Enhanced Cas12a multi-gene regulation using a CRISPR array separator
Figures
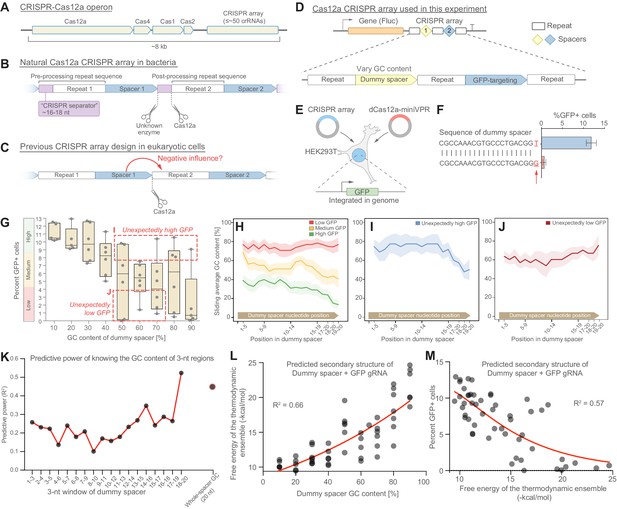
gRNA performance is affected by the GC content of the upstream spacer.
(A) The CRISPR-Cas12a operon consists of Cas genes and a CRISPR array. (B) Each gRNA consists of a repeat and a spacer. Pre-processing repeats contain a ~16–18 nt fragment, here denoted CRISPR separator, which gets excised by Cas12a and an unknown enzyme. (C) The separator has previously been omitted when expressing Cas12a arrays in mammalian cells. We asked if the separator serves to insulate gRNAs from the negative influence of secondary structure in spacers. (D) We designed CRISPR arrays consisting of two gRNAs, the first with a non-targeting dummy spacer, and the second targeting the promoter of GFP, genomically integrated in HEK293T cells. (E) Experimental setup; Lb-dCas12a-miniVPR was used to activate GFP, and GFP fluorescence was analyzed as a measure of array performance. (F) CRISPR arrays can display hypersensitivity to the composition of the dummy spacer. In extreme cases, replacing the last nucleotide from T to G can lead to almost complete abrogation of GFP activation. (G) A library of 51 CRISPR arrays, each with a dummy spacer of different GC content. A strong negative correlation is seen between the GC content of the dummy spacer and GFP fluorescence. Each dot represents one of the 51 CRISPR arrays (average of three replicates). Arrays were divided into three groups based on the level of GFP fluorescence they enabled. Boxes indicate two groups that were analyzed in (I and J). (H–J) For each group, the average GC content of a sliding 5-nt window was calculated. The best-performing arrays were the ones where the dummy spacer happened to have low GC content at its 3’ end (H). Some arrays showed unexpectedly high or low GFP activity for the GC content of their dummy spacers (G). These arrays contain low (I) or high (J) GC content at the very 3’ end of their dummy spacers, suggesting that the GC content of the last few bases is an important predictor of array performance. Shaded regions in (H–J) represent standard error. (K) The predictive power of knowing the GC content of 3-nt regions in the dummy gRNA (Materials and methods). Merely knowing the GC content of the las three bases is more predictive than knowing the overall GC content. (L) A plot showing the relationship between GC content of the 51 dummy spacers and the secondary structures they are predicted to form with the GFP-targeting gRNA (the larger the value on the y-axis, the more stable the predicted secondary structure). (M) This predicted secondary structure formation is anticorrelated with performance of the GFP-targeting spacer, suggesting that strong secondary structures is what impedes array performance.
-
Figure 1—source data 1
Raw data used for panel G.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig1-data1-v3.xlsx
-
Figure 1—source data 2
Raw data used for panel L.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig1-data2-v3.xlsx
-
Figure 1—source data 3
Raw data used for panel M.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig1-data3-v3.xlsx
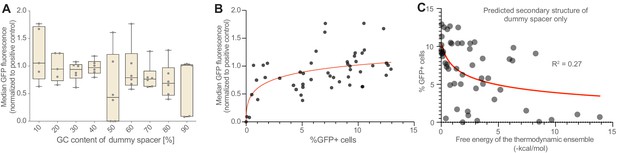
CRISPR activation of genomically integrated GFP.
(A) The negative correlation between GFP activation and dummy spacer GC content (Figure 1G) is seen also when measuring median GFP fluorescence, though percentage of cells that activate GFP is a more sensitive measure of CRISPR activation effects than the median GFP fluorescence (B), at least in this experimental system. (C) Predicted secondary structure formation is anticorrelated with array performance (Figure 1M); this correlation is weaker if only the dummy spacer (rather than the dummy spacer plus the subsequent GFP-targeting gRNA) is included in the secondary structure prediction tool (R2 = 0.27 vs 0.57; compare Figure 1M). This suggests that secondary structures that involve both the dummy spacer and the GFP-targeting gRNA are particularly disruptive.
-
Figure 1—figure supplement 1—source data 1
Raw data used for panel C.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig1-figsupp1-data1-v3.xlsx
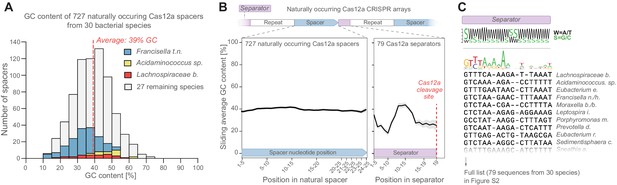
Natural CRISPR arrays contain separator sequences with low GC content.
(A) The GC content of naturally occurring CRISPR-Cas12a spacers display no obvious depletion of high-GC spacers (though spacer GC content is weakly correlated with overall genomic GC content, Figure 2—figure supplement 1A). Spacers from commonly used Cas12a variants (Lb, As, Fn) are highlighted in color. (B) Neither do these spacers show low GC content at their 3’ ends. But the separator sequences of these gRNAs have low GC content. (C) This is seen also in a multiple-sequence alignment of 79 natural separator sequences, which suggests that one purpose of the CRISPR separator is to act as an insulator between adjacent gRNAs in a Cas12a CRISPR array.
-
Figure 2—source data 1
Raw data used for panel A.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig2-data1-v3.xlsx
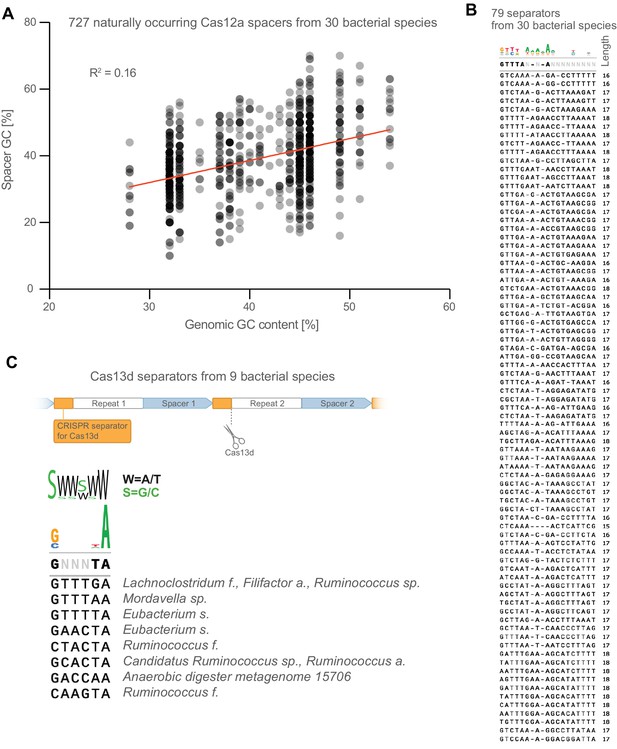
Further analysis of naturally occurring crRNA sequences.
(A) The GC content of naturally occurring spacers is weakly correlated with overall genomic GC content of their bacterial host. (B) Full list of CRISPR-separator sequences from Figure 2C. (C) CRISPR separators from the type VI CRISPR Cas13d are also AT-rich, particularly the two bases at the very 3’ end.
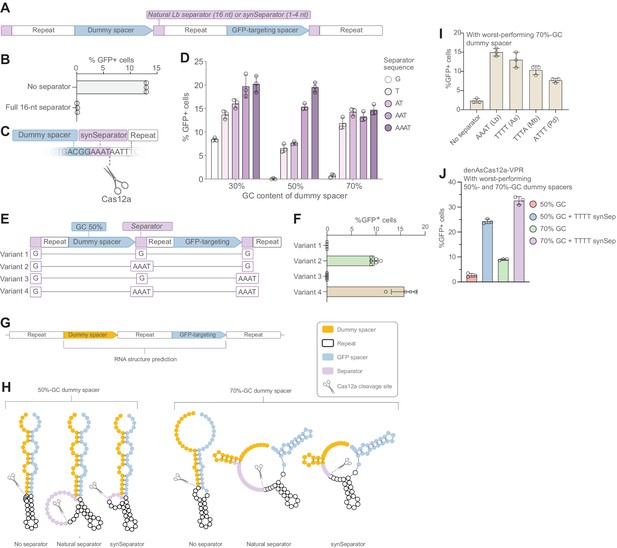
A synthetic separator improves CRISPR array performance in human cells.
(A) A diagram showing how we introduced either the full, 16-nt separator sequence from L. bacterium or a short 1–4 nt synthetic separator (synSeparator) between each gRNA in the CRISPR array. (B) The full, 16-nt separator sequence almost completely eliminates GFP activation. (C) Diagram showing inclusion of synSeparator sequences from the 3’ end of the L. bacterium in the CRISPR array. Each array contained a synthetic separator (G, T, AT, AAT, or AAAT) upstream of every repeat. The GC content of the dummy spacer was 30%, 50%, or 70%. The most non-permissive 50%- and 70%-GC spacers from Figure 1G were used. (D) AT-rich synSeparators improve GFP activation compared with a single G nucleotide. (E, F) This effect is caused by the separator upstream of the GFP-targeting gRNA: Adding the synSeparator only downstream of the GFP-targeting gRNA leads to no improvement. This suggests that the separator acts by improving gRNA processing rather than by generating a 3’ AAAT overhang on the GFP-targeting spacer itself. (G) RNA secondary structure prediction of two of the most disruptive dummy spacers (50% and 70% GC) together with the subsequent GFP-targeting gRNA. (H) This suggests that these spacers form secondary structures that involve the GFP-targeting spacer. Both the natural L. bacterium separator and the AAAT synSeparator are predicted to break up these structures or form protrusions that may facilitate Cas12a access to its cleavage site. (I) SynSeparators derived from other bacterial species (see Figure 2C) can rescue poor GFP activation caused by a non-permissive dummy spacer (70% GC) in a CRISPR array (array design as in A). (J) The enhanced Cas12a protein from A. species (Kleinstiver et al., 2019) is also sensitive to GC content of a dummy spacer (array design as in A) and its performance can be rescued using a TTTT synSeparator derived from its natural separator.
-
Figure 3—source data 1
Raw data used for panel D.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig3-data1-v3.xlsx
-
Figure 3—source data 2
Raw data used for panel F.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig3-data2-v3.xlsx
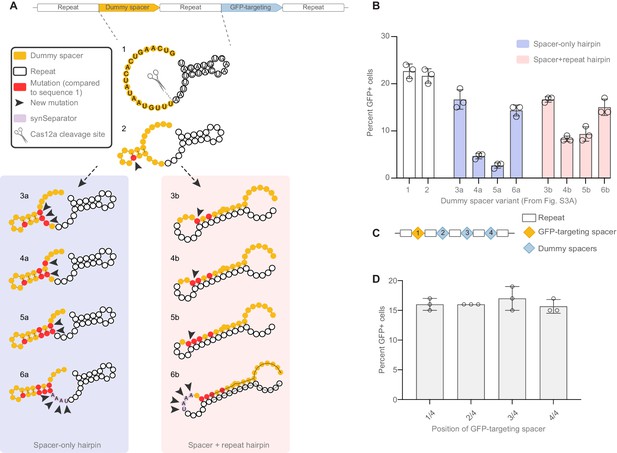
Targeted design of disruptive secondary structure, and no positional effect of crRNAs within a CRISPR array.
(A) We designed short CRISPR arrays where we gradually mutated the dummy spacer in order to create secondary structures that would be localized within the dummy spacer (3a–6a) or would compete with the hairpin normally formed in the GFP crRNA’s repeat region (3b–6b). Each iteration shows targeted mutations intended to make the secondary structure tighter and tighter. Arrowheads highlight the new mutations introduced in every iteration of the array; red highlights all mutations accumulated compared to the dummy spacer in sequence 1 (which has 30% GC). (B) GFP activation decreases as secondary structures get tighter and tighter, indicating that the GFP-targeting crRNA’s performance is sensitive both to secondary structures limited to the dummy spacer, and to hairpins involving the GFP crRNA itself. Adding an AAAT synSeparator rescues array performance in both cases. (C) Four-crRNA arrays were designed where the GFP-targeting crRNA occupies different positions. (D) No positional effect is seen from changing the spacer’s position.
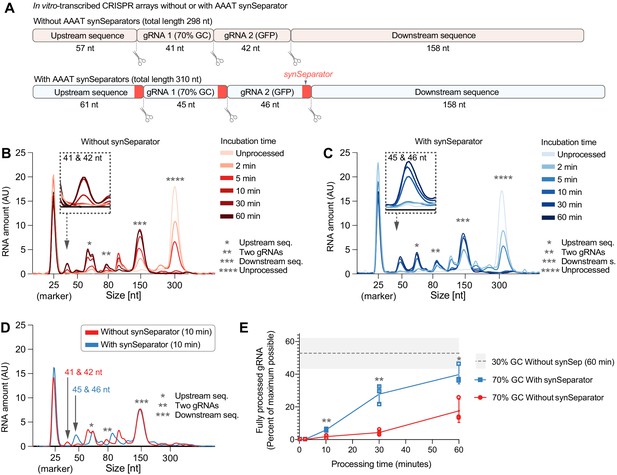
The synSeparator improves CRISPR array processing.
(A) In vitro-transcribed CRISPR arrays containing one of the worst-performing dummy spacers (70% GC), either with or without the AAAT synSeparator upstream of each Cas12a cleavage site. (B, C) Representative Bioanalyzer electropherograms show how both CRISPR arrays become more fully processed with longer Cas12a incubation times (Asterisks highlight specific cleavage products. Note the 25-nt RNA marker, which is not part of the CRISPR array, and how its peak height varies due to sample normalization. See Figure 4—figure supplement 1 for separate plots). However, the synSeparator-containing array is processed more efficiently, seen clearly after a 10 min incubation (D; note that the synSeparator remains attached to the gRNA and slightly increases its length). Bioanalyzer peak data were used to calculate how much of the maximum possible processing had occurred at different time points (E; calculations presented in Materials and methods; n = 3 replicates for each time point). For reference, a gray dotted line shows how much processing had occurred in a CRISPR array containing a 30%-GC dummy spacer and no synSeparator after a 60 min incubation (n = 5 replicates). AU, arbitrary units. Error bars and the gray area in (E) indicate standard deviation.
-
Figure 4—source data 1
Raw data used for panel A.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig4-data1-v3.xlsx
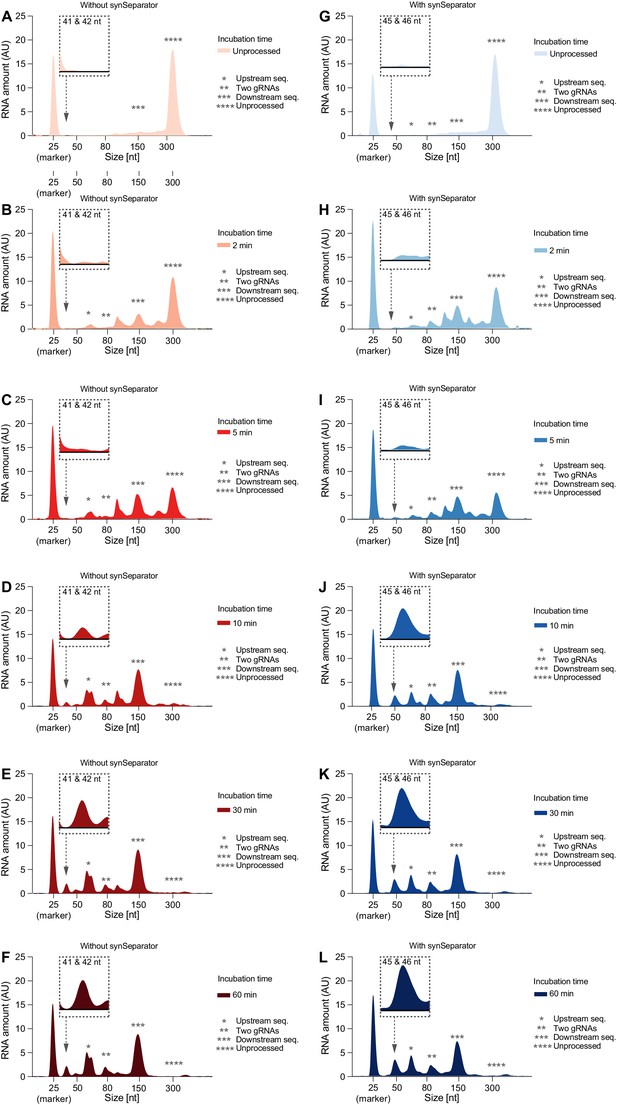
Individual Bioanalyzer electropherograms of Cas12a cleavage time series.
Representative electropherograms of (C) leavage products of the two-gRNA array from Figure 4, either without (A–F) or with (G–L) the AAAT synSeparator. As Cas12a incubation time increases, the array gets more and more fully processed.
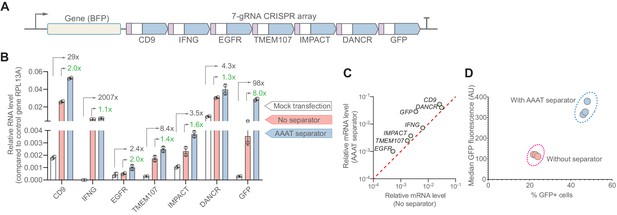
The synthetic separator improves multiplexed gene activation in human cells.
(A) We designed a 7-gRNA array to activate seven endogenous/reporter genes in HEK293T cells with and without the AAAT synSeparator between each gRNA. (B) For all target genes, the synSeparator improves target gene activation level, as measured by RT-qPCR. Arrows indicate the expression fold-change achieved with the synSeparator compared to mock transfection (black values) or the CRISPR array not containing the synSeparator (green values). (C) Plot showing that the improvement of CRISPR activation is consistent across genes. (D) This improvement is also seen on the protein level for the target gene GFP, as measured by GFP fluorescence and percent GFP-positive cells.
-
Figure 5—source data 1
Raw data used for panel B.
- https://cdn.elifesciences.org/articles/66406/elife-66406-fig5-data1-v3.xlsx
Tables
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (Homo sapiens) | HEK Lenti-X 293T (modified) | Takara Bio | 632,180 | Genomically integrated TRE3G-GFP |
| Commercial assay or kit | iScript cDNA Synthesis Kit | Bio-Rad | 1708890 | |
| Commercial assay or kit | RNeasey Plus Mini Kit | Qiagen | 74,134 | |
| Commercial assay or kit | iTaq Universal SYBR Green Supermix | Bio-Rad | 1725120 | |
| Peptide, recombinant protein | T7 DNA Ligase | NEB | M0318S | |
| Peptide, recombinant protein | T4 Polynucleotide Kinase | NEB | M0201L | |
| Commercial assay or kit | NucleoSpin Gel and PCR Clean-up | Takara | 32–740609.50 | |
| Commercial assay or kit | In-Fusion HD Cloning Plus | Takara | 638,920 | |
| Commercial assay or kit | KAPA HiFi Hot-start PCR kit | Roche | 07958897001 | |
| Commercial assay or kit | HiScribe T7 Quick High Yield RNA Synthesis Kit | NEB | E2040S | |
| Commercial assay or kit | MEGAclear Transcription Clean-Up Kit | Thermo Fisher | AM1908 | |
| Peptide, recombinant protein | EnGen Lba Cas12a (Cpf1) - 2000 pmol | NEB | M0653T | |
| Peptide, recombinant protein | Recombinant RNase inhibitor | Takara Bio | 2313 A | |
| Commercial assay or kit | RNA Nano 6,000 chips | Agilent | 5067–1511 | |
| Sequence-based reagent | Low Range ssRNA Ladder | NEB | N0364S | |
| Commercial assay or kit | TransIT-LT1 transfection reagent | Mirus Bio | MIR2304-70 |
Additional files
-
Supplementary file 1
Spacer, repeat and separator sequences used and analyzed in this study.
- https://cdn.elifesciences.org/articles/66406/elife-66406-supp1-v3.xlsx
-
Supplementary file 2
Naturally occurring CRISPR-Cas sequences analyzed in this study.
- https://cdn.elifesciences.org/articles/66406/elife-66406-supp2-v3.xlsx
-
Supplementary file 3
Spacers and qPCR primers used for endogenous gene activation (related to Figure 5).
- https://cdn.elifesciences.org/articles/66406/elife-66406-supp3-v3.xlsx
-
Supplementary file 4
CAGp-FireflyLuciferase-Array-Terminator (Figure 3C).
- https://cdn.elifesciences.org/articles/66406/elife-66406-supp4-v3.docx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/66406/elife-66406-transrepform1-v3.docx





