1-KR (for details, see Materials and methods). Mean ± SD (n = 3).
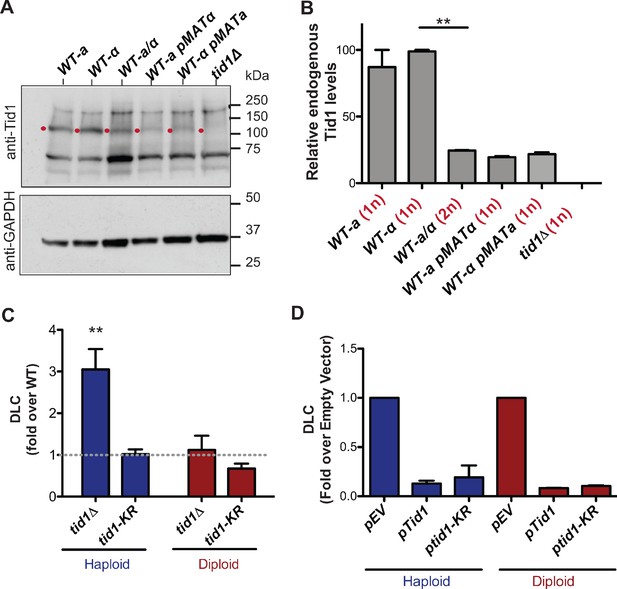
Tid1 affects D-loops in vivo in a concentration dependent manner.
(A) Western blots showing staining with anti-Tid1 or anti-GAPDH antibodies. The band depicting endogenous Tid1 protein (107.9 kDa mol. wt.) is indicated by a red dot. The position of Tid1 was validated by comparison with purified Tid1 (not shown). The other bands are non-specific bands from antibody staining. WT represents wild-type yeast, along with its mating type status. WT-a pMATα indicates MATa haploid yeast transformed with MATα-expressing plasmid. (B) Quantitation of endogenous Tid1 levels normalized to the Tid1 levels in haploid WT-α strain. It shows MAT-heterozygosity dependent suppression of Tid1 expression. Error bars indicate mean ± SD (n = 3). ** p-value<0.005 with two-tailed t-test. (C) The D-loop Capture (DLC) signal obtained from the DLC assay is normalized to WT DLC signal (indicated by gray dotted line) for both haploid and diploid yeast. The DLC signal was measured 2 hr post DSB induction. Error bars indicate mean ± SD (n = 5). The haploid data is adopted from Piazza et al., 2019. (D) DLC signal obtained from the DLC assay performed in haploid and diploid yeast transformed with plasmids. pEV, pTid1 and pTid1-KR indicates yeast transformed with an empty vector, a vector containing GST-Tid1 or GST-Tid1-KR under a galactose promoter respectively. The DLC signal was normalized to the sample containing empty vector. The DLC signal was measured 2 hr post DSB induction with galactose. Mean ± SD (n = 2).
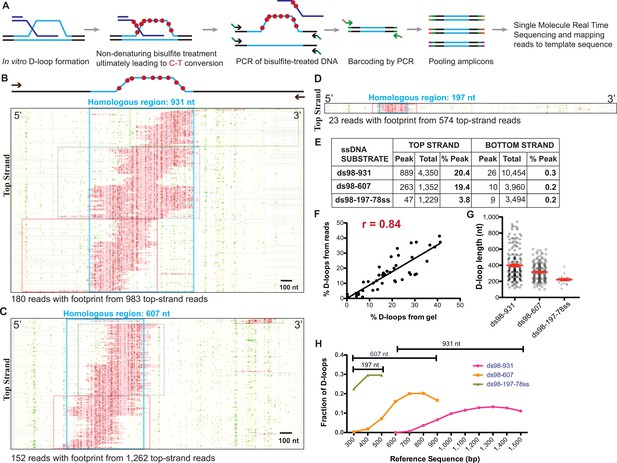
Single-molecule D-loop Mapping Assay to map D-loop length, position and distribution in vitro.
(A) Schematic of the D-loop Mapping Assay using D-loops formed on linear donors in vitro as described in Figure 1A as the starting input (for details, see Materials and methods; Shah et al., 2020). (B) Footprint map depicting reads with a D-loop footprint. Reads are derived from an in vitro Rad51-Rad54-mediated D-loop sample containing ds98-931 substrate and a linear donor. Only reads from the top strand of dsDNA donor that contain a footprint are shown here. Here and in all subsequent figures with a footprint map, each horizontal line represents one read molecule (or amplicon). The position of each cytosine across the read sequence is indicated by yellow lines. The status of each cytosine along the sequence is color-coded with green representing C-T conversions. The status of cytosine is changed to red if the C-T conversions cross the peak threshold and are thus defined as D-loop footprints. Unless otherwise mentioned the peak threshold is t40w50 (40% cytosines converted to thymine in a stretch of 50 consecutive cytosines). The reads are clustered based on the position of footprints in 5′ to 3′ direction. The faintly colored boxes indicate the clusters. The blue box represents dsDNA region homologous to 931 nt on the invading substrate. Scale bar is 100 nt. For bottom strand reads, refer to Figure 4—figure supplement 1A. (C) Footprint map depicting reads containing D-loop footprints from an in vitro Rad51- and Rad54-mediated D-loop reaction performed with ds98-607 substrate and a linear donor. (D) Footprint map depicting reads containing D-loop footprints from an in vitro Rad51- and Rad54-mediated D-loop reaction performed with ds98-197-78ss substrate and a linear donor. Note that a smaller fraction of reads with footprints was observed for the ds98-197-78ss substrate due to its lower D-loop formation efficiency (Wright and Heyer, 2014). (E) Table summarizing the total number of reads containing a footprint as ‘peak’ and the total number of reads analyzed as ‘total’ for each strand. ‘% D-loops/% Peak’ indicate the percentage of reads containing a footprint. The data represents a cumulation from >3 independent replicates. The percentage of D-loops are calculated by dividing the number of reads with footprint by the total number of reads for that strand. Shown are cumulative data from three to five independent replicates, of which one to two overlap with the data reported in the accompanying manuscript (Shah et al., 2020). (F) Dot plot indicating correlation between the percentage of reads containing D-loop footprint by DMA and the percentage of D-loops seen on the gel relative to the uninvaded dsDNA. Pearson coefficient’s r = 0.84 for 42 XY pairs. p-value<0.0001. (G) Dot plot indicating individual D-loop lengths measured across substrates with varying homology lengths. D-loop length was measured in nt based on the footprint size called by the DMA assay. The data represents a cumulation from >3 independent replicates. Red error bars indicate mean ± SEM. (H) Distribution of D-loop footprints across the region of homology (as indicated by a capped line for each substrate type), with an enrichment at the 3′-end. The distribution is measured by binning each footprint in 100 nt bins across the homology non-exclusively and depicted as the percentage of total D-loops within each bin. ‘Reference sequence’ indicates the position on dsDNA donor.
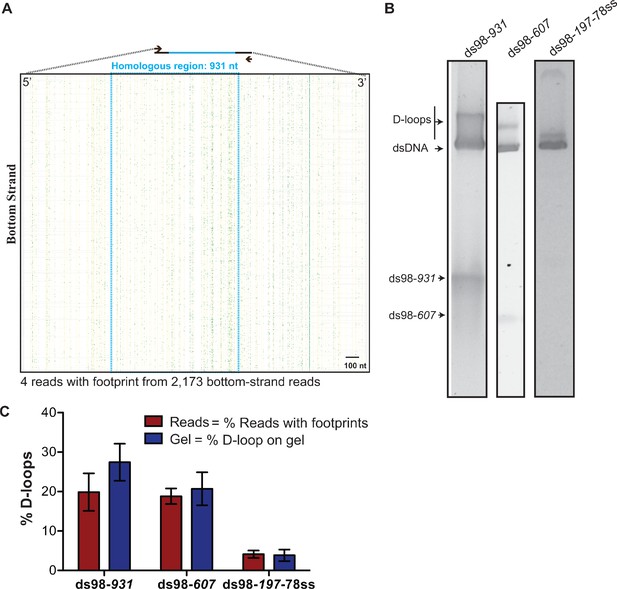
D-loops formed with various length substrates.
(A) Footprint map of reads from the bottom strand of dsDNA donor from the ds98-931 D-loop reaction. Lack of red lines indicates lack of D-loop footprints on the bottom strand. Since <5 reads had a footprint, a footprint map for footprint containing reads could not be created. The blue dotted box indicates region of homology with the substrate. Each horizontal line represents a read in 5′–3′ direction. The yellow vertical lines indicate position of cytosine in original sequence of the bottom strand. In each read, the yellow mark of cytosine is changed to green when a C-T conversion is observed. These green marks indicate cytosine modification from breathing of DNA. (B) Agarose gel depicting one-third volume of the D-loop reaction used for the D-loop Mapping Assay. (C) Quantitation of D-loops from the gel in (B) being compared to the percentage of reads with D-loop footprint as observed by DMA. The percentage of D-loops from DMA is comparable to the quantitation of D-loops from the gel for each substrate type. Mean ± SD (n = 3).
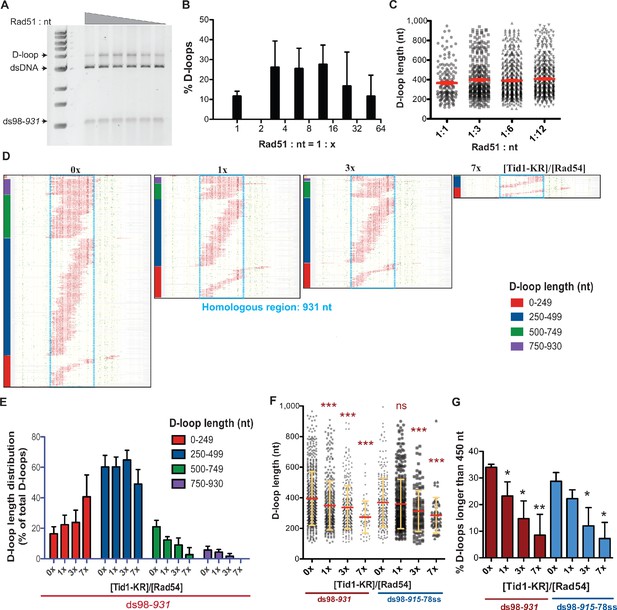
Tid1-KR restricts formation of longer D-loops.
(A) Gel stained with SYBR gold depicting D-loops formed with decreasing concentrations of Rad51. (B) Quantitation of D-loops from the gel in (A), where 1 Rad51 to 3 nt (Rad51: nt = 1:3) is a saturating Rad51 concentration. Mean ± SD (n = 3). (C) Dot plot showing the D-loop lengths of D-loop footprints seen with varying Rad51 concentrations in DMA assay. In red is Mean ± SEM (n = 3). Refer to Figure 5—figure supplement 1 for the corresponding footprint maps. (D) Footprint map of reads containing D-loop footprints in DMA assay. The D-loops were formed from a ds98-931 substrate and Rad51: nt = 1:12 concentration, followed with Tid1-KR titration. The reads were clustered based on D-loop length and position. The colored bars on the left indicate four different length clusters (0–249, 250–499, 500–749 and 750–930 nt). The footprints for each sample depict a cumulation from four independent replicates. (E) Quantitation of the distribution of D-loop lengths within each length cluster as a percentage of total D-loops for each sample in (D). The color of the bars correlates with the cluster bars on the footprint maps in (D). Mean ± SD (n = 4). (F) Dot plot depicting the D-loop lengths observed by DMA, from both ds98-931 and ds98-915-78ss substrates. In red is mean ± SEM, in yellow is mean ± SD (n = 4). *** indicates p-value<0.0005, with a two-tailed paired t-test, in comparison to ‘No Tid1-KR’ sample. (G) Percentage of D-loop footprints that are longer than 450 nt from samples in (F). Mean ± SD (n = 4). * indicates p-value<0.05, **<0.005, with a two-tailed paired t-test, in comparison to ‘No Tid1-KR’ sample. Refer to Figure 5—figure supplement 2 for the distribution of D-loops formed from ds98-931 and ds98-915-78ss substrates. Refer to Figure 5—figure supplement 3 for D-loop footprints observed in presence of WT Tid1.
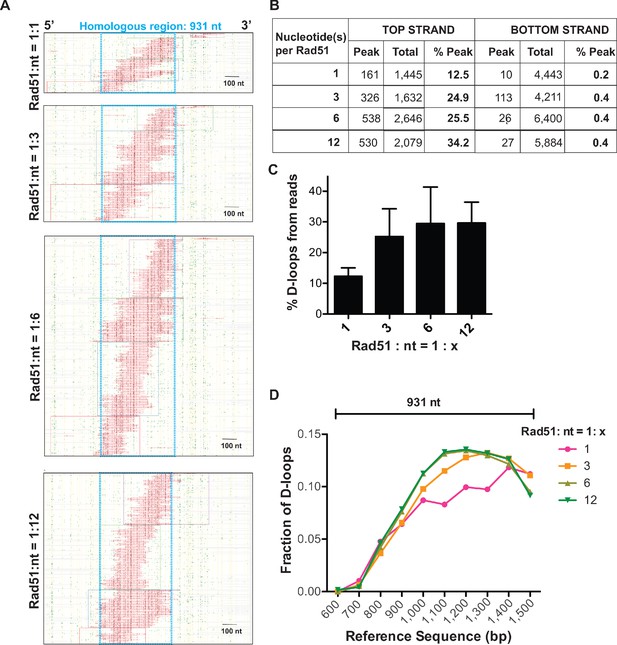
Changes in Rad51 concentration does not significantly alter D-loop characteristics in the D-loop Mapping Assay.
(A) Footprint maps of D-loop footprints from the top strand reads for D-loops formed in presence of varying Rad51 concentrations. To form the D-loops, ds98-931 substrate and a linear donor were used. The footprint map for each sample depict a cumulation from three independent replicates. (B) Table summarizing the total number of reads containing a footprint as ‘peak’ and the total number of reads analyzed as ‘total’ for each strand. ‘% D-loops/% Peak’ indicate the percentage of reads containing a footprint. The data represents a cumulation from three independent replicates. (C) Percentage of top strand reads containing D-loop footprints (percentage of D-loops from DMA assay). Mean ± SD (n = 3). (D) Distribution of D-loop footprints across the 5′−3′ region of homology (indicated by a capped line). The distribution is measured by binning each footprint in 100 nt bins across the homology non-exclusively and depicted as the percentage of total D-loops within each bin. ‘Reference sequence’ indicates the position on dsDNA donor. The ‘Rad51: nt = 1: 1’ sample shows variations due to an under-sampling effect.
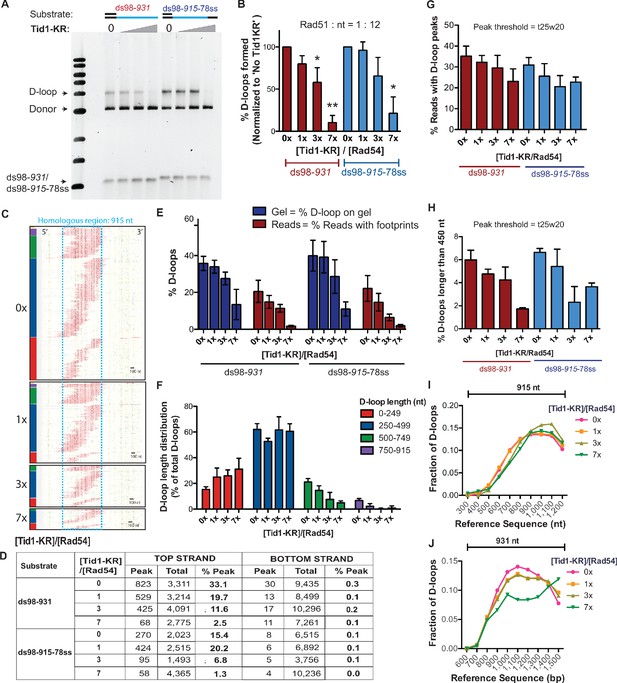
Tid1-KR limits D-loop length but does not alter the distribution of D-loop position.
(A) SYBR-gold-stained gel depicting D-loops formed in presence of Rad51: nt = 1:12, Tid1-KR titration and linear dsDNA. The same D-loop samples were used for the D-loop Mapping Assay (in Figure 5C, D). (B) Quantitation of D-loops from the gel in (A), normalized to the ‘No Tid1-KR’ sample for each paired reaction. Mean ± SD (n = 3). * indicates p-value<0.05, **<0.005 with a two-tailed t-test, in comparison to ‘No Tid1-KR’ sample. (C) Footprint maps of reads containing D-loop footprints from the D-loop reaction performed using ds98-915-78ss substrate as in (A). The reads were clustered based on D-loop length and position. The colored bars on the left indicate four different length clusters (red 0–249, blue 250–499, green 500–749 and purple 750–930 nt). The footprints for each sample depict a cumulation from four independent replicates. (D) Table summarizing the total number of reads containing a footprint as ‘peak’ and the total number of reads analyzed as ‘total’ for each strand. ‘% D-loops/% Peak’ indicate the percentage of reads containing a footprint. The data represents a cumulation from >3 independent replicates. (E) Quantitation of D-loops from the gel in (A) as compared to the percentage of top strand reads containing D-loop footprints (% D-loops from DMA) for each substrate. Mean ± SEM (n > 3). (F) Quantitation of the distribution of D-loop lengths within each length cluster as a percentage of total D-loops for each sample in (C). The color of the bars correlates with the cluster bars on the footprint maps in (C). Mean ± SD (n = 4). (G) Quantitation of D-loop levels from the DMA, where a footprint was defined using a t25w20 peak threshold instead of t40w50. (H) Percentage of D-loop footprints that are longer than 450 nt from samples in (G), where a peak threshold of t25w20 was used. (I, J) Distribution of the position of D-loop footprints across the region of homology (indicated by a capped line) for D-loops from ds98-915-78ss and ds98-931 substrates, respectively. The distribution was measured by binning each footprint in 100 nt bins across the homology non-exclusively and depicted as the fraction of total D-loops within each bin. ‘Reference sequence’ indicates the position on dsDNA donor. The distribution depicts an enrichment of D-loops at the 3′-end of homology. In (J), the ‘7x’ sample shows large variations due to an under-sampling effect.
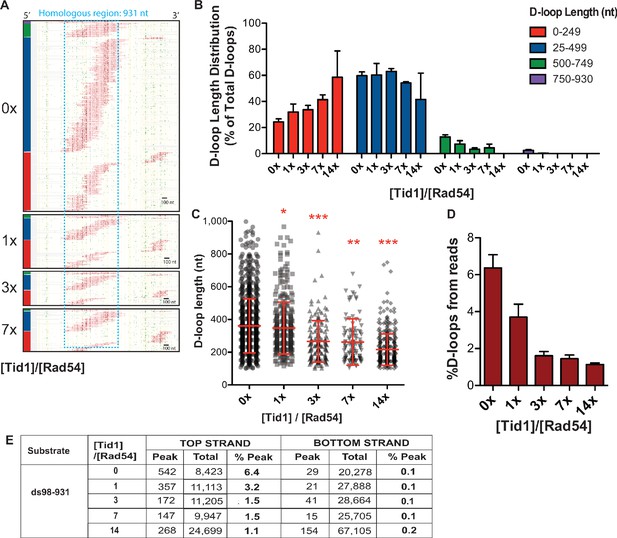
Tid1 limits D-loop length.
(A) Footprint map of D-loop footprints from the top strand reads of D-loop samples formed in presence of increasing concentration of Tid1 and ds98-931 substrate. The reads were clustered based on D-loop length and position. The colored bars on the left indicate four different length (red 0–249, blue 250–499, green 500–749 and purple 750–930 nt). The footprints for each sample depict a cumulation from two or three independent replicates. (B) Quantitation of the distribution of D-loop lengths within each length cluster as a percentage of total D-loops for each sample in (C). The color of the bars correlates with the cluster bars on the footprint maps in (A). Mean ± SD (n = 2 or 3). (C) Dot plot showing a distribution of D-loop lengths seen with a Tid1 titration in the DMA assay. In red is Mean ± SD (n = 2 or 3). * indicates p-value<0.05, **<0.005, ***<0.0005 with a two-tailed paired t-test, in comparison to ‘No Tid1’ sample. (D) Percentage of top-strand reads containing a D-loop footprint in DMA assay. (E) Table summarizing the total number of reads containing a footprint as ‘peak’ and the total number of reads analyzed as ‘total’ for each strand. ‘% D-loops/% Peak’ indicate the percentage of reads containing a footprint. The data represents a cumulation from two or three independent replicates for D-loop samples formed in presence of increasing concentration of Tid1 and the ds98-931 substrate.
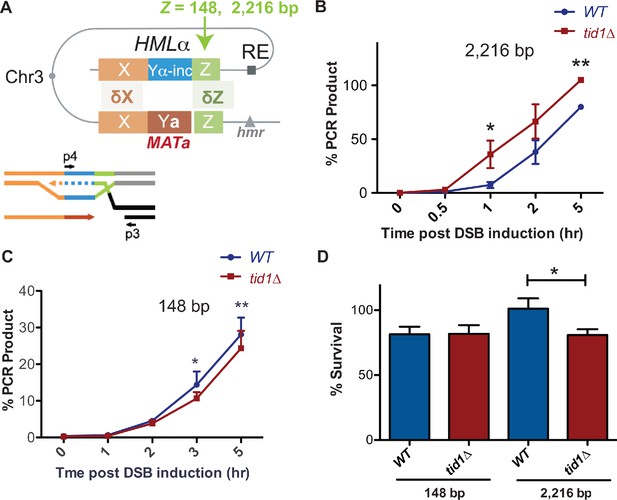
Tid1 affects kinetics of D-loop extension and cell survival depending on the length of homology between the donor and MAT during mating type switch.
(A) Schematic of the primer extension assay adopted from Mehta et al., 2017. Homology length to the invading Z-end (green) was altered at HML to be either 148 bp or 2216 bp. Arrowhead indicates HO-cut site. Primers (indicated by black arrows as p3 and p4) specific to the newly synthesized DNA (shown by dotted lines) after strand invasion into HML was used to quantify extended D-loops by the primer extension assay. (B, C) Kinetics of new DNA synthesis post D-loop formation as measured the primer extension assay. Graphs show qRT-PCR product at intervals post HO endonuclease induction by galactose in strains with 148 bp or 2216 bp homology at the Z-end. The amount of PCR product obtained from a switched MATα-inc colony was set to 100%. The Cp values were normalized to Arginine PCR product formation as in Mehta et al., 2017. Mean ± SD (n = 3). (D) Viability of strains having 148 bp or 2216 bp homology at the Z-end between HML and MAT loci after HO endonuclease induction by galactose. Mean ± SD (n ≥ 3). * indicates p-value<0.05 with paired two-tailed t-test.
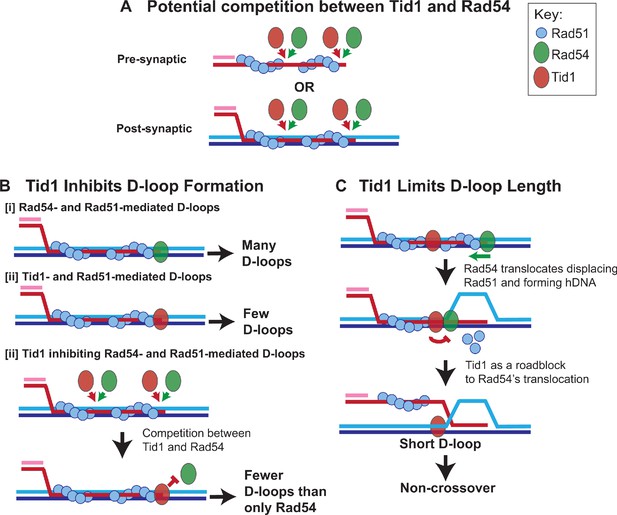
Model: Tid1 competes with Rad54 in binding to the filament ends, limits D-loop length and may promote non-crossover outcome.
(A) Model depicts Tid1 competing with Rad54 for binding to Rad51 filament ends either at pre-synaptic or post-synaptic stage. Since Rad51 is not as cooperative as its bacterial homolog, RecA (Galletto et al., 2006; Sanchez et al., 2013), gaps in the filament can be expected, that provide potential binding sites for Rad54 and/or Tid1. Both Tid1 (Chi et al., 2006; Petukhova et al., 2000; Santa Maria et al., 2013) and Rad54 (Raschle et al., 2004) interact with Rad51 via their N-terminal domain and are recruited to DSB sites (Kwon et al., 2008). (B) Model depicting in vitro D-loop levels when formed in presence of Rad54 and/or Tid1 and Rad51. (C) A ‘roadblock model’ explaining the effect of Tid1 on D-loop length. Tid1 potentially bound intermittently within a pre- or post-synaptic Rad51 filament, can act as a physical roadblock to Rad54 translocation. Rad54 translocation is stimulated by Rad51, simultaneously displacing Rad51 and forming a hDNA (Wright and Heyer, 2014). When Rad54 encounters Tid1, the N-terminal domain disengages resulting in the formation of shorter D-loops. Short, dynamic D-loops can subsequently prevent dHJ formation and the possibility of a crossover outcome.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (Saccharomyces cerevisiae) | See Supplementary file 1 | |||
| Antibody | Tid1 antibody (rabbit polyclonal) | Heyer laboratory | 1:200 | |
| Antibody | GAPDH antibody (mouse monoclonal) | Invitrogen | Catalog: #MA5-15738 | 1:5000 |
| Recombinant DNA reagent | pWDH597 | Wright and Heyer, 2014 | Amp, URA3 markers | Plasmid for overexpression of S. cerevisiae Tid1/Tid1-KR N-terminally tagged with GST, removable by cleavage with PreScission protease. |
| Recombinant DNA reagent | pUC19 | Addgene | Catalog: #50005 | Plasmid used to test topoisomerase contamination of purified protein |
| Recombinant DNA reagent | pBSphix1200 | Wright and Heyer, 2014 | Amp | Plasmid used as dsDNA donor in D-loop assay in supercoiled or linear form |
| Sequence-based reagent | 100-mer | Wright and Heyer, 2014 | ctggtcataatcatggtggcgaataagtacgcgttcttgcaaatcaccagaaggcggttcctgaatgaatgggaagccttcaagaaggtgataagcagga | |
| Sequence-based reagent | ds98-197-78ss | Wright and Heyer, 2014 | Homologous sequence, 197 nt: ctggtcataatcatggtggcgaataagtacgcgttcttgcaaatcaccagaaggcggttcctgaatgaatgggaagccttcaagaaggtgataagcaggagaaacatacgaaggcgcataacgataccactgaccctcagcaatcttaaacttcttagacgaatcaccagaacggaaaacatccttcatagaaattt | |
| Sequence-based reagent | ds98-607 | Wright and Heyer, 2014 | Homologous sequence, 607 nt: gaagtcatgattgaatcgcgagtggtcggcagattgcgataaacggtcacattaaatttaacctgactattccactgcaacaactgaacggactggaaacactggtcataatcatggtggcgaataagtacgcgttcttgcaaatcaccagaaggcggttcctgaatgaatgggaagccttcaagaaggtgataagcaggagaaacatacgaaggcgcataacgataccactgaccctcagcaatcttaaacttcttagacgaatcaccagaacggaaaacatccttcatagaaatttcacgcggcggcaagttgccatacaaaacagggtcgccagcaatatcggtataagtcaaagcacctttagcgttaaggtactgaatctctttagtcgcagtaggcggaaaacgaacaagcgcaagagtaaacatagtgccatgctcaggaacaaagaaacgcggcacagaatgtttataggtctgttgaacacgaccagaaaactggcctaacgacgtttggtcagttccatcaacatcatagccagatgcccagagattagagcgcatgacaagtaaaggacggttgtcagcgtcataagaggttttac | |
| Sequence-based reagent | ds98-931 | Heyer laboratory | Homologous sequence, 931 nt: gaacggaaaacatccttcatagaaatttcacgcggcggcaagttgccatacaaaacagggtcgccagcaatatcggtataagtcaaagcacctttagcgttaaggtactgaatctctttagtcgcagtaggcggaaaacgaacaagcgcaagagtaaacatagtgccatgctcaggaacaaagaaacgcggcacagaatgtttataggtctgttgaacacgaccagaaaactggcctaacgacgtttggtcagttccatcaacatcatagccagatgcccagagattagagcgcatgacaagtaaaggacggttgtcagcgtcataagaggttttacctccaaatgaagaaataacatcatggtaacgctgcatgaagtaatcacgttcttggtcagtatgcaaattagcataagcagcttgcagacccataatgtcaatagatgtggtagaagtcgtcatttggcgagaaagctcagtctcaggaggaagcggagcagtccaaatgtttttgagatggcagcaacggaaaccataacgagcatcatcttgattaagctcattagggttagcctcggtacggtcaggcatccacggcgctttaaaatagttgttatagatattcaaataaccctgaaacaaatgcttagggattttattggtatcagggttaatcgtgccaagaaaagcggcatggtcaatataaccagtagtgttaacagtcgggagaggagtggcattaacaccatccttcatgaacttaatccactgttcaccataaacgtgacgatgagggacataaaaagtaaaaatgtctacagtagagtcaatagcaaggccacgacgcaatggagaaagacggagagcgccaacggcgtccatctcgaaggagtcgccagcgataaccggagtagttgaaatggtaataagac | |
| Sequence-based reagent | ds98-915 | Heyer laboratory | Homologous sequence, 915 nt: gaagtcatgattgaatcgcgagtggtcggcagattgcgataaacggtcacattaaatttaacctgactattccactgcaacaactgaacggactggaaacactggtcataatcatggtggcgaataagtacgcgttcttgcaaatcaccagaaggcggttcctgaatgaatgggaagccttcaagaaggtgataagcaggagaaacatacgaaggcgcataacgataccactgaccctcagcaatcttaaacttcttagacgaatcaccagaacggaaaacatccttcatagaaatttcacgcggcggcaagttgccatacaaaacagggtcgccagcaatatcggtataagtcaaagcacctttagcgttaaggtactgaatctctttagtcgcagtaggcggaaaacgaacaagcgcaagagtaaacatagtgccatgctcaggaacaaagaaacgcggcacagaatgtttataggtctgttgaacacgaccagaaaactggcctaacgacgtttggtcagttccatcaacatcatagccagatgcccagagattagagcgcatgacaagtaaaggacggttgtcagcgtcataagaggttttacctccaaatgaagaaataacatcatggtaacgctgcatgaagtaatcacgttcttggtcagtatgcaaattagcataagcagcttgcagacccataatgtcaatagatgtggtagaagtcgtcatttggcgagaaagctcagtctcaggaggaagcggagcagtccaaatgtttttgagatggcagcaacggaaaccataacgagcatcatcttgattaagctcattagggttagcctcggtacggtcaggcatccacggcgctttaaaatagttgttatagatattcaaataaccctgaaacaaatgc | |
| Sequence-based reagent | ds98-915-78ss | Heyer laboratory | Homologous sequence, 915 nt: gaagtcatgattgaatcgcgagtggtcggcagattgcgataaacggtcacattaaatttaacctgactattccactgcaacaactgaacggactggaaacactggtcataatcatggtggcgaataagtacgcgttcttgcaaatcaccagaaggcggttcctgaatgaatgggaagccttcaagaaggtgataagcaggagaaacatacgaaggcgcataacgataccactgaccctcagcaatcttaaacttcttagacgaatcaccagaacggaaaacatccttcatagaaatttcacgcggcggcaagttgccatacaaaacagggtcgccagcaatatcggtataagtcaaagcacctttagcgttaaggtactgaatctctttagtcgcagtaggcggaaaacgaacaagcgcaagagtaaacatagtgccatgctcaggaacaaagaaacgcggcacagaatgtttataggtctgttgaacacgaccagaaaactggcctaacgacgtttggtcagttccatcaacatcatagccagatgcccagagattagagcgcatgacaagtaaaggacggttgtcagcgtcataagaggttttacctccaaatgaagaaataacatcatggtaacgctgcatgaagtaatcacgttcttggtcagtatgcaaattagcataagcagcttgcagacccataatgtcaatagatgtggtagaagtcgtcatttggcgagaaagctcagtctcaggaggaagcggagcagtccaaatgtttttgagatggcagcaacggaaaccataacgagcatcatcttgattaagctcattagggttagcctcggtacggtcaggcatccacggcgctttaaaatagttgttatagatattcaaataaccctgaaacaaatgc | |
| Sequence-based reagent | HOCSp3; MATp13 | Mehta et al., 2017 | Primer pair for measuring extension of D-loops. HOCsp3: GACAAAATGCAGCACGGAAT MATp13: GTTAAGATAAGAACAAAGAAgGATGCT | |
| Sequence-based reagent | olWDH1760 olWDH1761 | Piazza et al., 2019 | Primer pair for reference locus ARG4 on Ch. VIII. olWDH1760: AGACAGAATTGGCAAAGATCC olWDH1761: GGCCAATTAGTTCACCAAGACG | |
| Sequence-based reagent | olWDH1766 olWDH1767 | Piazza et al., 2019 | Primer pair for measuring dsDNA integrity at the HOcs. olWDH1766: GTTTCAGCTTTCCGCAACAG olWDH1767: GGCGAGGTATTGGATAGTTCC | |
| Peptide, recombinant protein | GST-Tid1 | Nimonkar et al., 2007 | ||
| Peptide, recombinant protein | GST-Tid1-K318R | Nimonkar et al., 2007 | ||
| Peptide, recombinant protein | Rad54 | Wright and Heyer, 2014 | ||
| Peptide, recombinant protein | Rad51 | Van Komen et al., 2006 | ||
| Peptide, recombinant protein | RPA | Binz et al., 2006 | ||
| Peptide, recombinant protein | Bsa1 | New England Biolabs | Catalog: #R0535S | To linearize pBSphix1200 |
| Peptide, recombinant protein | T4 Polynucleotide kinase | New England Biolabs | Catalog: #M0201S | |
| Peptide, recombinant protein | Phusion-U polymerase | Thermo Fischer | Catalog: #PN-F555S | |
| Chemical compound, drug | NADH | Sigma | Catalog: #606-68-6 | ATPase assay |
| Chemical compound, drug | Sera-Mag SpeedBead Carboxylate-Modified Magnetic particles, hydrophobic | Sigma | Catalog: #PN-65152105050250 | DMA |
| Chemical compound, drug | AMPure PB | Pacific Biosciences | Catalog: #100-265-900 | DMA |
| Commercial assay or kit | Baker Flex, Cellulose PEI-F | Fischer Scientific | Catalog: #9004-34-6 | ATPase assay |
| Commercial assay or kit | Epitect Bisulfite kit | Qiagen | Catalog: #59104 | DMA |
| Commercial assay or kit | SMRTbell Template Prep Kit 1.0 | Pacific Biosciences | Catalog: #100-259-100 | DMA |
Additional files
-
Source data 1
Source data for all the figures.
- https://cdn.elifesciences.org/articles/59112/elife-59112-data1-v2.xlsx
-
Supplementary file 1
Saccharomyces cerevisiae strains used in this work.
1 W303 strain background. 2 S288c strain background. 3 Obtained from Mehta et al., 2017.
- https://cdn.elifesciences.org/articles/59112/elife-59112-supp1-v2.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/59112/elife-59112-transrepform-v2.docx





