µ-Theraphotoxin Pn3a inhibition of CaV3.3 channels reveals a novel isoform-selective drug binding site
Abstract
Low voltage-activated calcium currents are mediated by T-type calcium channels CaV3.1, CaV3.2, and CaV3.3, which modulate a variety of physiological processes including sleep, cardiac pace-making, pain, and epilepsy. CaV3 isoforms’ biophysical properties, overlapping expression, and lack of subtype-selective pharmacology hinder the determination of their specific physiological roles in health and disease. We have identified μ-theraphotoxin Pn3a as the first subtype-selective spider venom peptide inhibitor of CaV3.3, with >100-fold lower potency against the other T-type isoforms. Pn3a modifies CaV3.3 gating through a depolarizing shift in the voltage dependence of activation thus decreasing CaV3.3-mediated currents in the normal range of activation potentials. Paddle chimeras of KV1.7 channels bearing voltage sensor sequences from all four CaV3.3 domains revealed preferential binding of Pn3a to the S3-S4 region of domain II (CaV3.3DII). This novel T-type channel pharmacological site was explored through computational docking simulations of Pn3a, site-directed mutagenesis, and full domain II swaps between CaV3 channels highlighting it as a subtype-specific pharmacophore. This research expands our understanding of T-type calcium channel pharmacology and supports the suitability of Pn3a as a molecular tool in the study of the physiological roles of CaV3.3 channels.
Editor's evaluation
Low voltage activated T-type calcium channels (CaV3.1-CaV3.3) are important to several physiological processes, but the functions of individual isoforms are difficult to distinguish pharmacologically. This study reports a spider toxin, Pn3a, which exhibits 100-fold selectivity for inhibiting CaV3.3 over CaV3.1 and CaV3.2 isoforms. This specificity is conferred by a specific interaction site on the Cav3.3 domain II voltage sensor.
https://doi.org/10.7554/eLife.74040.sa0Introduction
Voltage-gated calcium (CaV) channels are activated by membrane depolarization and are involved in many physiological processes including contraction, secretion, neurotransmitter release, and gene expression (Catterall et al., 2005). In contrast to high voltage-activated (HVA) Ca2+ currents mediated by L-, N-, P/Q-, and R-type calcium channels that require large depolarizations, low voltage-activated (LVA) currents mediated by T-type Ca2+ channels are activated by small membrane depolarization and display distinctively faster activation and inactivation kinetics. T-type channel activation near resting membrane potentials generates low-threshold Ca2+ spikes responsible for the conspicuous burst firing and low-frequency oscillatory discharges observed in thalamic, olivary, and cerebellar neurons (Park et al., 2010; Dreyfus et al., 2010; Molineux et al., 2006; Lee et al., 2014). Thus, the biophysical properties of the T-type channels make them important regulators of cardiac and neuronal excitability (Perez-Reyes, 2003; Huguenard, 1996; Coulter et al., 1989) and therefore key pharmacological targets for the treatment of neurological and psychiatric disorders (Zamponi, 2016; Maksemous et al., 2022).
T-type calcium channels display small single-channel conductance (T stands for transient or tiny) (Perez-Reyes et al., 1998) and are encoded by the CACNA1G (CaV3.1), CACNA1H (CaV3.2), and CACNA1I (CaV3.3) genes. The CaV3s have similar channel activation and inactivation kinetics and are ubiquitously expressed in the nervous, neuroendocrine, reproductive, and cardiovascular systems (Perez-Reyes, 2003; Hansen, 2015). In heterologous expression systems, CaV3.3-mediated currents display the slowest activation and inactivation kinetics and recover faster from inactivation (Kozlov et al., 1999), however, due to these relatively small differences, they are nearly indistinguishable from the other T-type isoform in native tissue. In the brain, abundant CACNA1I transcripts display remarkable regional distribution and appear to prevail in distal dendrites (Perez-Reyes, 2003; Lee et al., 1999), where CaV3.3 currents mediate the major sleep-spindle pacemaker in the thalamus (Astori et al., 2011). Transgenic mice lacking CaV3.3 channels display severe alterations to sleep-spindle generator rhythmogenic properties posing this T-type channel isoform as a critical target in the study of brain function and development (Astori et al., 2011). However, the expression of CaV3.3 channels overlaps with that of CaV3.1 and/or CaV3.2, which together with the lack of robust, isoform-selective pharmacology has hampered the elucidation of their specific contributions to cellular physiology.
T-type calcium channels share the characteristic modular topology of other voltage-gated ion channels (VGICs) (Catterall et al., 2005) that consists of a voltage sensor (VS) module formed by transmembrane segments S1 through S4 and a pore module (PM) composed of the transmembrane segments S5 and S6 connected by a re-entrant pore loop. The VS controls channel opening in response to changes in membrane potential and the PM provides aqueous passage for ions across the lipid membrane. The tetrameric arrangement of four PMs lining the permeation pathway surrounded by four VSs in either swapped or non-swapped configuration enables VGIC function (for review, see Barros et al., 2019). In voltage-gated potassium (KV) and transient receptor potential (TRP) channels, each monomer (1 × VS + 1 × PM) is encoded by a core α-subunit; whereas in voltage-gated sodium (NaV) and CaV channels, the α-subunit contains the four homologous, but not identical, domains (DI-DIV) joined through large intracellular linkers (Catterall, 2000; Catterall et al., 2005).
Natural compounds that evolved to occlude ion channel’s PM or to interact with their VS are distinguished broadly as pore blockers and gating modifiers, respectively. In VGICs, the extracellularly exposed areas of the VS are pharmacological targets of neurotoxins and synthetic compounds where at least three distinct pharmacological sites have been described in NaV channels (Catterall et al., 2007). Within the VS is a conserved S3b-S4 paddle motif which can be transplanted into the VS of other VGICs and retain toxin sensitivity (Bosmans et al., 2008). α-Scorpion toxins, sea-anemone toxins, and numerous spider toxins interact with NaV channel site 3, located in the extracellular loop between DIV-S3 and DIV-S4 thereby interfering with the conformational changes that couple channel activation to fast inactivation (Hanck and Sheets, 2007). Binding of β-scorpion toxins to the S1-S2 and S3-S4 loops of domain II (site 4) shifts the voltage dependence of channel activation towards depolarized voltages reducing the maximal current at normal activation potentials. Lastly, pharmacological site 6 (located near site 3) is targeted by the δ-conotoxins that slow NaV channel inactivation (Terlau et al., 1996). All these gating modifier peptides (GMPs) appear to ‘hold’ the VS in different conformations leading to their mechanism of modulation to be recognized as ‘VS trapping’ (Cestèle et al., 1998), a phenomenon also observed in the interaction of theraphotoxins with KV2.1 (Swartz and MacKinnon, 1997) and agatoxins with CaV channels (McDonough et al., 1997a).
Sequence and functional conservation between the VSs lead to promiscuous interactions between peptides and small molecules across VGIC families. Examples of these include ProTx-I (Thrixopelma pruriens, NaV/KV/CaV/TRPA1) (Bladen et al., 2014; Bosmans et al., 2008; Gui et al., 2014; Middleton et al., 2002); ProTx-II (T. pruriens, NaV/CaV) (Bladen et al., 2014; Middleton et al., 2002); Kurtoxin (Parabuthus transvaalicus, NaV/CaV) (Chuang et al., 1998); Hanatoxin (Grammostola spatulata, KV/NaV/CaV) (Bosmans et al., 2008; Li-Smerin and Swartz, 1998; Swartz and MacKinnon, 1997), and Pm1a (Pelinobius muticus, NaV/KV) (Finol-Urdaneta et al., 2022). Furthermore, small molecules such as capsaicin, capsazepine (TRPV1/KV/CaV) (Caterina et al., 1997; Kuenzi and Dale, 1996; McArthur et al., 2019), and A803467 (NaV, CaV) (Bladen and Zamponi, 2012) amongst others are known to interact across VGIC families.
Shared ancestry and sequence conservation within the voltage sensing machinery have been used to rationalize commonalities in structure, gating kinetics, and pharmacophores between NaV and T-type channels (Bladen and Zamponi, 2012). Several NaV-active GMPs were shown to inhibit CaV3.1 channels through interactions with the channel’s domain III (CaV3.1DIII) (Salari et al., 2016); whereas the potent NaV1.7 inhibitor, µ-theraphotoxin Pn3a (Pamphobeteus nigricolour) (Deuis et al., 2017), also interacts with HVA CaV channels (McArthur et al., 2020).
In this study, we have used whole-cell patch clamp electrophysiology, mutagenesis, and computational docking to probe Pn3a’s interactions with the LVA CaV3 channels. Our results support the use of Pn3a as a molecular tool for the study of CaV3.3-mediated currents in native cells and highlight a previously unrecognized pharmacophore that may enable selective targeting of T-type channel isoforms.
Results
Pn3a selectively inhibits CaV3.3 channels
Functional assessment of Pn3a activity was examined on depolarization-activated calcium currents (ICa) through the human T-type calcium channel isoforms: CaV3.1, CaV3.2, and CaV3.3. Whole-cell currents were elicited by a 100 ms test pulse to −20 mV from a holding potential (Vh) of −90 mV at a frequency of 0.2 Hz and recorded at room temperature (20–22°C, Figure 1A). Pn3a (10 µM) strongly inhibited CaV3.3-mediated currents (90.2% ± 1.9%, n=5) with negligible effects over the two other T-type isoforms (CaV3.1: 3.8% ± 1.8%, n=5; CaV3.2: 2.5% ± 2.5%, n=5) (Figure 1A and B). These results indicate that Pn3a has >100-fold preference for CaV3.3 over the other highly homologous CaV3 isoforms. Scaled CaV3.3 macroscopic currents recorded in the absence (control) and presence of Pn3a (3 µM) display similar macroscopic activation kinetics (τact, control = 5.95 ± 0.54 ms vs. τact, Pn3a = 6.37 ± 0.44 ms; p=0.32, n=5, paired t-test), whereas Pn3a slowed macroscopic inactivation kinetics (τinact control = 39.91 ± 8.25 ms vs. τinact, Pn3a = 60.25 ± 16.02 ms; p=0.04, n=5, paired t-test) (representative currents provided in the inset, Figure 1A). The preferential actions of Pn3a highlight its potential as a molecular probe to isolate and study the contribution of CaV3.3 currents in native cells.
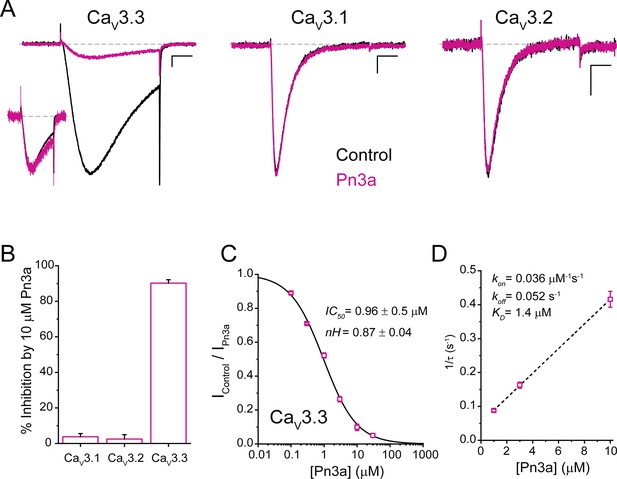
Pn3a preferentially inhibits human CaV3.3-mediated Ca2+ currents.
(A) CaV3.3 (left), CaV3.1 (middle), and CaV3.2 (right) currents elicited by 100 ms step depolarization to −20 mV (Vh −90 mV, 0.2 Hz) in the absence (control, black) and presence of 3 µM Pn3a (pink). Scale bars: 0.2 nA, 20 ms. The inset shows scaled CaV3.3 currents in control and in the presence of Pn3a with similar kinetics. (B) Bar graph summarizing % inhibition by 10 µM Pn3a of the three CaV3 isoforms. (C) Concentration-response curve for Pn3a inhibition of CaV3.3 currents. (D) Kinetics of Pn3a inhibition of CaV3.3. Kobs was determined at three concentrations and fit with a linear equation where Kobs = kon ∙ [Pn3a]+koff. Data shown as mean ± SEM (n=5).
-
Figure 1—source data 1
Pn3a inhibition of T-type calcium channels.
- https://cdn.elifesciences.org/articles/74040/elife-74040-fig1-data1-v2.xlsx
Pn3a inhibitory potency against CaV3.3 currents was assessed at increasing peptide concentrations from which a concentration-response curve was built (Figure 1C). Fit to a standard Hill equation rendered an IC50 value of 0.96±0.05 µM (nH 0.87±0.04, n=5 per concentration) for the inhibition of CaV3.3 channels. This Hill coefficient is consistent with a 1:1 stoichiometry between Pn3a toxin and CaV3.3 channels.
The change in CaV3.3 peak current amplitude during Pn3a washin and washout enables the assessment of Pn3a binding to CaV3.3 channels. The inhibition of CaV3.3 ICa by Pn3a was mono-exponential with progressively faster time constants (τobs) at increasing peptide concentration (Figure 1D). The on- and off-rate constants (kon and koff) were determined from the fit to the linear plot of 1/τobs vs. [Pn3a] (Figure 1D, n=5 per concentration), where kon is the slope and koff is the y-intercept. The linear regression line results in a kon of 0.036 µM–1 s–1 and koff of 0.052 s–1 which yield a KD of 1.4 µM, in close agreement with the IC50 value obtained (Figure 1C). koff was confirmed by fitting the washout to a single exponential (koff = 0.055 ± 0.002 s–1). Hence, Pn3a inhibits CaV3.3-mediated currents without apparent actions on CaV3.1 or CaV3.2 exposed to up to 10 µM peptide.
Pn3a modifies the gating of CaV3.3
The voltage dependence of CaV3.3 activation, deactivation, inactivation, and recovery from inactivation were investigated in the absence and presence of Pn3a (3 µM) (Figure 2 and Table 1). CaV3.3 activation is shifted ~13 mV to more depolarized potentials (control V0.5 = −28.3±0.4 mV, n=5, vs. Pn3a V0.5 = −15.3±0.4 mV, n=5; p<0.0001), indicating that a stronger depolarization is required to enable channel opening in the presence of Pn3a (Figure 2A and C). The voltage dependence of CaV3.3 steady-state inactivation (SSI) was not affected by exposure to the spider peptide (control V0.5 = −56.3±0.3 mV, n=5, vs. Pn3a V0.5 = −56.6±0.4 mV, n=5, p=0.57, Figure 2B–C). Consistent with the observed slowing of inactivation, CaV3.3-mediated currents inactivated by a 200 ms pre-pulse to –20 mV recovered faster in the presence of Pn3a (τ=0.22 ± 0.01 s, n=5) than under control conditions (τ=0.34 ± 0.01 s, n=5; p<0.0001) (Figure 2D–E), suggesting a potential Pn3a-dependent destabilization of the inactivated state.
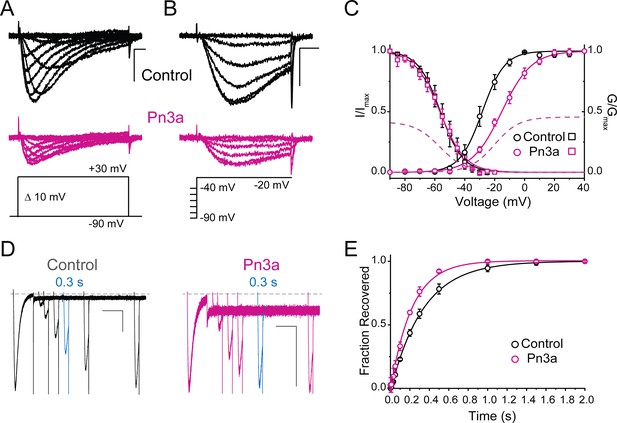
Pn3a produces a depolarizing shift in the voltage dependence of activation and speeds up recovery from inactivation of CaV3.3.
(A–C) Effect of Pn3a on the voltage dependence of CaV3.3. Representative currents from (A) activation or (B) steady-state inactivation protocols, in the absence (top: control, black) and presence of 3 µM Pn3a (middle: Pn3a, pink) using the standard protocols (bottom). Scale bars: 0.5 nA, 10 ms. (C) Activation (circles) and steady-state inactivation (squares) relationships for CaV3.3 in the absence (black) and presence of 3 µM Pn3a (pink). Dashed lines represent non-normalized activation and inactivation curves of Pn3a. (D) Representative recovery from inactivation currents in control (left) and presence of 3 µM Pn3a (right) (trace shown in blue highlights the current recovered after 0.3 s). Scale bars: 0.2 nA, 200 ms. (E) Recovery from inactivation in the absence (black) and presence of 3 µM Pn3a (pink). Data shown as mean ± SEM (n=5).
-
Figure 2—source data 1
Pn3a voltage effects o activation, steady-state inactivation, and recovery from inactivation.
- https://cdn.elifesciences.org/articles/74040/elife-74040-fig2-data1-v2.xlsx
Activation and inactivation values of CaV3.3 and Kv1.7-CaV3.3DI-IV chimers in the absence (control) and presence of Pn3a (3 µM for CaV3.3; 1 µM for KV1.7-CaV3.3D1-IV chimeras).
| Control Pn3a | |||
|---|---|---|---|
| hCaV3.3 | V0.5 activation | −28.3±0.4 mV (5) | −15.3±0.4 mV (5) * |
| ka Slope factor | 7.5±0.3 (5) | 10.0±0.3 (5) * | |
| V0.5 SSI | −56.3±0.3 mV (5) | −56.6±0.4 mV (5) | |
| ka Slope factor | 7.5±0.2 (5) | 8.4±0.4 (5) | |
| τ Recovery | 0.34±0.01ms (5) | 0.22±0.01 (5) * | |
| KV1.7 | V0.5 activation | −10.5±0.6 mV (7) | - |
| ka Slope factor | 9.9±0.5 (7) | - | |
| KV1.7-CaV3.3DI | V0.5 activation | 91.3±0.4 mV (5) | 88.2±0.3 mV (5) |
| ka Slope factor | 1.0±0.3 (5) | 11.8±0.2 (5) | |
| KV1.7CaV3.3DII | V0.5 activation | 97.3±0.4 mV (9) | 112.6±0.4 mV (7) * |
| ka Slope factor | 14.1±0.3 (9) | 12.3±0.4 (7) * | |
| KV1.7-CaV3.3DIII | V0.5 activation | 71.3±0.8 mV (7) | 74.7±0.9 mV (5) |
| ka Slope factor | 21.1±0.7 (7) | 22.0±0.8 (5) | |
| KV1.7-CaV3.3DIV | V0.5 activation | −60.8±0.8 mV (5) | −60.8±0.4 mV (5) |
| ka Slope factor | 10.1±0.7 (5) | 10.0±0.4 (5) | |
-
*
Significant determined from paired t-test with significance threshold set to p<0.05.
The voltage dependence of Pn3a inhibition of ICa and Na+-mediated (INa) CaV3.3 currents was examined at various test potentials (Figure 3A and Figure 3—figure supplement 1). Pn3A displayed a decrease in CaV3.3 ICa inhibition at more depolarized potentials consistent with Pn3a preferentially binding to the down-state of the VS. To examine the voltage dependence across a larger voltage range, extracellular Ca2+ was removed, permitting Na+ to be the primary charge carrier through the open CaV3.3. Similar to that seen for ICa, INa showed a similar voltage dependence, with stronger inhibition at more negative potentials (Figure 3—figure supplement 1).
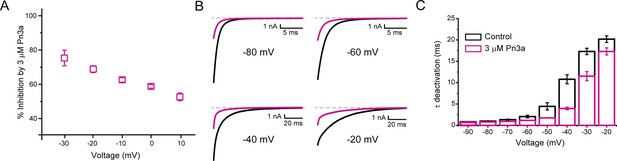
Pn3a inhibition of CaV3.3 is stronger at hyperpolarized potentials and speeds up channel deactivation.
(A) Voltage dependence of Pn3a inhibition of CaV3.3-mediated Ca2+ currents. (B) Representative CaV3.3 tail currents at different voltages in the absence (black) and presence (pink) of 3 µM Pn3a. (C) Summary of the time constant (τ) of CaV3.3 deactivation upon return to the holding potential (−90 mV) plotted against the activating (pre-pulse) potential in the absence (control, black) and the presence of Pn3a (3 µM, pink). Data shown as mean ± SEM (n=5).
-
Figure 3—source data 1
Voltage dependence of Pn3a inhibition.
- https://cdn.elifesciences.org/articles/74040/elife-74040-fig3-data1-v2.xlsx
The time course of tail current decay reflects the rate of channels leaving the open state (test potential) and entering the closed state (deactivation) upon return to the holding potential. We examined the modulation of CaV3.3 channel deactivation through tail current kinetic analysis. Pn3a-modified CaV3.3 currents displayed a faster deactivation time constant (τdeactivation) across all potentials tested compared to control (Figure 3B–C). This data suggests that Pn3a binding may destabilize the open state or stabilize the closed one. This results that Pn3a is a CaV3.3 GMP that decreases channel availability by increasing the energy required for channel opening.
Pn3a interacts with CaV3.3DII S3-S4 paddles
To ascertain Pn3a’s pharmacophore on CaV3.3 channels, we applied the chimeric approach of transplanting its four S3-S4 paddles into a KV channel based on a similar template to that previously described for KV2.1/CaV3.1 chimeras (Salari et al., 2016). We substituted portions of the S3b-S4 from CaV3.3 DI to DIV into the KV1.7 channel backbone. The sequence alignment of CaV3.3 DI-DIV S3-S4 segments and the corresponding extracellular region of KV1.7 is presented in Figure 4A.
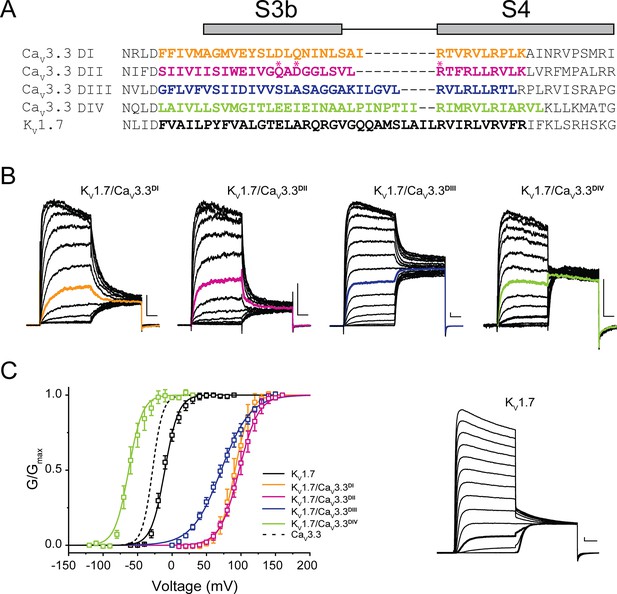
Chimeric constructs of the KV1.7 channel with the CaV3.3 voltage sensor paddles.
(A) Sequence alignment between paddle regions of CaV3.3 and KV1.7. The coloured bolded sequences from CaV3.3 DI (yellow), CaV3.3 DII (pink), CaV3.3 DIII (blue), and CaV3.3 DIV (green) were grafted onto KV1.7 (black). (B) Representative current traces in response to 50 ms long I-V protocols used to evaluate the voltage dependence of activation of all constructs (Vh = −80 mV). KV1.7/CaV3.3DI (0–160 mV), KV1.7/CaV3.3DII (0–160 mV), KV1.7/CaV3.3DIII (0–160 mV) and KV1.7/CaV3.3DIV (−120 to 30 mV). The traces highlighted in colour correspond to currents near half-activation potential (V0.5). (C) Activation curves for all chimeras: KV1.7/CaV3.3DI (yellow), KV1.7/CaV3.3DII (pink), KV1.7/CaV3.3DIII (blue), and KV1.7/CaV3.3DIV (green) and the parental channel KV1.7 (black). The dotted line corresponds to CaV3.3 activation for reference. The inset contains representative KV1.7 currents (−60 to 90 mV). All scale bars: 1 nA, 10 ms.
All four KV1.7/CaV3.3 chimeric constructs were functional, mediating large K+ currents that displayed distinct gating properties from those of the parental wild-type KV1.7 (V0.5 = −10.5±0.6 mV, n=7; Figure 4B, inset). Briefly, chimeric KV1.7/CaV3.3DI, KV1.7/CaV3.3DII and KV1.7/CaV3.3DIII displayed 80–100 mV depolarizing shifts in channel activation (KV1.7/CaV3.3DI V0.5=91.3 ± 0.4 mV, n=5; KV1.7/CaV3.3DII V0.5=97.3 ± 0.4 mV, n=9; KV1.7/CaV3.3DIII V0.5=71.3 ± 0.8 mV, n=7, two-way ANOVA, p<0.05), whereas KV1.7/CaV3.3DIV activation was ~50 mV hyperpolarized (KV1.7/CaV3.3DIV V0.5 = −60.8±0.8 mV, n=5; two-way ANOVA, p<0.05) (Figure 4). Our results are in broad agreement with previous reports for CaV3.1, where DIV displays a hyperpolarizing shift and DI-III show a variable depolarizing shift in V0.5 (Salari et al., 2016).
KV1.7/CaV3.3DI-IV chimeras and the parental KV1.7 construct were analysed in control and after exposure to Pn3a (1 µM) and representative currents (Vh = −100 mV, with test pulse to +40 mV (KV1.7), +120 mV (KV1.7/CaV3.3DI-III), or 0 mV (KV1.7/CaV3.3DIV)) in both conditions are included as insets (Figure 5). Similar to the parental channel, peak currents of KV1.7/CaV3.3DI and KV1.7/CaV3.3DIV chimaeras were insensitive to Pn3a (KV1.7: 1.0% ± 0.8%, n=6; CaV3.3DI: 0.6% ± 0.5%, n=5, and CaV3.3DIV: 0.4% ± 0.2%, n=5) (Figure 5A, D and E), with KV1.7/CaV3.3DIII displaying modest peptide-dependent inhibition (12.0% ± 0.6%, n=6) (Figure 5C).
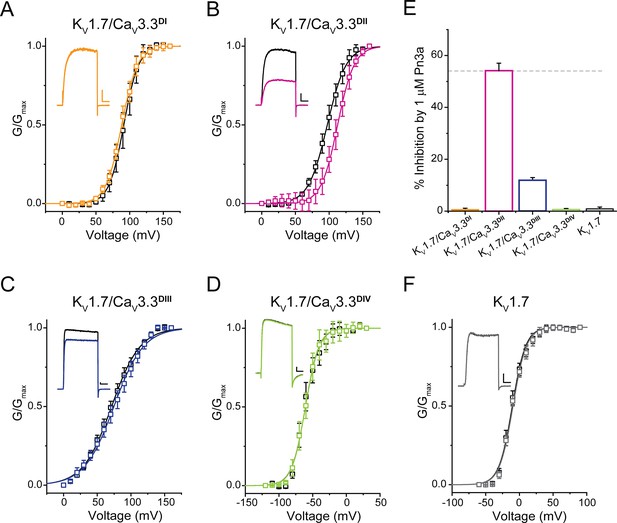
Pn3a shifts the activation of KV1.7/CaV3.3DII chimeric channels.
G/Gmax-V relationships and representative currents obtained in the absence (control, black) and presence of 1 µM Pn3a (pink) for (A) KV1.7/CaV3.3DI (90 mV), (B) KV1.7/CaV3.3DII (100 mV), (C) KV1.7/CaV3.3DIII (70 mV), and (D) KV1.7/CaV3.3DIV (−60 mV). (E) Bar graph showing percent inhibition by 1 µM Pn3a of KV1.7/CaV3.3DI-IV chimeras and KV1.7 wt. (F) KV1.7 G/Gmax-V relationship and current traces obtained in the absence (control) and presence of Pn3a (–10 mV). All scale bars: 1 nA, 10 ms.
-
Figure 5—source data 1
Voltage dependence of Pn3a inhibition of Kv1.7/Cav3.3 Chimera.
- https://cdn.elifesciences.org/articles/74040/elife-74040-fig5-data1-v2.xlsx
In contrast, currents mediated by KV1.7/CaV3.3DII were significantly reduced (54.2% ± 2.9%, n=7) in the presence of Pn3a (Figure 5B). Furthermore, conductance-voltage relationships for each S3-S4 paddle chimera in the absence and presence of Pn3a were built from peak currents in response to standard I-V protocols (KV1.7 –60 to +90 mV; KV1.7/CaV3.3DI-III 0 to +160 mV; and KV1.7/CaV3.3DIV −120 to +30 mV; Vh = −100 mV, 0.2 Hz). The latter analysis revealed an ~15 mV rightward shift in the half voltage of activation (V0.5) exclusively in the CaV3.3DII chimera when exposed to Pn3a (control V0.5=97.3 ± 0.4 mV, n=9; vs. Pn3a V0.5=112.6 ± 0.4 mV, n=7, p<0.0001), strongly suggesting that Pn3a interacts with the S3-S4 region of domain II in CaV3.3. These results suggest that the positive shift in activation V0.5 observed in the full length and DII chimera likely underpins Pn3a’s mechanism of CaV3.3 inhibition, highlighting the differences in pharmacological properties of each voltage-sensing module.
Molecular determinants of Pn3a inhibition of CaV3.3
The critical role of DII was further supported by examining the effects of Pn3a on full-length CaV3 channel constructs where the whole DII was swapped between isoforms. Replacement of CaV3.3DII with either CaV3.1DII or CaV3.2DII resulted in partial (CaV3.3/CaV3.1DII: 33% ± 4.1%, n=5, one-way ANOVA p<0.0001) or complete (CaV3.3/CaV3.2DII: 2.2% ± 1.5%, n=5, one-way ANOVA p<0.0001) loss of inhibition compared to the full-length CaV3.3 (90.2 ± 1.9% n=5) in the presence of 10 μM Pn3a (Figure 6E). The reverse constructs where CaV3.3DII was grafted onto CaV3.1 (CaV3.1/CaV3.3DII) or CaV3.2 (CaV3.2/CaV3.3DII) afforded Pn3a sensitivity to the parental CaV3 channel such that currents mediated by CaV3.1/CaV3.3DII were inhibited by 71.4% ± 4.1% (n=5), and CaV3.2/CaV3.3DII by 68.0% ± 3.6% (n=5) (Figure 6F). Thus, verifying that CaV3.3DII is required for Pn3a interaction with T-type calcium channels.
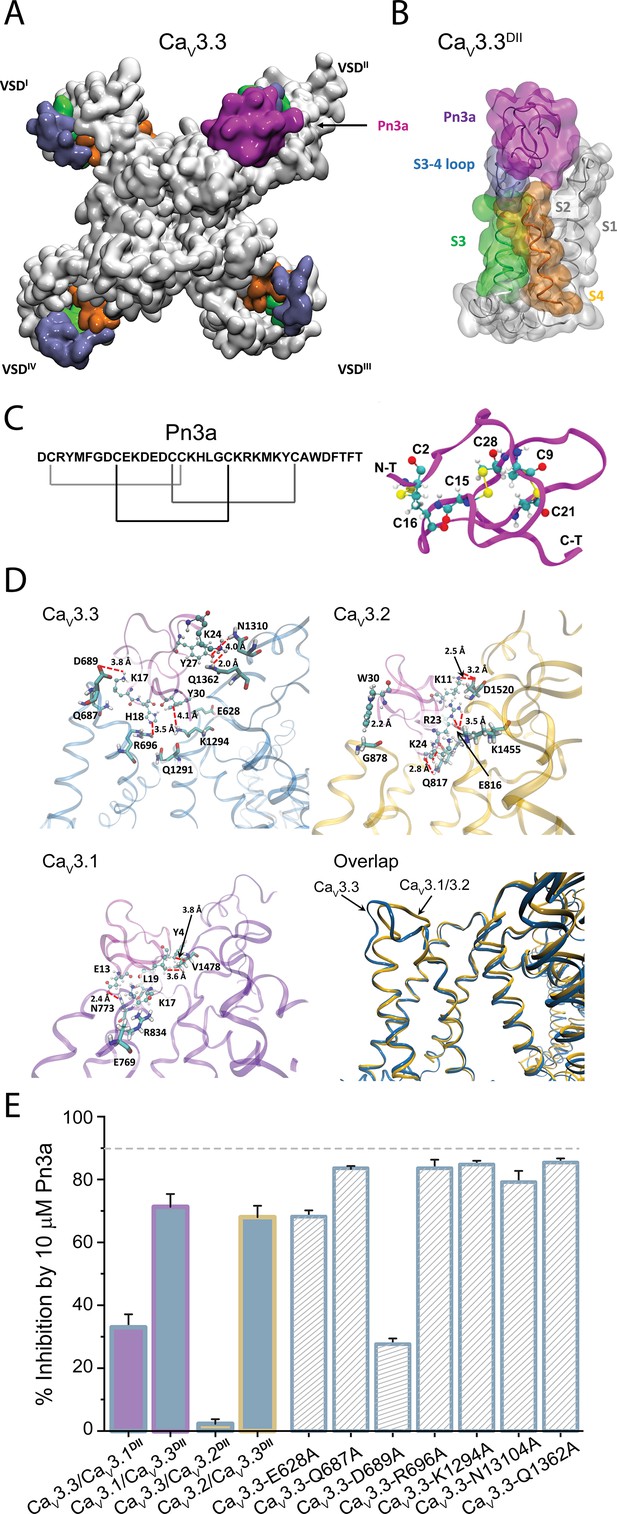
Pn3a and CaV3.3 interactions and subtype specificity.
(A) Top view of Pn3a (magenta) binding to CaV3.3DII. The S3 (lime), S3-4 loop (ice blue), and S4 (pink) are highlighted. (B) Side view of the binding conformation of Pn3a to CaV3.3DII. (C) Left: Pn3a amino acid sequence and disulfide connectivity. Right: 3D structure of Pn3a (PDB ID: 5T4R Deuis et al., 2017) showing cysteine residues in CPK linked by salt bridges (yellow). (D) Pairwise amino acid interactions between Pn3a (magenta) with the extracellular loop of CaV3.3DII (top left, blue), CaV3.2DII (top right, yellow), and CaV3.1DII (bottom left, purple). Overlap of the Pn3a-bound ribbon structures of all CaV3 highlighting differences in extracellular S3-S4 loops between CaV3.3 and the other two CaV3 isoforms (bottom right). S1, S1-2 loop, and S2 were removed for clarity. (E) Bar graph showing percent inhibition by 10 µM Pn3a of CaV3DII domain-swapped CaV3 constructs and point mutants.
-
Figure 6—source data 1
Pn3a docking coordinates for T-type calcium channels.
- https://cdn.elifesciences.org/articles/74040/elife-74040-fig6-data1-v2.zip
Molecular docking was used to assess the most energetically favoured binding poses for Pn3a on the extracellular-facing pockets of DII of the three channel isoforms, CaV3.3, CaV3.2, and CaV3.1 (Figure 6 and Figure 6—figure supplements 1–3). Comparative analyses of toxin interactions with the other T-type calcium channel members showed that Pn3a forms fewer contacts with the DII S3-S4 linker of CaV3.1 and CaV3.2, compared to CaV3.3 (Figure 6D). Evaluation of the inter-residue contacts between the docked Pn3a and CaV3.3DII revealed several receptor residues predicted to form close contacts with the toxin (Figure 6D, upper left inset). In particular, Pn3a-K17 is predicted to be in close proximity to D689 and may form an especially strong interaction due to the possibility of a salt bridge contact. Guided by the docking results, point mutations of selected Cav3.3DII amino acids (E628A, Q687A, D689A, R696A, K1294A, N1310A, and Q1362A) were explored within full-length Cav3.3 channels. In comparison to wild-type CaV3.3, CaV3.3-D689A (27.6% ± 1.8%, n=5, one-way ANOVA p<0.0001) and CaV3.3-E628A (68.2% ± 2.0%, n=5, one-way ANOVA p<0.0001) displayed significant reductions in Pn3a inhibition (Figure 6F), whereas the peptide activity in the other mutants studied (CaV3.3-Q687A: 83.6% ± 0.5%, n=5; CaV3.3-R696A: 83.6% ± 2.7%, n=5; CaV3.3-K1294A: 84.8% ± 1.2%, n=5; CaV3.3-N1310A 79.2% ± 3.5%, n=5; and CaV3.3-Q1362A 85.4% ± 1.3%, n=5) was unaffected. The relevance of CaV3.3-D689 interaction with Pn3a was further substantiated by the loss of inhibition in the analogous charge neutralization KV1.7/Cav3.3DII chimeric construct KV1.7/CaV3.3DII-D262A (Figure 5—figure supplement 1), thus pinpointing key molecular determinants responsible for Pn3a’s selective inhibition of CaV3.3.
Discussion
The present study demonstrates the functional activity of Pn3a as a gating modifier inhibitor of human CaV3.3 channels with >100-fold higher activity over CaV3.1 and CaV3.2. Our chimeric approaches revealed Pn3a’s preference for CaV3.3DII S3-S4 paddle region, whereas comparative molecular docking amongst isoforms identified a novel binding site putatively determining Pn3a’s proclivity towards CaV3.3. Thus, this investigation highlights Pn3a as the first molecular probe available for the study of CaV3.3 contribution in native cells. We propose that the unique features of Pn3a’s CaV3.3 specificity may be exploited to design isoform-selective T-type calcium channel modulators.
CaV3 subtype selectivity
µ-Theraphotoxin Pn3a was originally described as a potent NaV1.7 inhibitor with an IC50 of 0.9 nM, while also inhibiting other tetrodotoxin-sensitive NaV channels (NaV1.1–1.4 and NaV1.6) with 10- to 100-fold less potency (Deuis et al., 2017). More recently Pn3a was shown to inhibit murine dorsal root ganglion HVA ICa and human CaV1.2, CaV1.3, CaV2.1, and CaV2.2 calcium channels expressed in HEK293T cells (IC50 3–10 μM) (McArthur et al., 2020). Perhaps a more interesting aspect of Pn3a activity is its >100-fold selectivity for human CaV3.3 over both CaV3.1 and CaV3.2 (Figure 1) which distinguishes this spider peptide as a unique isoform-selective modulator of T-type calcium channels. Only a handful of venom-derived peptides have been shown to interact with T-type calcium channels. The scorpion peptide Kurtoxin was the first GMP reported to modulate CaV3 channels with a high affinity for CaV3.1 and CaV3.2 channels (Chuang et al., 1998). The closely related peptide KLI, from Parabuthus granulatus, was later shown to inhibit CaV3.3 with an IC50 ~ 450 nM (Olamendi-Portugal et al., 2002). However, the T-type isoform selectivity and binding site of these two peptides were not comprehensively documented at the time. The activity of spider Protoxins I and II against the three CaV3 channels revealed that ProTx-I preferentially modulates CaV3.1 channels, whereas ProTx-II targets CaV3.2 (Bladen et al., 2014; Salari et al., 2016). Thus, Pn3a complements the molecular toolbox for the study of T-type calcium channels in native tissues.
Voltage dependence and current kinetics
Pn3a inhibits CaV3.3 by inducing a depolarizing shift in CaV3.3 voltage dependence of activation in a manner analogous to Kurtoxin, Protoxin I, and Protoxin II actions on CaV3.1 channels (Edgerton et al., 2010; Chuang et al., 1998). This is also consistent with Pn3a’s modulation of NaV1.7 (Deuis et al., 2017) but not of CaV2.2 channels for which a hyperpolarizing shift in the voltage dependence of inactivation was associated with current inhibition (McArthur et al., 2020). In contrast to the twofold slower recovery from inactivation reported for Pn3a-bound NaV1.7 channels, currents mediated by Pn3a-modified CaV3.3 channels recover ~1.5-fold faster from inactivation (Figure 2D/E). This and the observed slowing of inactivation (at high Po) are suggestive of a toxin-induced destabilization of the inactivated state. Nevertheless, given Pn3a’s voltage dependence of Cav3.3 inhibition (Figure 3A), voltage-dependent unbinding of the toxin at more positive potentials could also contribute to an apparent faster recovery from inactivation as reported for CaV active toxins such as AgaIVA (McDonough et al., 1997b) and SNX482 (Bourinet et al., 2001).
Pn3a inhibition of CaV3.3 was voltage-dependent with greater inhibition at more negative potentials (Figure 3A) similar to Kurtoxin-inhibited CaV3.1 channel currents (Chuang et al., 1998). However, a delay in CaV3.1 activation kinetics was apparent in the presence of both Kurtoxin and ProTx-II (Edgerton et al., 2010; Chuang et al., 1998) but not in Pn3a-modified CaV3.3. The spider peptides Pn3a (this study) and ProTx-II (Edgerton et al., 2010) appear to slow T-type channel deactivation suggestive of stabilization of the channel’s closed state, whereas Kurtoxin, from scorpion venom, does not affect CaV3.1 channel closure (Chuang et al., 1998) highlighting incompletely understood aspects of GMP/VGIC interactions.
Interaction of Pn3a with CaV3.3 VS paddles
The portability of the S3-S4 paddle region was shown more than 20 years ago (Swartz and MacKinnon, 1997). To date, most studies, if not all, have used the KV2.1 channel backbone for the identification of GMPs binding sites in NaV and CaV channels expressed in Xenopus oocytes (Bosmans et al., 2008; Salari et al., 2016). For our chimera studies, we selected the KV1.7 backbone given the remarkable scarcity of GMPs interacting with KV1 channels (Finol-Urdaneta et al., 2020). The robust expression of this channel in heterologous systems (Finol-Urdaneta et al., 2006; Finol-Urdaneta et al., 2012) and resistance to Pn3a modulation (Figure 5E/F) make it suitable to assess peptide binding to CaV3.3 paddle motifs in mammalian cells. The generated KV1.7/CaV3.3 constructs resulted in four functional, voltage-gated chimeric potassium channels that exhibited distinct gating properties to those of the parental scaffold reflecting the acquisition of the grafted S3-S4 paddle regions from DI to DIV of CaV3.3. Namely, KV1.7/CaV3.3DI-DIII all displayed >80 mV depolarizing shifts in channel activation compared to wild-type KV1.7, whereas the CaV3.3DIV bearing chimeric construct presented a comparable magnitude shift in the hyperpolarizing direction (Figure 4).
Analogous chimeric approaches of KV2.1/CaV3.1DI-IV have indicated that ProTx-II, PaTx-1, GsAF-I, and GsAF-II exert their inhibitory actions predominantly through interaction with CaV3.1DIII (Salari et al., 2016), whereas Pn3a does so by targeting KV2.1/NaV1.7DII and KV2.1/NaV1.7DIV (Deuis et al., 2017). Here, we observe modest Pn3a inhibition of KV1.7/CaV3.3DIII-mediated currents and potent effects on KV1.7/CaV3.3DII chimeras with current inhibition coupled to toxin-induced rightward shift in the voltage dependence of activation as evidence of preferential interactions with CaV3.3DII (Figure 5B/F). A substantial body of literature has shown that DI-DIII and DIV are important for NaV activation and inactivation, respectively (Ahern et al., 2016). Thus, the predominant effects of Pn3a on CaV3.3 activation are consistent with its interaction with DII of this channel.
Furthermore, it has been shown that ProTx-I inhibits CaV3.3/CaV3.1DIV chimeric channels while interacting less potently with CaV3.3/CaV3.1DII. However, a clear binding site could not be delineated through mutation of individual CaV3.1DII residues as those did not result in measurable changes in toxin affinity (Bladen et al., 2014). Our domain swap experiments verified that Pn3a’s actions are largely determined by CaV3.3DII from which aspartate in position 689 constitutes an important interaction site as shown by its alanine replacement in CaV3.3 and KV1.7/CaV3.3DII. It can be surmised that subtle, but significant GMP/VGIC interaction differences highlight incompletely understood idiosyncrasies related to molecular aspects of ion channel function between isoforms as well as how peptide interactions may be affected by the experimental manipulation and conditions used for their study.
A novel binding site with CaV3 subtype selectivity
Our molecular docking calculations suggest that Pn3a binds within the groove formed between extracellular linkers S1-S2 and S3-S4 supported by electrostatic interactions with the CaV3.3DII S3-S4 paddle, which bears some similarity with the CaV3.1/ProTx-II complex in which favourable binding sites occur in CaV3.1’s DII and DIV (Bladen et al., 2014).
The interaction of Pn3a with CaV3.3 channels involves substantial interactions within the S3-S4 paddle (K17-Q687) and S4 (H18-R696) of CaV3.3DII, while a close contact may also exist between the sidechain of D689 and the sidechain of Pn3a-K17 (Figure 6D, upper left panel). The latter is consistent with the establishment of a salt bridge between these residues. The loss of Pn3a inhibition upon charge neutralization at this position lends support to this docking model.
The proximity between cationic residues (K17 and H18) on loop 3 of Pn3a and the CaV3.3DII S3-S4 paddle places this peptide at the hollow between S1-S2 and S3-S4 linkers in a binding conformation similar to other tarantula toxins targeting NaV channel site 4, like HwTX-IV, ProTx-II, and HNTX-III, that bear critical basic and hydrophobic amino acids within loop 4 that interact with acidic residues on the NaVDII’s S3-S4 linker (Deng et al., 2013; Liu et al., 2013; Bosmans and Swartz, 2010; Figure 6—figure supplements 1 and 2). Hence, the predicted Pn3a interaction with CaV3.3 appears to share overall similarities with previously identified spider peptide toxins modulating NaV and KV channels in which the common bioactive surface consists of positively charged and hydrophobic residues (Smith et al., 2007; Corzo et al., 2005).
The findings presented here establish Pn3a as a gating modifier modulator of CaV3.3 channels interacting with the paddle motif of DII’s VS module through putative stabilization of the closed/resting state and concomitant channel current inhibition. Pn3a’s>100-fold higher potency against CaV3.3 over the other two CaV3 isoforms is rationalized through recognition of a previously unknown drug binding site that may be exploited in the design of isoform-selective CaV3 channel modulators.
Materials and methods
Cell lines, culture, and transfections
Request a detailed protocolHuman embryonic kidney (HEK293T, authenticated by STR profiling and mycoplasm free) cells containing the SV40 Large T-antigen were cultured and transfected by calcium phosphate method as reported previously (McArthur et al., 2018). In brief, cells were cultured at 37°C, 5% CO2 in Dulbecco’s modified Eagle’s medium (DMEM, Invitrogen Life Technologies, VIC, Australia), supplemented with 10% fetal bovine serum (FBS, Bovigen, VIC, Australia), 1% GlutaMAX and penicillin-streptomycin (Invitrogen). HEK293T cells were then transiently co-transfected with the different CaV channel isoforms and green fluorescent protein (GFP) for visualization, using the calcium phosphate method. cDNAs encoding human CaV3.1 (provided by Dr G Zamponi), human CaV3.2 (a1Ha-pcDNA3 was a gift from Dr E Perez-Reyes, Addgene #45809) (Cribbs et al., 1998), human CaV3.3 (a1Ic-HE3-pcDNA3 also from Dr E Perez-Reyes, Addgene #45810) (Gomora et al., 2002) in combination with GFP. CaV3DII swap constructs and KV1.7/CaV3.3 chimeras were custom synthesized by GeneScript, NJ.
Chinese hamster ovary (CHO) cells were used to express KV1.7 and KV1.7-CaV3.3 paddle chimeras. Cell culture conditions were the same as the HEK293T cells except DMEM was substituted with DMEM/F12 (Invitrogen). CHO cells were transfected with cDNAs encoding KV1.7 and KV1.7/CaV3.3DI-DIV chimeras using Lipofectamine 2000 (Invitrogen) as per the manufacturer’s protocol and used for recordings 12–48 hr post-transfection.
Electrophysiology
Request a detailed protocolWhole-cell patch clamp configuration was used to record calcium (ICa) or potassium (IK) currents in transiently transfected HEK293T cells. Recordings were made using a MultiClamp 700B amplifier, digitized with a DigiData1440, and controlled using Clampex11.1 software (Molecular Devices, San Jose, CA). Whole-cell currents were sampled at 100 kHz and then filtered to 10 kHz, with leak and capacitive currents subtracted using a −P/4 protocol for Ca2+ currents and uncorrected for K+ currents. All recordings were series compensated 60–80%. External solution for ICa contained in mM: 100 NaCl, 10 CaCl2, 1 MgCl2, 5 CsCl, 30 TEA-Cl, 10 D-glucose and 10 HEPES, pH 7.3 with TEA-OH. External solution for IK contained in mM: 140 NaCl, 5 KCl, 1 MgCl2, 2 CaCl2, 10 glucose, and 10 HEPES, pH 7.3 with NaOH. Fire-polished borosilicate (1B150F-4, World Precision Instruments, Sarasota, FL) patch pipettes were used with resistance of 1–3 MΩ. Intracellular recording solution contained as follows (mM): 140 KGluconate, 5 NaCl, 2 MgCl2, 5 EGTA, and 10 HEPES, pH 7.2 with KOH. Cells were continuously perfused with extracellular solution at a rate of 1.2 ml/min, while toxin application was superfused onto the cell through a capillary tube attached to a syringe pump (2 µl/min), directly onto the cell being recorded.
For experiments on CaV3s, all cells were held at −90 mV. To examine the onset of block, test pulses (100 ms, 0.5 Hz) to −20 mV were applied. To generate activation curves, cells were pulsed from −90 to +40 mV in 10 mV increments at 0.5 Hz. SSI curves were generated by measuring the peak current from a test pulse to −20 mV when preceded by a 1 s pre-pulse from −90 to 20 mV (0.1 Hz). Recovery from inactivation curves was produced by varying the time (0–2 s) between two depolarizing pules to −20 mV (P1 200 ms, P2 50 ms).
For experiments on KV1.7 and CaV3.3 DI-IV chimeras, cells were held at −80 mV and activation curves were generated by applying a 50 ms pre-pulse to varying potentials depending on the channel construct examined (KV1.7 –60 to 90 mV, DI-III 0 to +160 mV, and DIV −120 to +30 mV) followed by a 50 ms test pulse to measure tail currents (KV1.7 0 mV, DI-III +80 mV, and DIV −20 mV). To measure the onset of Pn3a inhibition, test pulses (50 ms) from a holding potential of −100 mV to a test potential determined for each channel construct (KV1.7 40 mV, DI-III 120 mV, and DIV 0 mV) were elicited at 0.2 Hz.
Structures and homology modelling of human T-type calcium channels and the docking of Pn3a
Request a detailed protocolThe cryo-EM structures of human CaV3.1 (PDB ID: 6KZO) (Zhao et al., 2019) and CaV3.3 (PDB ID: 7WLI) (He et al., 2022) were used for docking calculations. The comparative model of humanCaV3.2 was built upon the cryo-EM structure of hCaV3.1 in the apo form (PDB ID: 6KZO) via the SWISS-MODEL server (https://swissmodel.expasy.org/). The 3D structure of μ-theraphotoxin (TRTX)-Pn3a (PDB ID: 5T4R) (Deuis et al., 2017) was docked into the three T-type calcium channels via Autodock Vina with a grid box of 40Å × 40Å × 40 Å. We also compared the binding mode and the binding affinity amongst the four VSs of CaV3.3, to identify which VS module Pn3a preferentially targets. The docking results were further analysed via Discovery Studio 2017R2 and visualized using Visual Molecular Dynamics (VMD) version 1.9.3 (Humphrey et al., 1996).
Data and statistical analysis
Request a detailed protocolAll data analysis and graphs were generated in OriginPro (Origin Lab Corporation, Northampton, MA). Concentration-response curves were generated by plotting peak current amplitudes in the presence of Pn3a (IPn3a), over the current before Pn3a application (IControl). The resulting curve was fit with a sigmoidal curve according to the following expression:
where IC50 is the half-maximal inhibitory concentration and n is the Hill coefficient. Activation (Equation 2) and SSI (Equation 3) curves were fit by the modified Boltzmann equation:
where I is the current or G is the conductance, Vm is the pre-pulse potential, V0.5 is the half-maximal activation potential, and ka is the slope factor. Recovery from inactivation plots was fit using a single exponential of the following equation:
where τ is the time constant and A is the amplitude. Statistical significance (p<0.05) was determined using paired or unpaired t-test or two-way ANOVA followed by a Tukey multiple comparison test if F achieves the level of statistical significance of p<0.05 and no variance inhomogeneity. All data are presented as mean ± SEM (n), where n is individual cells with all experimental results containing n≥5 individual cells.
Data availability
All data generated or analysed during this study are included in the manuscript and supporting file.
References
-
The hitchhiker’s guide to the voltage-gated sodium channel galaxyThe Journal of General Physiology 147:1–24.https://doi.org/10.1085/jgp.201511492
-
New structures and gating of voltage-dependent potassium (Kv) channels and their relatives: a multi-domain and dynamic questionInternational Journal of Molecular Sciences 20:2.https://doi.org/10.3390/ijms20020248
-
Common mechanisms of drug interactions with sodium and T-type calcium channelsMolecular Pharmacology 82:481–487.https://doi.org/10.1124/mol.112.079715
-
Targeting voltage sensors in sodium channels with spider toxinsTrends in Pharmacological Sciences 31:175–182.https://doi.org/10.1016/j.tips.2009.12.007
-
Voltage-gated ion channels and gating modifier toxinsToxicon: Journal of the International Society on Toxinology 49:124–141.https://doi.org/10.1016/j.toxicon.2006.09.022
-
Inhibition of T-type voltage-gated calcium channels by a new scorpion toxinNature Neuroscience 1:668–674.https://doi.org/10.1038/3669
-
Molecular and functional differences between heartMolecular and Functional Differences between Heart mKv1.7 channel isoform Channel IsoformsThe Journal of General Physiology 128:133–145.https://doi.org/10.1085/jgp.200609498
-
Block of Kv1.7 potassium currents increases glucose-stimulated insulin secretionEMBO Molecular Medicine 4:424–434.https://doi.org/10.1002/emmm.201200218
-
Functional importance of T-type voltage-gated calcium channels in the cardiovascular and renal system: news from the world of knockout miceAmerican Journal of Physiology. Regulatory, Integrative and Comparative Physiology 308:R227–R237.https://doi.org/10.1152/ajpregu.00276.2014
-
Structure, gating, and pharmacology of human CaV3.3 channelNature Communications 13:e28.https://doi.org/10.1038/s41467-022-29728-0
-
Low-threshold calcium currents in central nervous system neuronsAnnual Review of Physiology 58:329–348.https://doi.org/10.1146/annurev.ph.58.030196.001553
-
VMD: visual molecular dynamicsJournal of Molecular Graphics 14:33–38.https://doi.org/10.1016/0263-7855(96)00018-5
-
Distinct kinetics of cloned T-type Ca2 + channels lead to differential Ca2 + entry and frequency-dependence during mock action potentialsThe European Journal of Neuroscience 11:4149–4158.https://doi.org/10.1046/j.1460-9568.1999.00841.x
-
Cloning and expression of a novel member of the low voltage-activated T-type calcium channel familyThe Journal of Neuroscience 19:1912–1921.https://doi.org/10.1523/JNEUROSCI.19-06-01912.1999
-
Investigation of CACNA1I cav3.3 dysfunction in hemiplegic migraCav3.3 Dysfunction in Hemiplegic MigraineFrontiers in Molecular Neuroscience 15:e2820.https://doi.org/10.3389/fnmol.2022.892820
-
Inhibition of human N- and T-type calcium channels by an ortho-phenoxyanilide derivative, MONIRO-1British Journal of Pharmacology 175:2284–2295.https://doi.org/10.1111/bph.13910
-
Analgesic transient receptor potential vanilloid-1-active compounds inhibit native and recombinant T-type calcium channelsBritish Journal of Pharmacology 176:2264–2278.https://doi.org/10.1111/bph.14676
-
Spider venom peptide Pn3a inhibition of primary afferent high voltage-activated calcium channelsFrontiers in Pharmacology 11:633679.https://doi.org/10.3389/fphar.2020.633679
-
Alteration of P-type calcium channel gating by the spider toxin omega-Aga-IVABiophysical Journal 72:2117–2128.https://doi.org/10.1016/S0006-3495(97)78854-4
-
Two tarantula peptides inhibit activation of multiple sodium channelsBiochemistry 41:14734–14747.https://doi.org/10.1021/bi026546a
-
Two new scorpion toxins that target voltage-gated Ca2+ and Na+ channelsBiochemical and Biophysical Research Communications 299:562–568.https://doi.org/10.1016/s0006-291x(02)02706-7
-
Molecular physiology of low-voltage-activated t-type calcium channelsPhysiological Reviews 83:117–161.https://doi.org/10.1152/physrev.00018.2002
-
Molecular interactions of the gating modifier toxin ProTx-II with NaV 1.5: implied existence of a novel toxin binding site coupled to activationThe Journal of Biological Chemistry 282:12687–12697.https://doi.org/10.1074/jbc.M610462200
-
Targeting voltage-gated calcium channels in neurological and psychiatric diseasesNature Reviews. Drug Discovery 15:19–34.https://doi.org/10.1038/nrd.2015.5
Article and author information
Author details
Funding
Rebecca L. Cooper Medical Research Foundation (PG2019396)
- Jeffrey R McArthur
National Health and Medical Research Council (APP1072113)
- David J Adams
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
We are grateful to Alexander Mueller (PhD, Vetter laboratory at the Institute of Molecular Bioscience, University of Queensland) for synthesizing the µ-Theraphotoxin Pn3a used in this research. This work was supported by the Rebecca Cooper Foundation for Medical Research Project Grant (PG2019396) to JRM and the National Health and Medical Research Council (NHMRC) Program Grant (APP1072113) to DJA. JRM and RKF-U are grateful to Emma and Zack O Yepugas for continuous support. Computational resources were provided by the National Computational Infrastructure (NCI) which is funded by the Australian Government, and the Pawsey Supercomputing Centre which is funded by the Australian Government and the Government of Western Australia. We acknowledge PRACE for awarding access to the Piz Daint cluster at the Swiss National Supercomputing Centre (CSCS), Switzerland.
Copyright
© 2022, McArthur et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 967
- views
-
- 253
- downloads
-
- 5
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 5
- citations for umbrella DOI https://doi.org/10.7554/eLife.74040






