Peer review process
Revised: This Reviewed Preprint has been revised by the authors in response to the previous round of peer review; the eLife assessment and the public reviews have been updated where necessary by the editors and peer reviewers.
Read more about eLife’s peer review process.Editors
- Reviewing EditorEmil LouUniversity of Minnesota, Minneapolis, United States of America
- Senior EditorWafik El-DeiryBrown University, Providence, United States of America
Reviewer #1 (Public Review):
Summary:
This finding shows a connection between cancer associated beta-catenin mutations extracellular vesicle secretion. A link between the beta-catenin mutation and expression of trafficking and exocytosis machinery. They used a multidisciplinary approach to explore expression levels of relevant proteins and single particle imaging to directly explore the release of extracellular vesicles. These results suggest a role of extracellular vesicles in immune evasion in liver cancer with the role needing to be further explored in other forms of cancer. I find this work to be compelling and of strong significance.
Strengths:
This paper uses multidisciplinary methods to demonstrate a compelling role of beta-catenin mutations in suppressing EV secretion in tumors. The results and imaging are extremely convincing and compelling.
Reviewer #2 (Public Review):
Summary:
Dantzer and colleagues are investigating the pivotal role of ß-catenin, a gene that undergoes mutation in various cancer cells, and its influence on promoting the evasion of immune cells. In their initial experiments, the authors developed a HepG2 mutated ß-catenin KD model, conducting transcriptional and proteomic analyses. The results revealed that the silencing of mutated ß-catenin in HepG2 cells led to an up-regulation in the expression of exosome biogenesis genes.
Furthermore, the researchers verified that these KD cells exhibited an increased production of exosomes, with the mutant form of ß-catenin concurrently decreasing the expression of SDC4 and Rab27a. Intriguingly, applying a GSK inhibitor to the cells resulted in reduced expression of SDC4 and Rab27a. Subsequent findings indicated that mutated ß-catenin actively facilitates immune escape through exosomes, and silencing exosome biogenesis correlates with a decrease in immune cell infiltration.
In a crucial clinical correlation, the study demonstrated that patients with ß-catenin mutations exhibited low levels of exosome biogenesis.
Strengths:
Overall, the data robustly supports the outlined conclusions, and the study is commendably designed and executed. However, there are a few suggestions for manuscript improvement.
Weaknesses: No weakness
Reviewer #3 (Public Review):
Summary:
In this very important study by Dantzer et al., 'Emerging role of oncogenic b-catenin in exosome biogenesis as a driver of immune escape in hepatocellular carcinoma' the authors define a role for oncogenic b-catenin on exosome biology and explore the link between reduce exosome secretion and tumor immune cell evasion. Using transcriptional and proteomic analysis of hepatocellular carcinoma cells with either oncogenic or wildtype b-catenin the authors find that oncogenic b-catenin negatively regulates exosome biogenesis.
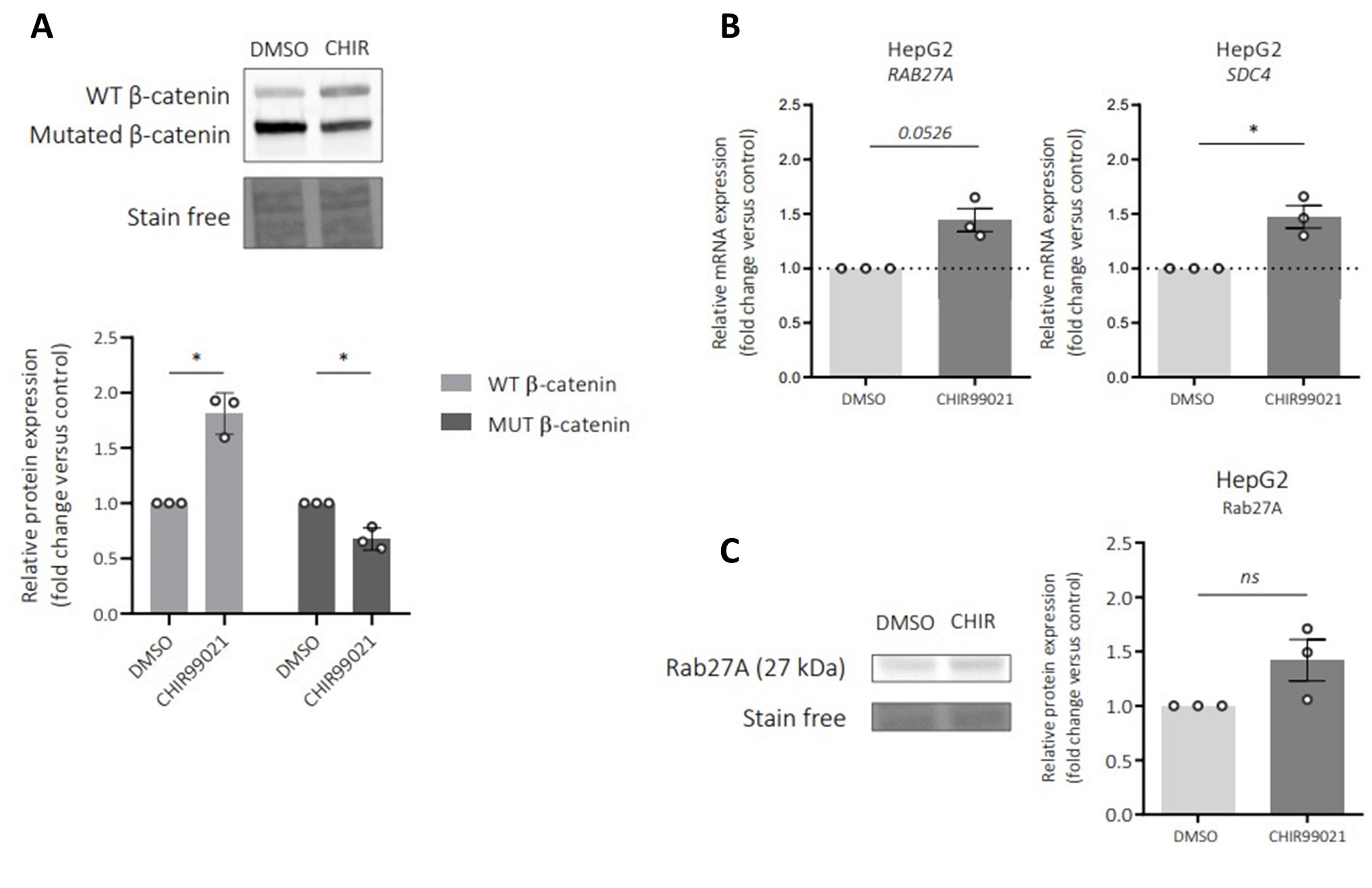
The authors can provide compelling evidence that oncogenic b-catenin in different hepatocellular carcinoma cells negatively regulates exosome biogenesis and secretion, by downregulation of, amongst others, SDC4 and RAB27A, two proteins involved in exosome biogenesis. The authors corroborate these results by inducing b-catenin activation using CHIR99021 in a hepatocarcinoma cell line with non-oncogenic bCatenin (Huh7 cells). The authors can further demonstrate convincingly that reduction in exosome release by hepatocarcinoma spheroids leads to a reduction in immune cell infiltration into the tumor spheroid.
Strengths:
This is a very important and well-conceived study, that appeals to a readership beyond the field of hepatocarcinoma. The authors demonstrate a compelling link between oncogenic bCatenin and exosome biogenesis. Their results are convincing and with well-designed control experiments. The authors included various complementary lines of investigation to verify their findings.
Weaknesses:
One limitation of this study is that the mechanistic relationship of exosome release and how they affect immune cells remains to be elucidated. In this context, the authors conclusions rest on the assumption that hepatocarcinoma immune evasion is based exclusively on the reduced number of exosomes. However, the authors do not analyze exosome composition between exosomes of wildtype and oncogenic background, which could be different.