Neuronal calmodulin levels are controlled by CAMTA transcription factors
Figures
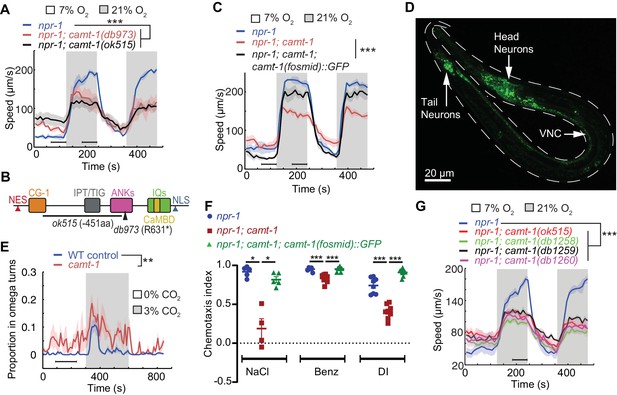
camt-1 mutants exhibit pleiotropic behavioral defects.
(A) camt-1(db973) and camt-1(ok515) mutants (see also (B)) exhibit altered locomotory responses to 21% O2 and hyperactive movement at 7% O2. (B) The domain organization of CAMT-1, highlighting camt-1 loss of function mutations used in this study. (C) A WT copy of the camt-1 genomic locus rescues the O2-response defects of camt-1(db973) mutants. (D) CAMT-1a::GFP driven from its endogenous regulatory sequences in a recombineered fosmid is expressed widely in the nervous system. (E) camt-1(db973) mutants exhibit an increased turning frequency both in the presence and absence of a CO2 stimulus. Assays were performed in 7% O2. (F) camt-1(ok515) mutants show defects in chemotaxis to NaCl, benzaldehyde (Benz), and diacetyl (DI), which can be rescued by expressing a WT copy of CAMT-1. Colored bars indicate the mean and error bars indicate the SEM. (G) The O2-response defects of mutants harboring amino acid substitutions in the CG-1 DNA-binding domain (db1258, db1259, and db1260 alleles; see also Figure 1—figure supplement 1B), are comparable to those of a camt-1(ok515) deletion mutant. (B, C, E, G) Lines indicate average speed and shaded regions SEM, black horizontal bars indicate time points used for statistical tests. (B, C, E–G) Mann-Whitney U-test, ns: p≥0.05, *: p<0.05, **: p<0.01, ***: p<0.001. Number of animals: n≥22 (A), n>41 (C), n≥23 (E), n≥4 assays for each genotype (F), n≥56 (G). ANK, ankyrin domain; CaMBD, calmodulin-binding domain; CG-1, DNA-binding domain; IPT/TIG, Ig-like, plexins, transcription factors or transcription factor immunoglobulin; IQ, calmodulin-binding motif; NES, nuclear export signal; NLS, nuclear localization signal; VNC, ventral nerve cord; WT, wild-type.
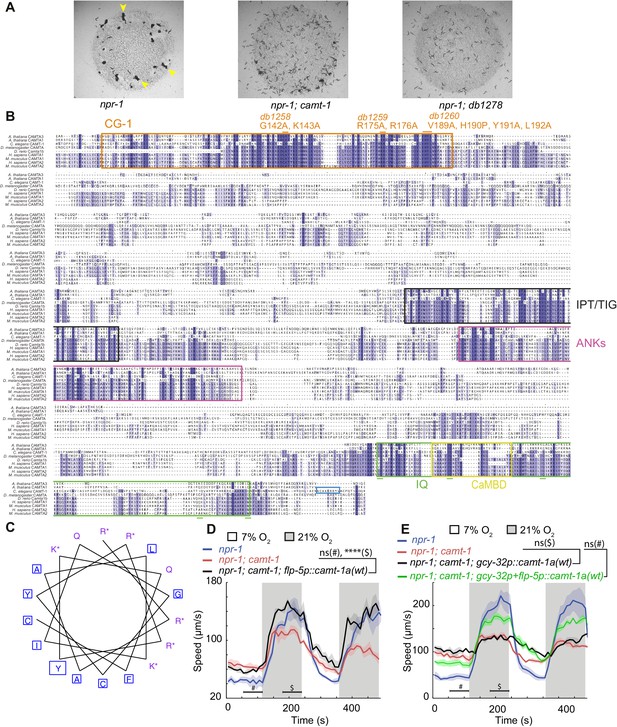
CAMT-1 structure.
(A) Worm clumps (arrowheads) formed by npr-1 null mutants and disruption of aggregation in npr-1; camt-1(ok515) and npr-1; db1278 double-mutants. db1278 allele contains three deletions at the cmd-1 promoter (see also Figure 6). (B) Sequence alignment of CAMTA proteins, highlighting the conserved CG-1 DNA-binding domain (orange), the IPT/TIG domain (black), Ankyrin repeats (purple), putative Ca2+-dependent CaM-binding domain (CaMBD) (yellow), and IQ region (green). CRISPR/Cas9-generated alleles encoding site-specific point mutations in the CG-1 domain are indicated by orange horizontal bars. Green horizontal bars indicate site-specific point mutations of isoleucine residues in IQ motifs to asparagine; these CRISPR/Cas9-generated changes are present in the syb1919 allele and denoted as camt-1(4IQ*). Domain predictions are based on Uniprot and Calmodulin Target Database (http://calcium.uhnres.utoronto.ca/ctdb/ctdb/home.html). (C) Helical wheel projection of CaMBD using Emboss peepwheel tool (http://emboss.bioinformatics.nl/cgi-in/emboss/pepwheel?_pref_hide_optional=0) showing a hydrophobic and a positively charged face. Hydrophobic residues are boxed while positively charged residues are marked with a star. (D, E) Defective responses of camt-1(ok515) mutants to 21% O2 are rescued by expression of CAMT-1a in RMG (using the flp-5 promoter, D) but not in O2-sensing neurons (using the gcy-32 promoter, E). The defective response of camt-1 mutants to 7% O2 are neither rescued by expression in RMG nor by expression in both RMG and O2-sensing neurons (gcy-32p+flp-5p::camt-1a(wt)). # and $ mark the time intervals used for statistical tests at 7% and 21% O2, respectively. For (D, E), Mann-Whitney U-test, ns: p≥0.05, ****: p<0.0001, n≥28 (D), n≥34 animals (E), average speed (line) and SEM (shaded regions) are plotted, black horizontal bars show time points used to test.
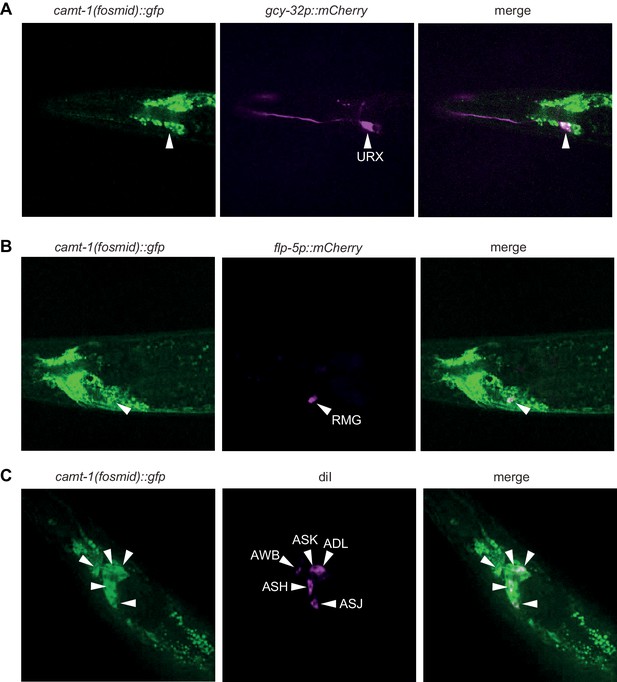
CAMT-1 is widely expressed in the nervous system.
A fosmid-based reporter that tags the CAMT-1a isoform C-terminally with GFP, co-localizes with markers for URX (gcy-32p::mCherry, A), RMG (flp-5p::mCherry, B), and ciliated sensory neurons (DiI, C).
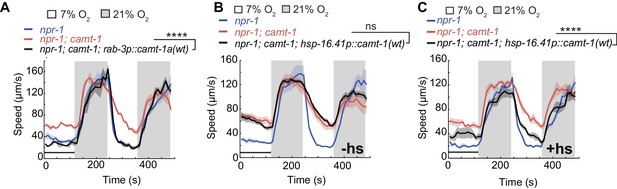
CAMT-1 acts in neurons and is not required developmentally to regulate the oxygen response.
(A) Pan-neuronal expression using the rab-3 promoter of the longest CAMT-1 isoform, CAMT-1a, in camt-1(db973) mutants, rescues O2 response defects. (B, C) Transgenic expression of CAMT-1 from the hsp-16.41 heat-shock promoter does not rescue the hyperactive locomotion of camt-1(ok515) mutants at 7% O2 without heat-shock (B). Heat-shock-induced expression of CAMT-1 in L4 animals rescues this phenotype in camt-1(ok515) mutants, although partially (C). Lines indicate average speed and shaded regions SEM. Black horizontal bars indicate time points used for statistical tests. Mann-Whitney U-test, ns: p≥0.05, ***: p<0.001. Number of animals: n≥39 (A), n≥158 (B), n≥56 (C). hs, heat-shock.
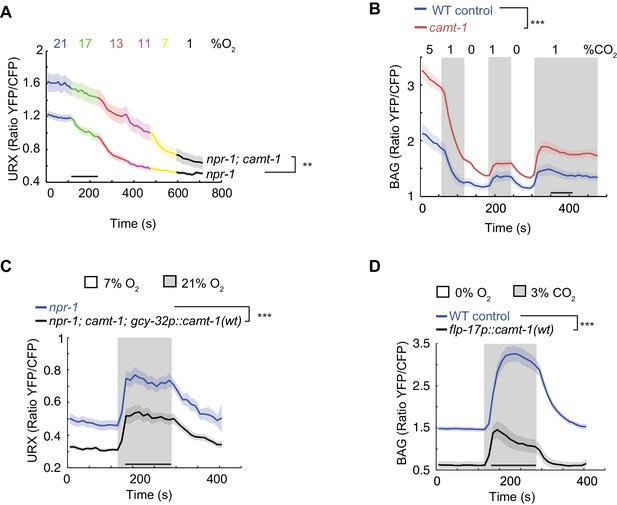
camt-1 mutants show altered Ca2+ traces in sensory neurons.
(A, B) The URX O2-sensing neurons (A) and the BAG CO2 sensors (B) show higher Ca2+ baselines and Ca2+ responses across a range of stimulus intensities in camt-1(db973) mutants. (C–D) Overexpressing wild-type camt-1 cDNA in O2-sensing (using gcy-32p, C) or BAG neurons (using flp-17p, D) strongly reduces Ca2+ levels in these neurons. n≥15 (A), n≥18 (B), n≥17 (C), and n≥20 animals (D). Strains express a Yellow Cameleon sensor in O2-sensing neurons (A, C), or in BAG (B, D) (see Materials and methods). Average YFP/CFP ratios (line) and SEM (shaded regions) are plotted. **: p<0.01, ***: p<0.001, Mann-Whitney U-test.
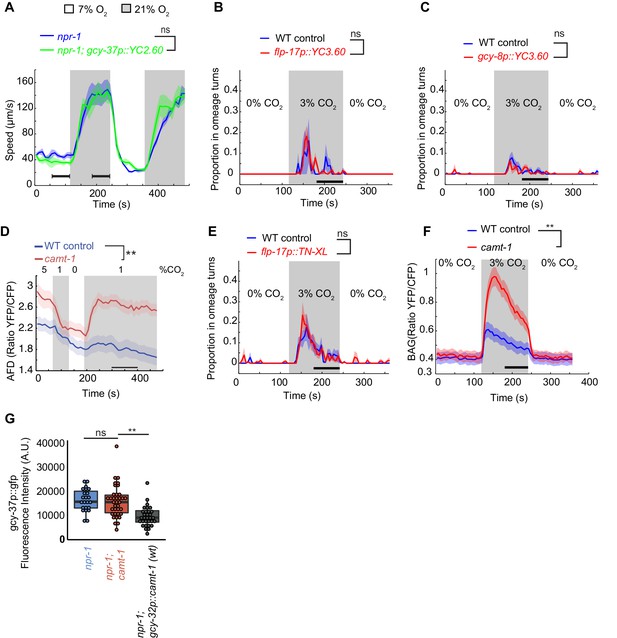
Analyses using Ca2+ imaging lines.
(A–C) Expression of Yellow Cameleon in AQR, PQR, and URX (A), BAG (B), and AFD (C) neurons did not affect the behavioral responses of npr-1 mutants to O2 (A) and of WT control worms to CO2 (B, C). (D) The AFD neuron shows enhanced Ca2+ responses across a range of CO2 stimulus intensities in camt-1(db973) mutants. (E) Expression of the Ca2+ sensor TN-XL in BAG neurons did not affect the response of Caenorhabditis elegans to CO2. (F) The TN-XL Ca2+ sensor, which uses troponin C instead of CaM to detect Ca2+ changes, reports enhanced CO2-evoked Ca2+ responses in BAG neurons in camt-1(ok515) mutants. (G) Expression from the gcy-37 promoter is reduced when CAMT-1 is overexpressed. Quantification of GFP expression driven from the gcy-37 promoter (the same promoter used to drive YC2.60 in the Ca2+ imaging strain) suggests that YC2.60 expression levels are not significantly affected by disrupting camt-1, but are reduced when CAMT-1 is overexpressed. camt-1(db973) mutants were used. Animal responses to CO2 were assayed in 7% O2. n≥34 (A), n≥25 (B), n≥65 (C), n≥15 (D), n≥52 (E), n≥13 (F), and n≥23 animals (G). The strains in (D) express YC3.60 in AFD and in (F) express TN-XL in BAG (see Materials and methods). Plots show average YFP/CFP ratios (line) and SEM (shaded regions). ns: p≥0.05, **: p<0.01, ***: p<0.001, Mann-Whitney U-test (A–F), ANOVA with Tukey’s post hoc HSD (G). WT, wild-type.
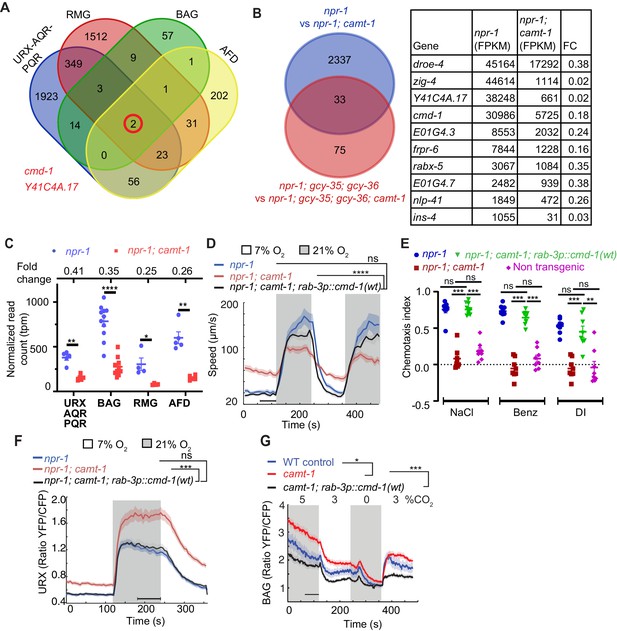
The pleiotropic phenotypes of camt-1 reflect a role in regulating the expression of calmodulin.
(A) Venn diagram showing numbers of genes differentially regulated by CAMT-1 in neuron types we profiled (URX/AQR/PQR, BAG, AFD, and RMG). Two genes, cmd-1 (calmodulin-1) and Y41C4A.17, show consistently altered expression in all neural types profiled. (B) Left: Venn diagram comparing the number of genes differentially regulated by CAMT-1 in URX/AQR/PQR neurons in npr-1 versus npr-1; gcy-35; gcy-36 genetic backgrounds. Right: The most highly expressed genes (read count>1000 FPKM) among the 33 loci regulated by CAMT-1 across all genotypes tested. (C) cmd-1 transcript read counts and FC (top) for URX/AQR/PQR, BAG, AFD, and RMG neurons in camt-1 mutants compared to controls. Each dot or square represents a separate RNA Seq experiment. (D, E) Supplementing CMD-1 expression in neurons using a rab-3p::cmd-1(wt) transgene rescues the O2-response (C) and chemotaxis (D) phenotypes of camt-1 mutants. (F, G) Supplementing CMD-1 expression in neurons also rescues the camt-1 Ca2+-response phenotypes of URX neurons to O2 (F) and of BAG neurons to CO2 (G). Responses to CO2 were assayed in 7% O2. ns: p≥0.05, *: p<0.05, **: p<0.01, ***: p<0.001, ****: p<0.0001, Mann-Whitney U-test (C–H). n≥4 replicates for all cell types (A, B, C), n≥103 (D), n=8 assays for each condition (E), n=32 for each genotype (F), n≥58 animals (G). camt-1 denotes camt-1(ok515). (D, F, G) Lines represent average speed and shaded regions the SEM, black horizontal bars indicate time points used for statistical tests. (C, E) Colored bars indicate the mean and error bars indicate the SEM. FC: fold change.
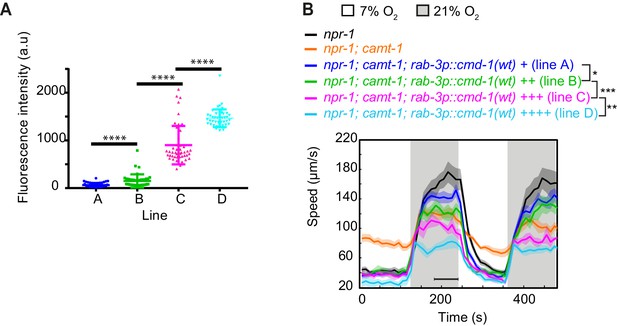
Neuronal calmodulin levels modify behavioral responses to O2.
(A, B) Levels of calmodulin (CMD-1) expression from a series of npr-1; camt-1(ok515) mutant transgenic lines bearing multi-copy arrays of a rab-3p::cmd-1(wt)::SL2::mCherry transgene, quantified using the mCherry fluorescence signal (A). The transgene is an operon that places DNA sequences encoding SL2-mCherry immediately downstream of the CMD-1 stop codon. (B) shows the speed of animals from the lines in (A) at 7% and 21% O2 concentrations. The behavior of npr-1 and npr-1; camt-1(ok515) animals are included for comparison. Note that the speed at 21% O2 is inversely correlated with the CMD-1 expression level (A, B). *: p<0.05, ***: p<0.001, ****: p<0.001, Mann-Whitney U-test.
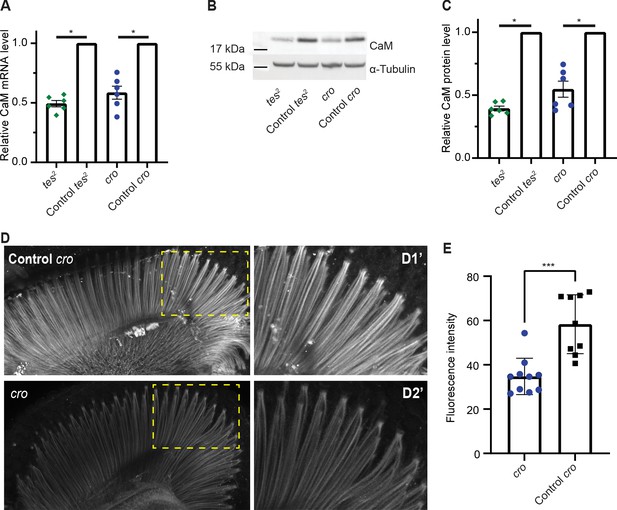
CAMTA regulates CaM expression in Drosophila.
(A) The Drosophila CAMTA mutants tes2 and cro show decreased CaM mRNA levels compared to control flies. mRNA levels in fly heads were measured by quantitative PCR. CAMTA mRNA levels were first normalized to RpL32 (rp49), the qPCR internal control, and then to the value of control flies. (B, C) tes2 and cro mutants show a decrease in CaM protein levels compared to control flies. Protein levels were determined using Western blot of proteins extracted from fly heads. (B) shows a representative picture and (C) shows quantification. CAMTA protein levels were first normalized to alpha-tubulin levels, then to the value of the control flies. (D, E) Immunostaining of fly retinae using CaM antibodies shows reduction of staining of rhabdomeres in cro mutants (see also Figure 5—figure supplement 1B). (D) shows representative pictures of control and cro retinae, respectively, with D1′ and D2′ are blow-ups of yellow rectangle in the left pictures. (E) shows quantification of CaM intensity. (A, C) *: p<0.05, one sample Wilcoxon test to control value of 1, n=6 for each genotype, colored bars indicate the mean and error bars indicate the SEM. (E) ***: p<0.001, Mann-Whitney U-test. w1118; cn1 and w1118; sb are control flies for tes2 (w1118; cn1; tes2) and cro (w1118; cro; sb) mutants, respectively. CaM, calmodulin; CAMTA, CaM-binding transcription activator.
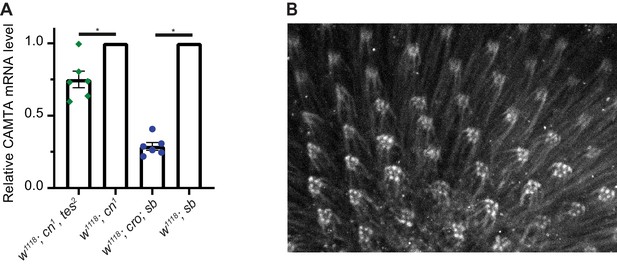
CAMTA and CaM expression in Drosophila.
(A) CAMTA mRNA levels in fly CAMTA mutants. Significant decreases in CAMTA mRNA levels, determined using quantitative PCR and normalized to RpL32 (rp49) mRNA, in tes2 and cro mutant flies, respectively, compared to controls. *: p<0.05, one sample Wilcoxon test to control value of 1. (B) Immunofluorescence staining of control fly retina using CaM antibodies highlights rhabdomeres. CaM, calmodulin; CAMTA, CaM-binding transcription activator.
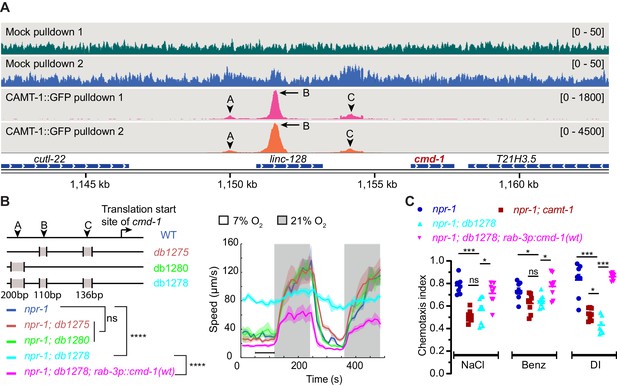
CAMT-1 directly activates calmodulin expression by binding multiple sites in the cmd-1 promoter.
(A) Coverage plots of chromatin pulldown samples showing enrichment at cmd-1 promoter in CAMT-1::GFP pulldown (peaks A, B, and C; arrows: major peaks, arrow heads: minor peaks) compared to a mock pulldown or input (see also Figure 6—figure supplement 1A). Bracketed numbers on the right indicate the scale (normalized read counts). (B) Left: CRISPR-generated strains deleted for one or more of the CAMT-1 ChIP-seq peaks A, B, and C shown in (A); deletions are not drawn to scale. Right: O2-evoked speed responses of the promoter deletion strains shown at left. The db1278 allele in which all three CAMT-1 peaks are deleted confers a strong phenotype that can be rescued by supplementing CMD-1 expression in the nervous system. The db1275 and db1280 alleles, which delete only one or two sites have no obvious phenotype. (C) The db1278 allele confers chemotaxis defects to NaCl, benzaldehyde, and diacetyl, similarly to camt-1(ok515) mutants, that can be rescued by supplementing CMD-1 expression in the nervous system. ns: p≥0.05, *: p<0.05, ***: p<0.001, ****: p<0.0001, Mann-Whitney U-test. n=2 (A), n≥49 (B), n=8 assays for each condition (C). (B) Lines represent average speed and shaded regions the SEM, black horizontal bars indicate time points used for statistical tests. (C) Colored bars indicate the mean and error bars indicate the SEM. ChIP-seq, chromatin immunoprecipitation sequencing.
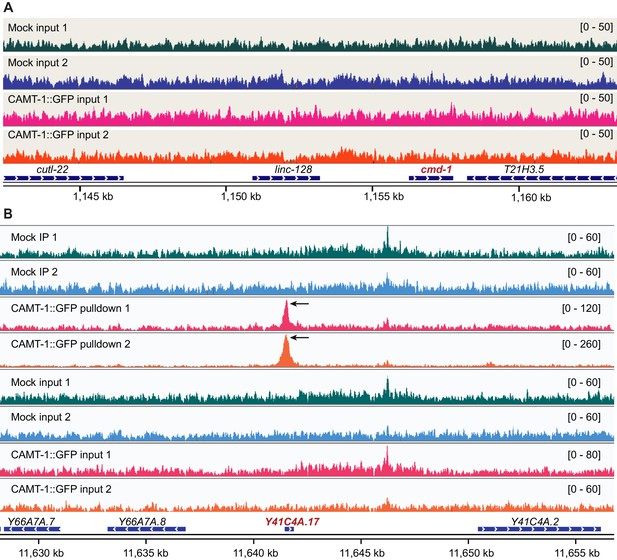
Control samples for ChIP-seq and CAMT-1 binding peak in Y41C4A.17 promotor region.
(A) Coverage plots for input samples corresponding to the ChIP-seq pulldown samples shown in Figure 6A. Numbers on the right indicate the scale (normalized read counts). (B) Coverage plots of pulldown (first four) and input (last four) samples of mock and CAMT-1::GFP showing a peak in the promoter region of the Y41C4A.17 gene in the CAMT-1::GFP pulldown samples (black arrows). Numbers on the right indicate the scale (normalized read counts). ChIP-seq, chromatin immunoprecipitation sequencing.
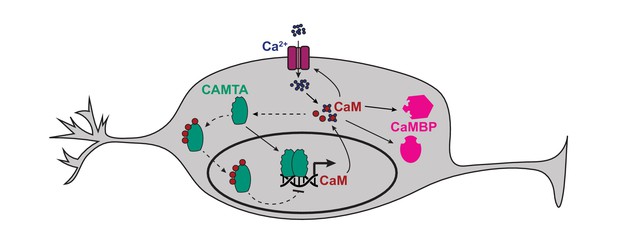
Model of how CAMT-1 may positively and negatively regulate levels of CaM in neurons.
The binding of four apo-CaM to CAMTA is hypothetical, and is based on published data obtained from plant and Drosophila CAMTAs. CaMBP: Other CaM-binding proteins. Further analysis is required to confirm if the negative feedback loop occurs at physiological CaM concentrations. CaM, calmodulin; CAMTA, CaM-binding transcription activator.
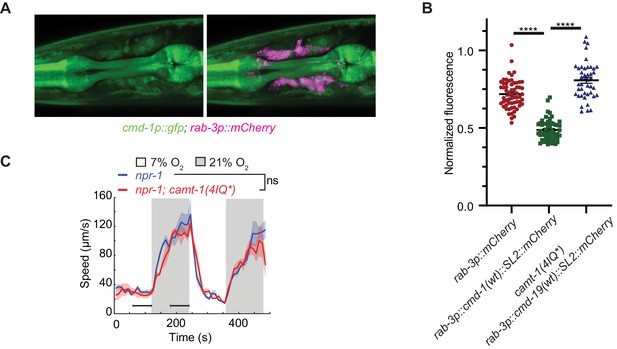
CAMT-1 can repress CMD-1/CaM expression at high CMD-1/CaM levels.
(A) A transcriptional reporter of CMD-1 (cmd-1p::gfp) shows expression in neurons, pharyngeal, and body wall muscles. Neurons are co-labeled with mCherry driven from rab-3p. (B) Overexpression of CMD-1 in neurons (rab-3p::cmd-1(wt)) inhibits the expression of a cmd-1 transcriptional reporter (shown in A). This inhibition can be abrogated by introducing mutations in the four IQ domains of CAMT-1 (as shown in Figure 1—figure supplement 1B). These lines express mCherry in neurons either directly from rab-3p or in an operon with CMD-1 (rab-3p::cmd-1::SL2::mCherry) and all have npr-1 background mutation. Y-axis shows neuronal fluorescence after normalization to pharyngeal fluorescence. (C) Binding of CAMT-1 to CaM via IQ motifs is not essential for oxygen escape behavior. Mutants with four IQ motifs mutated exhibit similar locomotory response to O2 as npr-1. ns: p≥0.05, ****: p<0.0001, Mann-Whitney U-test, n≥43 (B), n≥37 (C). Colored bars indicate the mean and error bars indicate the SEM (B). Plots show average speed (line) and SEM (shaded regions), black horizontal bars show time points used for statistical tests (C). CaM, calmodulin.
Additional files
-
Supplementary file 1
The 100 most highly expressed genes (in order of decreasing read counts, in TPM) from neuron-specific RNA profiling.
- https://cdn.elifesciences.org/articles/68238/elife-68238-supp1-v3.xlsx
-
Supplementary file 2
Genes showing differential expression between camt-1 and WT in the profiled neural types.
- https://cdn.elifesciences.org/articles/68238/elife-68238-supp2-v3.xlsx
-
Supplementary file 3
Genomic locations differentially bound by CAMT-1 identified using the DiffBind algorithm for ChIP-seq data with a false discovery rate (FDR) threshold of 0.05.
Genes overlapping or within 10 kb downstream of these sites are reported. The table is sorted in the order of increasing FDR. Note that the CAMT-1 binding site at the cmd-1 promoter was annotated with the overlapping long intervening noncoding RNA linc-128.
- https://cdn.elifesciences.org/articles/68238/elife-68238-supp3-v3.xlsx
-
Supplementary file 4
List of C. elegans strains used in this study.
- https://cdn.elifesciences.org/articles/68238/elife-68238-supp4-v3.xlsx
-
Supplementary file 5
Exact p-values and n numbers for experiments reported in this study.
- https://cdn.elifesciences.org/articles/68238/elife-68238-supp5-v3.xlsx
-
Transparent reporting form
- https://cdn.elifesciences.org/articles/68238/elife-68238-transrepform-v3.docx





