Letter from the Chairman
One of the most important highlights for eLife in 2017 was welcoming a fourth funding organisation to our initiative. The Knut and Alice Wallenberg Foundation has joined the Howard Hughes Medical Institute, the Max Planck Society and Wellcome to support the work that eLife is doing to improve the communication and conduct of biomedical research.
The key motivation for this decision by the Wallenberg Foundation is our work in technology and product development, specifically towards the creation of open infrastructure to support the complete publishing process, from submission and peer review, to formatting and presentation of the work. Widespread use of such infrastructure could bring substantial benefits to research communication by reducing costs, encouraging consistency and supporting the shift towards greater accessibility for research.
Widespread use of such infrastructure could bring substantial benefits to research communication by reducing costs, encouraging consistency and supporting the shift towards greater accessibility for research.
In the area of technology development, eLife achieved an important milestone in 2017 with the release of a fully redesigned journal website. Created using best practices in user experience design, the new eLife website is designed to provide readers with a first-class reading experience, enabling humans and machines to consume the latest research more effectively. Behind the outward appearance of the site, our engineers have optimised the architecture and operation of the software to allow very rapid delivery of content especially in settings where bandwidth is limited.
Alongside these developments, eLife reached some important landmarks in publishing in 2017. We marked five years since the journal first launched with an editorial from Editor-in-Chief, Randy Schekman, along with a collection of articles discussing recent efforts to change peer review, including eLife’s consultative approach. Despite the introduction of the publication fee in 2017, submissions to the journal remained strong – only slightly down on 2016 – and we published over 1,300 research articles. The journal provided access to a wealth of influential, diverse and beautiful work, including the visualisation of lymphatic vessels around the human brain, evolution of warm-bloodedness, killing of cancer cells by RNA interference and symbiosis between fungi and plants (see “The science”, below).
In his editorial, Randy reflected on our first five years and how he judges the success of eLife not by any single metric, but by “the support of our editorial board and the authors who send us their outstanding work”. This sentiment sits squarely with eLife’s mission, which ultimately is about helping to create a healthy and vibrant scientific culture, in which science is evaluated not on the basis of where it is published, but on its strengths and the way it has influenced other research. With these aims in mind, research is selected and reviewed under the guidance of the working scientists who run eLife.
At the time of writing, 2018 promises to be no less momentous a year in the development of eLife. The very recent announcement that Randy Schekman will be stepping down next year means that we are just beginning the search for another outstanding individual to lead eLife in the pursuit of our mission. The Board and everyone associated with eLife owes Randy a huge debt of gratitude for everything that he has done to take us this far. This year, we have also appointed an early-career researcher to the eLife Board of Directors, conducted our first innovation sprint aimed at exploring new approaches to open communication in science, and launched the trial of a radical new approach to peer review.
eLife was created to innovate and experiment; as we continue to explore and learn how research communication can be optimised in a digital, networked environment, we hope that our own work – like that of the research community we serve – provides a valuable platform that others can build on.

Modernising technology for research communication
Product and technology development remain a strong focus, given eLife’s ambitions to use digital tools to improve research communication. Our intention is not just to provide the best experience we can for eLife’s own journal, but also to make tools that others can use, engage with and contribute to.
Our intention is not just to provide the best experience we can for eLife’s own journal, but also to make tools that others can use, engage with and contribute to.
The code and resources associated with the new eLife website launched in 2017 – including the designs – are freely available under liberal license terms. But that is not enough to ensure these tools are readily usable by others; our teams are now working hard to engage with potential collaborators and identify the additional work that we need to do to make them as adaptable as possible.
We are committed to working with like-minded partners and were involved in a number of major collaborations in 2017:
- Our work with Hypothesis reached fruition a few months ago with the incorporation of the annotation tool across the journal website. Importantly, a growing number of publishers are adopting these tools, such as The American Diabetes Association and Cambridge University Press.
- We joined the Substance Consortium, which is developing text editors to support web-based publishing, and initiated a specific project with Substance and Stencila to create a prototype for a reproducible research article that incorporates executable code and data.
- We joined forces with the Collaborative Knowledge Foundation (Coko) and their partners – including California Digital Library, Hindawi, EuropePMC – to create open infrastructure for the editorial and peer review processes.
To build on these efforts further, eLife piloted some data science projects in 2017, which led to the recruitment of our first data scientist. Techniques such as machine learning, natural language processing and computer vision are all being explored through the Science Beam project to see how they might contribute to more effective and efficient publishing, discovery and re-use of scientific findings.
eLife is pleased to participate in the growing community of individuals and organisations committed to the development of open infrastructure for research communication. We believe an open and interoperable foundation is needed to support the creation of new services – commercial and non-commercial – that will modernise and improve research communication.
We believe an open and interoperable foundation is needed to support the creation of new services – commercial and non-commercial – that will modernise and improve research communication.
Convening and supporting communities
As a non-profit, mission-driven organisation, eLife’s success is enhanced and enriched by our ability to convene communities of different interested groups and work with them on developing new solutions, as the following examples demonstrate.

2017 in numbers.
Changing peer review
eLife’s work involves a team of almost 400 editors, as well as thousands of reviewers and authors. With their support, we have pioneered a consultative peer review approach and are constantly refining our processes with the hope that it will be easier for other journals to adopt them. The peer review trial that eLife has recently launched is also motivated by a desire to explore new ways to improve peer review, which might be more effective not just for eLife, but across many selective journals.
Reforming research assessment
In 2017 eLife joined forces with several other organisations to reinvigorate the San Francisco Declaration on Research Assessment (DORA) and launch a fully redesigned DORA website. DORA is shifting focus from collecting signatures in support of research assessment reform towards an effort to highlight examples of good practices across the world. In this way, we hope to help shift attention away from the venue of publication as a proxy for research quality to a broader view of research accomplishment.
Unlocking citation data
Our work with the Initiative for Open Citations (I4OC) continued during 2017 and, just as 2018 started, we were able to announce that more than 50% of the citation data deposited with Crossref is now openly available – over 500 million references. The opening up of this data could have substantial benefits, such as increasing the transparency of research evaluation, and will support research into areas as fundamental as the evolution of research and knowledge.
Backing tool creators
eLife is pursuing a number of collaborations across the world involving developers in the USA, Canada, New Zealand, and various countries in Europe. With the appointment of an eLife innovation officer at the end of 2016, we also expanded our outreach in 2017 to early-stage projects, publishing contributions about many promising new projects on eLife Labs, and most recently ran the first eLife Innovation Sprint. This community of developers, researchers and others helps eLife explore how new tools can be devised and adopted to enhance how new results are presented, shared and used.
Partnering with early-career researchers
Finally, we continue to strengthen our work with the early-career community, which is critical to eLife’s long-term mission. The kinds of shifts in culture and evaluation that we hope to see will take time, and the current early-career community can help to shape and then benefit from these changes. We cover early-career issues in interviews and other articles in our Magazine, and host regular webinars. We also took advantage of our fifth anniversary and interviewed some of eLife’s initial early-career authors, learning how these scientists have progressed and how they have built on their earlier work.
Most important of all, we are steadily increasing the role of this community in the operation and governance of eLife:
- We ran the first elections for the Early-Career Advisory Committee in 2017 and received an overwhelming response, which led to the creation of a related initiative: the eLife Ambassadors.
- The early-career reviewer pool is expanding, with several individuals being invited to the Board of Reviewing Editors, and a new initiative to involve early-stage peer reviewers in Evolutionary Biology was launched early in 2018.
- We recruited an early-career researcher to the eLife Board of Directors, so that we have a voice from the early-career community at the highest level of governance in eLife.
The Community page of the newly designed eLife website highlights all of the ways in which we work together with the early-career research community.
Advancing reproducibility
At the beginning of 2017, eLife published the first of the Replication Studies that form part of the Reproducibility Project: Cancer Biology. This long-term project is being conducted by the Center for Open Science and aims to provide a transparent assessment of biomedical research reproducibility. There was substantial interest from the scientific media in the initial results of this project, and, although there has been much discussion about a potential “reproducibility crisis” in science in recent years, the reporting of the first replications studies was balanced and well-informed, thanks in particular to the efforts of eLife Senior Editor Sean Morrison. A Nature editorial also highlighted the value of the decision letters and responses that eLife routinely publishes along with the research articles.
To strengthen the integrity and reliability of the work that we publish in eLife, in 2017 we improved our policies in several areas:
- Recommending that authors deposit software in Github so that the correct version is available to other researchers;
- Publishing transparent reporting forms that authors are required to provide with their submissions;
- Encouraging authors to provide a key resources table listing the reagents, materials and software that are essential to reproduce the results presented.
There are many approaches that will help to encourage greater reproducibility in science, but one of the more novel ideas was a project called ReFigure that eLife supported and for which a prototype was launched in 2017. ReFigure is a way to link figures across the published literature; one of the use cases is to link experiments that are attempts at reproduction, which could prove valuable to other researchers who are interested in building on a published experiment.
The steps that eLife and others are taking to improve the reliability of published research will be all the more successful when assessment practices are also reformed and recognise the researchers who make strenuous efforts to ensure that their work is reproducible.
The science
On Jan 6, 2015, Eve Marder and Vivek Malhotra published an editorial that described the kinds of qualities the editors look for in the research articles they wish to publish.
“An eLife paper should give the reader the pleasure of reading about elegant or clever experiments, of learning something new, of being challenged to think about their subject in a new way, or of seeing a particularly stunning image that has meaning because it shows some of the secrets of life.”
Along those lines, in 2017, eLife published research that has…
...identified two members of a large gene family in fission yeast (Schizosaccharomyces pombe) that act as spore killers. Hu et al. investigated these genes, called cw9 and cw27 and belonging to the wtf gene family, and found they enhance their own transmission by impeding the maturation of spores not inheriting them. Together with a related study by Nuckolls et al., the findings shed new light on the causes of reproductive isolation, and will add to our understanding of speciation and evolution in general.
...provided new insight on the evolution of endothermy (or ‘warm-bloodedness’) in mammals. Rey et al.’s analysis of therapsid fossils suggests that endothermy first emerged among our ancestors towards the end of the Permian period, and may have helped the therapsid species be among the few to survive the mass extinction event at the end of the Permian.

Images: Christina/ CC BY 0 and Ghedoghedo/ CC BY 4.0 .
...revealed that host plant lipids are transferred to symbiotic fungi and needed for their development. Keymer, Pimprikar et al. studied how arbuscules (fungal structures inside the plant cells) form in the Lotus japonicus plant. They carried out fatty acid transfer experiments to show how the plant feeds the fungus with fat molecules as well as sugars.
...reported that the function of RNG105/caprin1 is essential for long-term memory formation. Nakayama, Ohashi et al. found that mice deficient in this protein in the cerebrum and hippocampus regions of the brain can form short-term memories lasting several minutes, but not long-term memories. These results pave the way for future research into human disease.
...indicated that an RNA interference off-target effect may be used to kill cancer cells. Putzbach, Gao, Patel et al. say future studies will determine whether this effect could be exploited to shrink tumours in animal models of cancer. If successful, this in turn could lead to new treatments for cancer patients.
...revealed the existence of lymphatic vessels in the dura mater of people and marmoset monkeys. Absinta, Ha et al. found a way to visualise these vessels using brain magnetic resonance imaging, making it possible to study how the brain removes waste products and circulates white blood cells, and to see whether this process is impaired in aging or disease.
...presented a new omics analysis to help increase understanding of the events that accompany dysfunction in oxidative phosphorylation (OXPHOS) and contribute to mitochondrial diseases. Kühl et al. identified a novel response to OXPHOS dysfunction across five different strains of mouse model, whereby enzymes in the mitochondria necessary for producing ubiquinone (Q) were severely reduced. This deficiency in Q could be a target for future drug development.
We also published a lengthy Feature Article on the Human Cell Atlas, an international project that aims to define all the cell types in the human body in terms of distinctive molecular profiles (such as gene expression profiles) and to connect this information with classical cellular descriptions (such as location and morphology). The availability of such a reference map of the molecular state of cells in healthy human tissues would benefit many areas of basic research, and also provide a framework for understanding cellular dysregulation in human disease.
eLife believes great science needs to be communicated well. We encourage understanding of research through podcasts, interviews, expert commentaries, plain-language summaries, and more. In 2017, we created a single point of access to all of this great material, through the eLife Magazine. And, in another celebration of our five years as a journal with all these features, we explored how other organisations are using plain-language summaries to help readers understand new research.
Financial Summary
Analysis of revenue
In an important step toward establishing our long-term sustainability, eLife introduced a publication fee in January 2017. As outlined in our communications in 2016, the $2,500 fee is designed to cover part of our publishing costs. (See “Setting a fee for publication”).
In 2017, 648 of the 1,307 articles published were subject to a fee, yielding the revenue described below. Approximately half of the articles we published in 2017 were initially submitted before the fee took effect. Others were Research Advances and Replication Studies, which were not subject to a fee, or the authors were granted a waiver.
Given the revenue from publication fees, we required less support from our funders than in 2016.
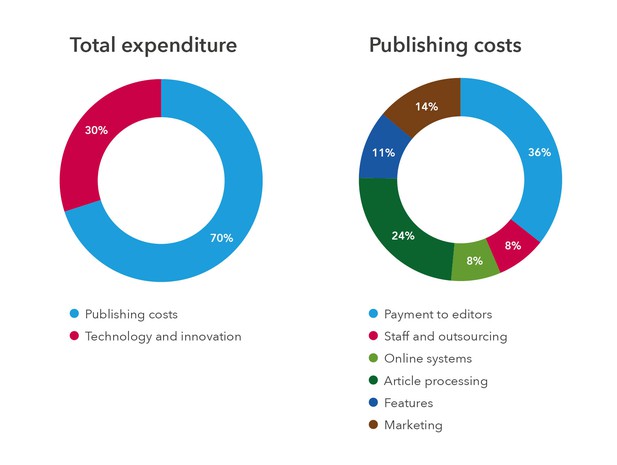
Analysis of expenditure
Our relative expenditure on publishing and technology and innovation remains the same as last year. Within the publishing activity, the major spending areas were also broadly the same as in 2016.
Revenue and expenditure, years ended December 31 2017 and 2016 (in £ thousands)
| Revenue | 2017 | 2016 |
|---|---|---|
| Grants | 3,832 | 4,641 |
| Publication fees | 1,233 | |
| Total | 5,065 | 4,641 |
| Expenditure | 2017 | 2016 |
| Publishing | 3,708 | 3,394 |
| Technology and innovation | 1,614 | 1,420 |
| Total | 5,322 | 4,814 |
| Changes in net assets | (257) | (173) |
The full audited accounts for eLife Sciences Publications Limited for 2017 are available below. As a US-registered tax-exempt organisation, we also publish detailed financial information in our Form 990:
For the latest news from eLife sign up to receive our bi-monthly newsletter. You can also follow @eLife on Twitter.




