An AMPK-dependent, non-canonical p53 pathway plays a key role in adipocyte metabolic reprogramming
Figures
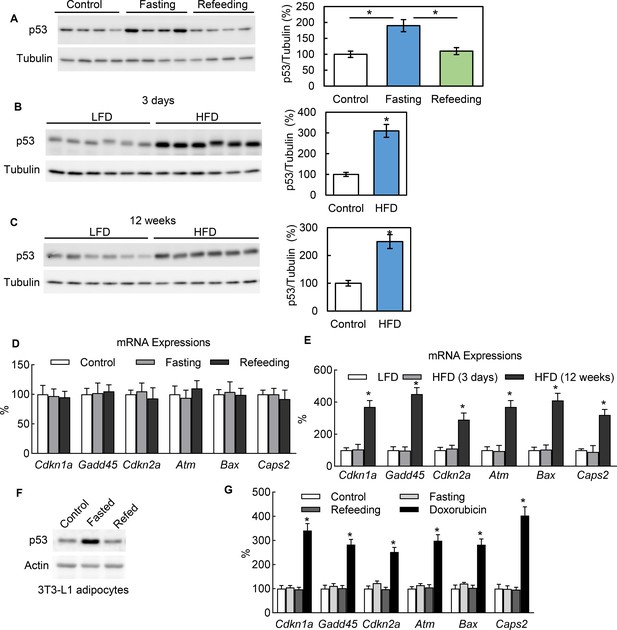
In adipocytes p53 can be up-regulated by nutrient starvation, but without canonical pathway activation.
(A) Three groups of WT C57 mice were either not treated (Control), fasted for 24 hr (Fasting), or followed by refeeding for 24 hr (Refeeding). Primary adipocytes isolated from epididymal fat tissue were subjected to western blot using indicated antibodies. (B–C) p53 protein levels in isolated primary adipocytes from mice fed a high-fat diet (HFD) for 3 days (B) or 12 weeks (C) were examined by western blots. The quantification of relative p53 expression levels was normalized by tubulin and shown in bar graphs. (D–E) Gene expressions of p53 canonical pathway were measured by RT-PCR in samples from A-C. (F) The protein expression levels of p53 and actin were measured from 3T3-L1 adipocytes cultured in three media conditions: normal culture (DMEM with 10% FBS), fasting (PBS with 1% BSA and ISO, 10 mM for 6 hr), and refeeding (normal plus insulin for 6 hr after fasting). (G) Gene expressions of p53 canonical pathway were measured by RT-PCR in samples from F.
-
Figure 1—source data 1
An Excel sheet with numerical quantification data.
- https://cdn.elifesciences.org/articles/63665/elife-63665-fig1-data1-v3.xlsx
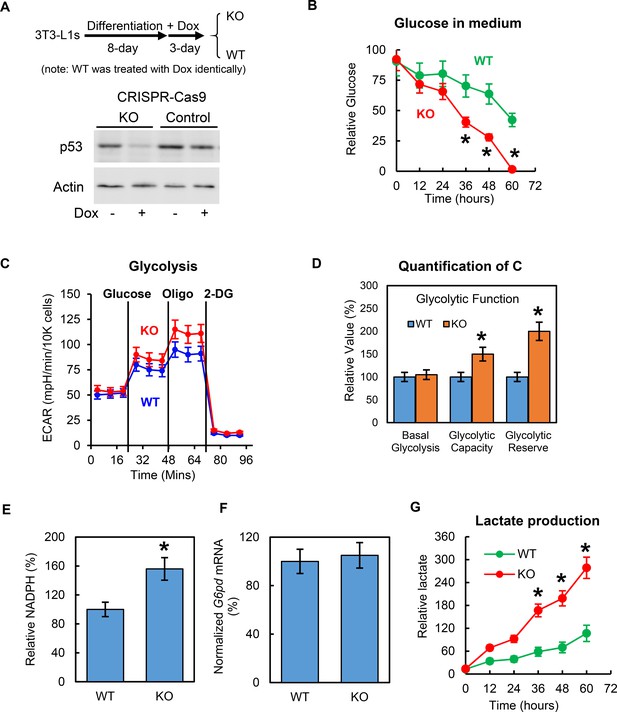
Deletion of p53 increases adipocyte glucose metabolism.
(A) In vitro working model for inducible p53 knockout (KO) in 3T3-L1 adipocytes. (B) p53-knockout (KO) and wild-type (WT) cells were cultured in no-glucose DMEM with 0.1% FBS for overnight, then glucose was added to media to a final concentration of 5 mM. 10–20 μl culture media were sampled every 12 hr. Glucose concentrations were measured using commercially available kits. (C–D) Glycolytic and mitochondrial functions were measured by Seahorse. Relative NADPH levels (E) and G6DP mRNA expression levels (F) were examined. (G) Lactate levels were measured similarly as in (B). (*p<0.05).
-
Figure 2—source data 1
An Excel sheet with numerical quantification data.
- https://cdn.elifesciences.org/articles/63665/elife-63665-fig2-data1-v3.xlsx
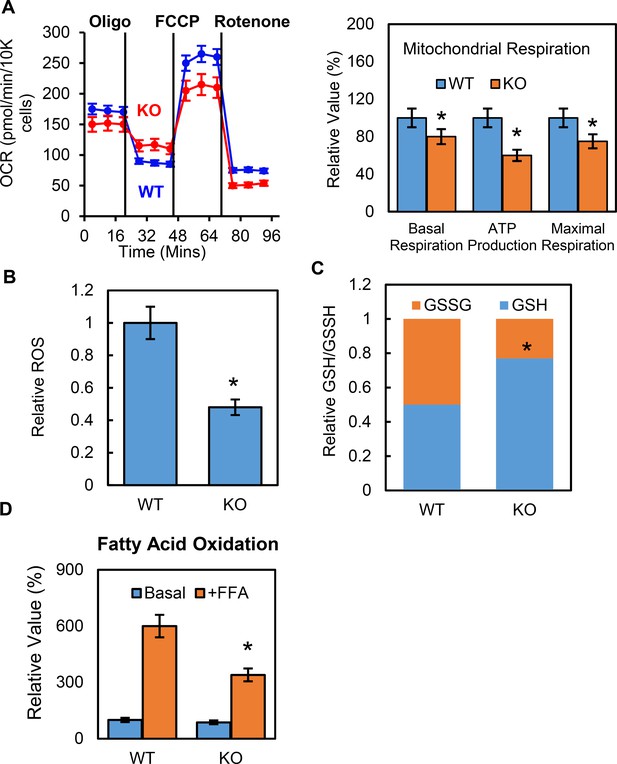
Deletion of p53 suppresses adipocyte mitochondrial respiration and fatty acid oxidation.
(A) Mitochondrial respiration was measured by Seahorse XF technology in p53-KO and WT control 3T3-L1 adipocytes. (B–C) ROS, GSSG, and GSH were measured using commercially available kits. (D) Fatty acid oxidation was measured using Seahorse XF technology. (*p<0.05).
-
Figure 3—source data 1
An Excel sheet with numerical quantification data.
- https://cdn.elifesciences.org/articles/63665/elife-63665-fig3-data1-v3.xlsx
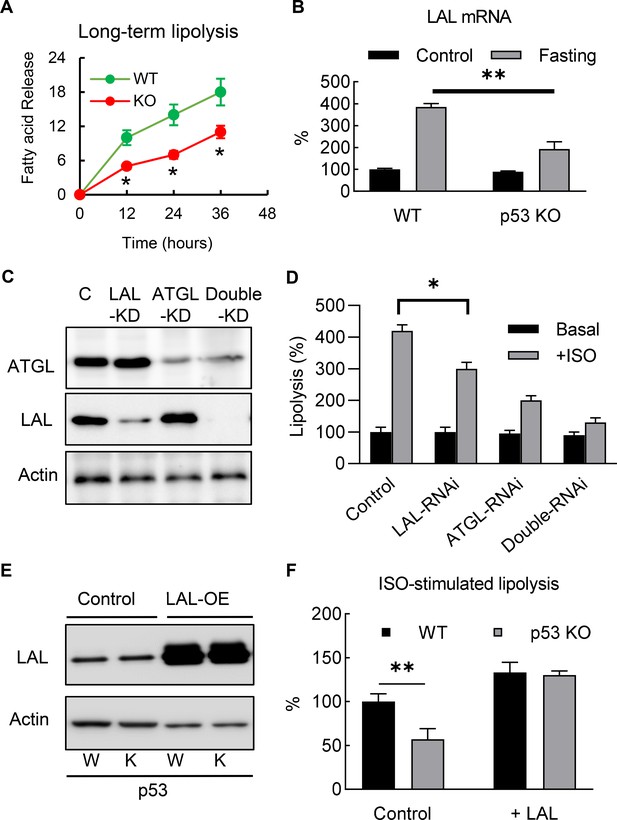
Deletion of p53 suppresses adipocyte lipid metabolism through the down-regulation of lysosomal acid lipase.
(A) Long-term lipolysis activities were measured in Iso-stimulated p53-knockout (KO) and wild-type (WT) control adipocytes. (B) LAL mRNA expression levels were examined in the samples from the end time point (36 hr) of A (C) Protein expression levels of LAL, ATGL, and actin were examined by western blots in control, LAL, ATGL, and LAL/ATGL double RNAi knockdown cells. (D) 3T3-L1 adipocytes from C were cultured with or without ISO stimulation (in PBS +1% BSA) for 18 hr and the concentrations of fatty acid in the medium were measured. (*p<0.05). (E) LAL was overexpressed in WT and p53–KO 3T3-L1 adipocytes, and (F) long-term lipolysis (18 hr) activities were measured by determining FFA levels. (*p<0.05, **p<0.01).
-
Figure 4—source data 1
An Excel sheet with numerical quantification data.
- https://cdn.elifesciences.org/articles/63665/elife-63665-fig4-data1-v3.xlsx
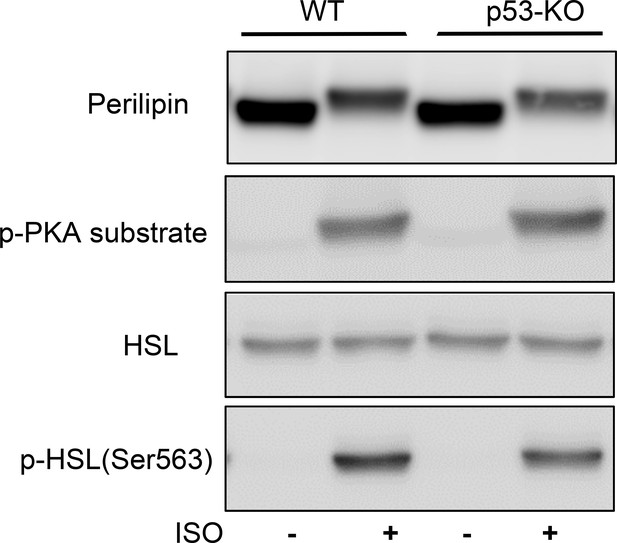
Deletion of p53 does not affect traditional neutral lipolysis signaling pathway.
Total and phosphor-perillipin and HSL expression level in ISO-stimulated p53-knockout (p53–KO) and wild-type (WT) control adipocytes were examined by western blots.
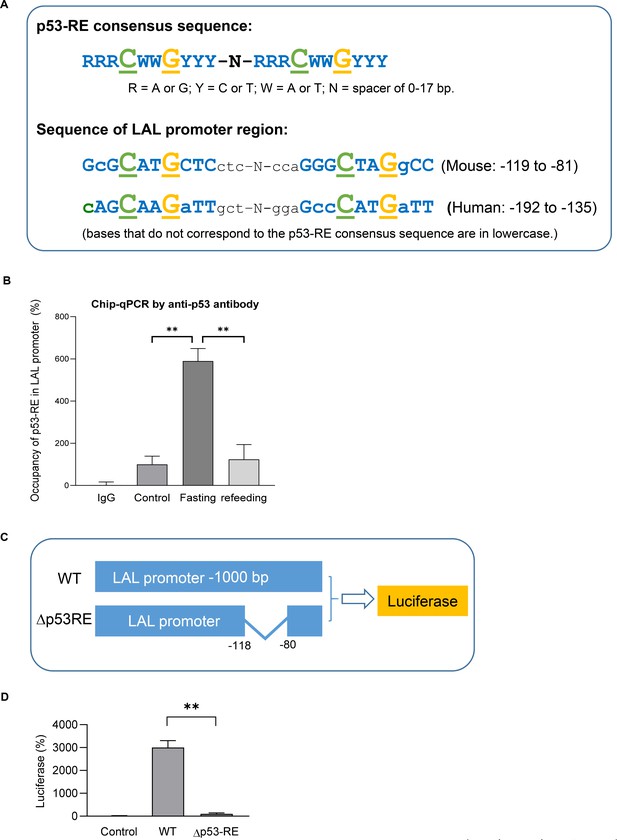
LAL is a transcription target of p53.
(A) Previous known p53 response-element (p53–RE) consists of two copies of the palindromic half-site RRRCWWGYYY where each p53 monomer binds five nucleotides (top). A highly p53RE-like sequence was found in LAL promoter region in both human and mouse (bottom). (B) Chip-qPCR assay using IgG or p53 antibody from control, 24 hr fasted, and refed mouse primary adipocytes. (C) Luciferase reporter constructs were generated by using 1 kb wild-type (WT) or p53-RE deleted (Δp53RE) LAL promoter sequence. (D) 3T3-L1 adipocyte were transiently transfected by electroporation with constructs from C together with pcDNA3.1-p53 construct. After 48 hr, cells were harvested in the reporter lysis buffer. Luciferase activity in cell lysates was assayed as described under ‘Experimental Procedures’ and normalized by protein concentrations. Data are presented for triplicate samples as mean ± S.D. (error bars). **, p<0.01.
-
Figure 5—source data 1
An Excel sheet with numerical quantification data.
- https://cdn.elifesciences.org/articles/63665/elife-63665-fig5-data1-v3.xlsx
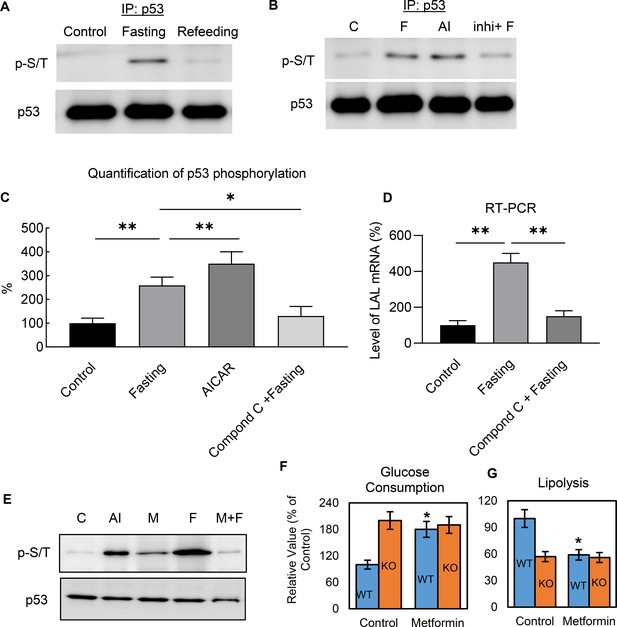
Metabolic regulation of p53 is mediated through AMPK-dependent phosphorylation.
(A–B) Nuclear fraction was prepared from control (C, High-glucose DMEM with 10% FBS), fasted (F, PBS with 1% BSA and ISO for 6 hr), AICAR (AI), or compound C treated before fasting (Inhi+F), 3T3-L1 adipocytes, following by immunoprecipitation and western blots. (C) Quantification of B. (D) LAL mRNA expression levels were examined by RT-PCR in the cells from B. (E) The effect of metformin on fasting-induced p53 phosphorylation was determined by immunoprecipitation and western blots. (F–G) Effects of metformin on glucose consumption and lipolysis in cultured 3T3-L1 WT and p53-KO adipocytes were determined. (*p<0.05).
-
Figure 6—source data 1
An Excel sheet with numerical quantification data.
- https://cdn.elifesciences.org/articles/63665/elife-63665-fig6-data1-v3.xlsx
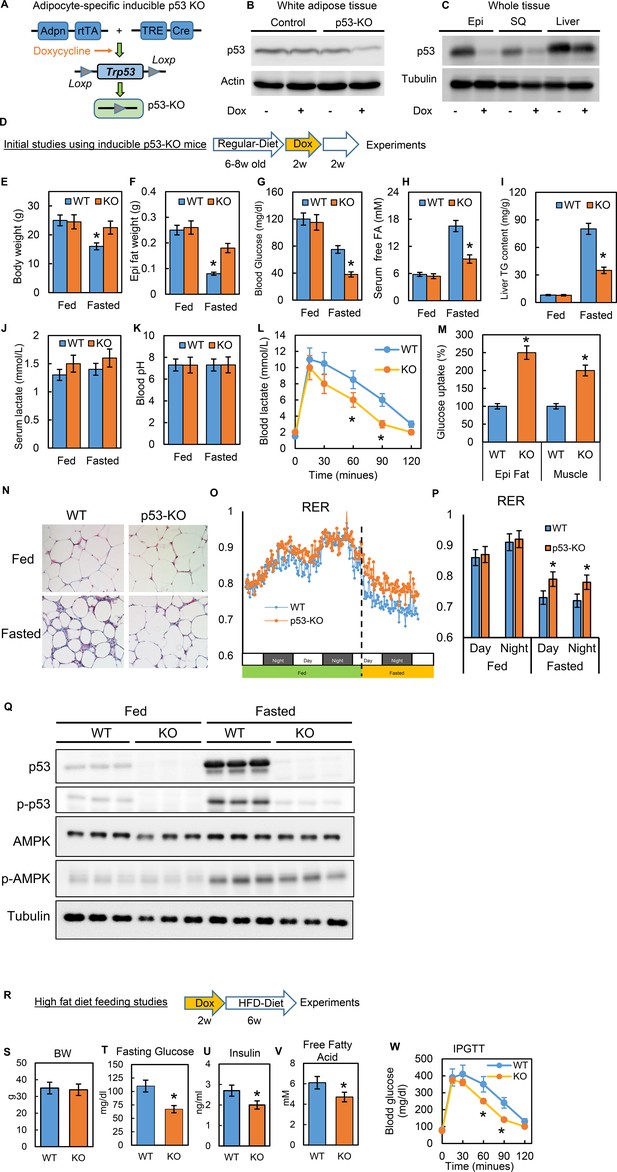
Inducible adipocyte-specific p53 knockout (p53-iAKO) mice show decreased lipolysis after fasting and improved metabolic phenotypes upon high-fat diet feeding.
(A) iAdipo-p53-KO (KO) mouse was generated by breeding doxycycline-inducible (TRE-Cre), adipocyte-specific (Adpn-rtTA), floxed-p53 lines together, and induced by doxycycline. (B) white adipose tissue and (C) whole tissue from control (rtTA/Cre only) and KO mice fed with or without doxycycline diet were subjected to western blot by using indicated antibodies. (Epi: epididymal fat; SQ: subcutaneous fat). (D) Initial study design. (E–K) KO and WT mice were fasted for 24 hr and refed for 24 hr or not; body weight (E), epididymal fat tissue weight (F), blood glucose(G), serum-free fatty acid (H), liver TG content (I), serum lactate concentration (J), and blood pH (K) were measured. Lactate tolerance test (L) and in-vivo glucose uptake assay (M) were performed in WT and KO mice. (N) The histology H-E staining was conducted in epididymal fat tissues from WT and p53-KO mice under feeding (fed) and fasting (Fasted, 24 hr) conditions. (O–P) Respiratory exchange ratio (RER) was measured in metabolic cage studies. (Q) Total and phosphor-p53, AMPK, and tubulin expression levels in fed and fasted p53-knockout (KO) and wild-type (WT) mice were examined by western blots. (R) Study design of high-fat diet feeding studies. Metabolic phenotyping in high-fat diet-fed p53-knockout (KO) mice compared to WT, including (S) body weight (BW), (T) fasting glucose, (U) insulin, (V) fatty acids, and (W) IPGTT. (n = 3–6, *p<0.05).
-
Figure 7—source data 1
An Excel sheet with numerical quantification data.
- https://cdn.elifesciences.org/articles/63665/elife-63665-fig7-data1-v3.xlsx
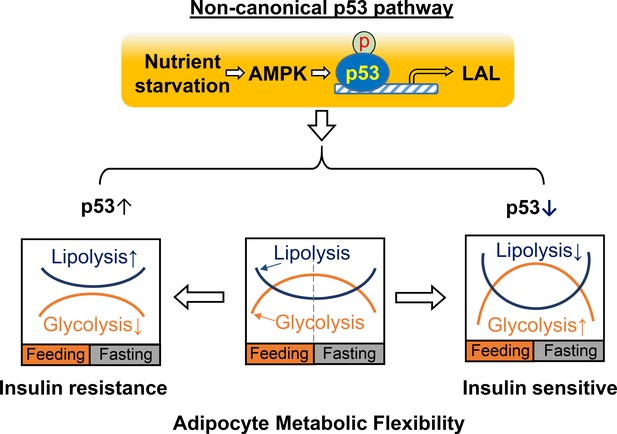
A working model for non-canonical p53 pathway-regulated adipocyte metabolic flexibility.
Nutrient starvation causes an AMPK-dependent p53 non-canonical pathway activation, which leads to the upregulation of LAL-mediated lysosome acid lipolysis. This pathway plays key role for adipocyte oscillating between glycolysis and lipolysis under feeding and fasting states. In insulin resistance, p53 is found to be upregulated, while p53 downregulation is associated with insulin sensitivity.
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Genetic reagent (Mus. musculus) | B6.Cg-Tg(tetO-cre) (TRE-Cre mouse) | Jackson Laboratory | JAX stock #006234; RRID:IMSR_JAX:006234 | |
| Genetic reagent (Mus. musculus) | Adipoq-rtTA mouse | Sun et al., 2012; Wang et al., 2013 | A kind gift from Dr. Scherer. | |
| Genetic reagent (Mus. musculus) | B6.129P2-Trp53tm1Brn/J (floxed-Trp53 mouse) | Jackson Laboratory | JAX stock #008462; RRID:IMSR_JAX:008462 | |
| other | Doxycycline diet | BioServ | #S3888 | |
| other | Control diet | BioServ | #S4207 | |
| other | 60% high-fat diet | Research Diets | D12492 | |
| Cell line (mouse) | 3T3-L1 cell line. (mouse) | Zenbio | Zenbio: SP-L1-F; RRID:CVCL_0123 | |
| Transfected construct (mouse) | Mouse Lentiviral Trp53 sgRNA, glycerol stock | GE Healthcare Dharmacon | GSGM11839-246656039 | Target sequence: CTGTACGGCGGTCTCTCCCA |
| Transfected construct (mouse) | Mouse Lentiviral Trp53 sgRNA, glycerol stock | GE Healthcare Dharmacon | GSGM11839-247039414 | Target sequence: CTCCAGAAGATATCCTGGTA |
| Transfected construct (mouse) | Mouse Lentiviral Trp53 sgRNA, glycerol stock | GE Healthcare Dharmacon | GSGM11839-246656045 | Target sequence: GTGATGGGAGCTAGCAGTTT |
| recombinant DNA reagent | Inducible Cas9 lentiviral vector plasmid DNA | GE Healthcare Dharmacon | CAS11229 | TRE3G dox-inducible promoter. |
| recombinant DNA reagent | MGC mouse Lipa cDNA plasmid DNA | GE Healthcare Dharmacon | MMM1013-202770272 | Mammalian Expression LAL by CMV promoter |
| Transfected construct (mouse) | siRNA: Lipa | Thermo Fisher Scientific | AM16708 | siRNA ID 75920 |
| Transfected construct (mouse) | siRNA: Lipa | Thermo Fisher Scientific | AM16708 | siRNA ID 157079 |
| Transfected construct (mouse) | siRNA: Lipa | Thermo Fisher Scientific | AM16708 | siRNA ID 75827 |
| Transfected construct (mouse) | siRNA: Pnpla2 | Thermo Fisher Scientific | AM16708 | siRNA ID 183465 |
| Transfected construct (mouse) | siRNA: Pnpla2 | Thermo Fisher Scientific | AM16708 | siRNA ID 183466 |
| Transfected construct (mouse) | siRNA: Pnpla2 | Thermo Fisher Scientific | AM16708 | siRNA ID 79116 |
| antibody | anti-p53 (rabbit polyclonal) | Santa Cruz | Santa Cruz: sc-6243; RRID:AB_653753 | WB. 4-degree overnight incubation only. 1:1000 |
| antibody | anti-p53 (mouse monoclonal) | Cell signaling | Cell signaling: 2524; RRID:AB_331743 | WB. 1:1000 |
| antibody | anti-p53 (mouse monoclonal) | Abcam | Abcam: ab26; RRID:AB_303198 | IP. 1:100 |
| antibody | anti-p53 (mouse monoclonal) | Thermo Fisher | Thermo Fisher: AHO0112; RRID:AB_2536305 | IP.1:100 |
| antibody | anti-Tubulin (mouse monoclonal) | Sigma | Sigma:T4026; RRID:AB_477577 | 1:2000 |
| antibody | anti-Actin (rabbit polyclonal) | Cell signaling | Cell signaling: 4967; RRID:AB_330288 | 1:1000 |
| antibody | anti-ATGL (rabbit polyclonal) | Cell signaling | Cell signaling: #2138; RRID:AB_2167955 | 1:1000 |
| antibody | anti-LAL (rabbit polyclonal) | Abcam | Abcam: ab154356; RRID:AB_154356 | 1:5000 |
| antibody | anti-Perilipin (Guinea Pig polyclonal) | Fitzgerald | Fitzgerald:20R-PP004; RRID:AB_1288416 | 1:5000 |
| antibody | anti- Phospho-(Ser/Thr) PKA Substrate (rabbit polyclonal) | Cell Signaling | Cell signaling: #9621; RRID:AB_330304 | 1:1000 |
| antibody | anti-HSL (rabbit polyclonal) | Cell Signaling | Cell signaling: #4107; RRID:AB_2296900 | 1:1000 |
| antibody | anti-p-HSL (Ser563) (rabbit polyclonal) | Cell Signaling | Cell signaling: #4139; RRID:AB_2135495 | 1:1000 |
| antibody | anti-AMPKα (rabbit polyclonal) | Cell Signaling | Cell signaling: #2532; RRID:AB_330331 | 1:1000 |
| antibody | Anti-phospho-AMPKα (Thr172) (rabbit polyclonal) | Cell Signaling | Cell signaling: #2535; RRID:AB_331250 | 1:1000 |
| sequence-based reagent | Lipa-mRNA-F | This paper | qPCR primers | TGTGACCGAGATAATCATGCG |
| sequence-based reagent | Lipa-mRNA-R | This paper | qPCR primers | GAAGATACACAACTGGTCTGGG |
| sequence-based reagent | Lipa-promoter-F | This paper | Chip-qPCR primers | AAGCTCTGGCTGGGCTTAGAG |
| sequence-based reagent | Lipa-promoter-R | This paper | Chip-qPCR primers | GCAGGCGAGCTTGGCCAACCT |





