Divergent kleisin subunits of cohesin specify mechanisms to tether and release meiotic chromosomes
Figures
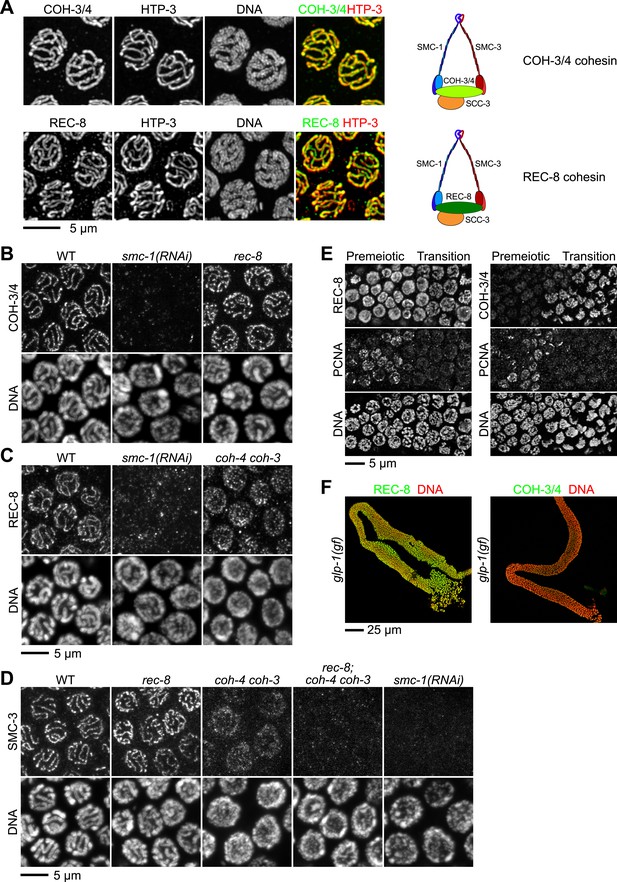
Multiple cohesin complexes that differ in their kleisin subunit bind to C. elegans meiotic chromosomes.
Interdependent loading of REC-8 and COH-3/4 with cohesin SMC proteins is demonstrated. Shown are Z-projected confocal sections through pachytene nuclei (A–D), the distal region of the gonad (E), and entire dissected gonads (F). (A) The predicted α-kleisins REC-8 and COH-3/4 are present along synapsed homologs in pachytene nuclei of wild-type animals and co-localize with the axis protein HTP-3, as expected for meiotic kleisins. COH-3/4 (B) and REC-8 (C) both require SMC-1 for their association with meiotic chromosomes but bind chromosomes independently. (D) SMC-3 associates with chromosomes of rec-8 and coh-4 coh-3 mutants, but SMC-3 staining is undetectable in kleisin triple mutants and smc-1(RNAi) animals. (E) The distal region of the gonad holds nuclei undergoing mitotic proliferation and premeiotic DNA replication (Premeiotic Zone) and nuclei that have entered prophase of meiosis I (Transition Zone). REC-8 is strongly expressed in all germline nuclei, including S phase nuclei, which express GFP::PCN-1. In contrast, COH-3/4 staining is undetectable in GFP::PCN-1 positive nuclei and first appears on meiotic chromosomes in the transition zone, indicating that COH-3/4 cohesin becomes cohesive independently of DNA replication. (F) glp-1(gf) mutations prevent initiation of meiosis; consequently, the gonad fills with mitotically proliferating germ cell nuclei. Robust expression of REC-8, but not COH-3/4, is detected in the mitotic nuclei of glp-1(gf) worms, indicating that COH-3/4 is first expressed during meiosis.
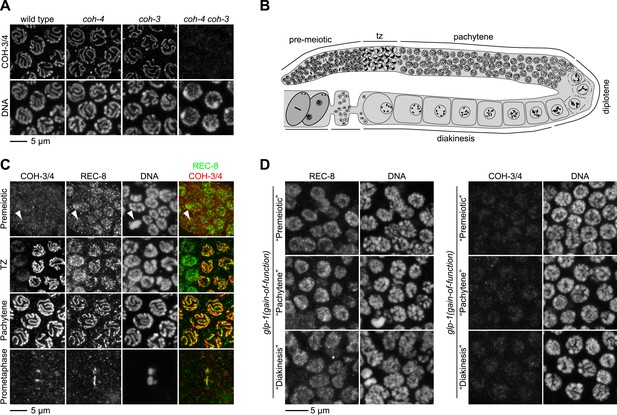
REC-8 accumulates in hermaphrodite gonads prior to the initiation of meiosis, while COH-3/4 becomes detectable only in meiotic nuclei.
(A) Rabbit polyclonal antibodies recognize both COH-3 and COH-4. COH-3/4 antibodies label meiotic chromosomes in wild-type animals as well as coh-3 and coh-4 single mutants. Antibody staining is undetectable only in coh-4 coh-3 double mutants. (B) Cartoon showing that premeiotic nuclei and nuclei in various stages of meiosis occupy distinct, predictable regions of the gonad in wild-type animals. (C) Confocal micrographs of germline nuclei stained with DAPI and antibodies to REC-8 and COH-3/4. COH-3/4 are undetectable in the distal-most region of the gonad, which contains mitotically-cycling germline stem cells and nuclei in premeiotic S phase. Arrowheads indicate a metaphase figure. COH-3/4 appear abruptly on chromosomes at the onset of meiotic prophase (transition zone, TZ), and COH-3/4 are detected along the entire chromosomal axis in pachytene. By prometaphase of meiosis I, COH-3/4 are detected only at the short arm. The pattern of COH-3/4 localization differs from that of REC-8 in two important ways. First, REC-8 is expressed in premeiotic nuclei, although it is unclear whether REC-8 is bound to chromosomes at this stage. Second, while COH-3/4 become restricted to the short arm by prometaphase I, REC-8 is removed from the short arm but persists at the long arm. (D) Gain-of-function alleles of glp-1 prevent initiation of meiosis, and consequently, the germline fills with mitotically proliferating nuclei. High magnification confocal images of regions of the gonad that would contain premeiotic, pachytene, or diakinesis nuclei in wild-type animals are shown. REC-8 is highly expressed in all of these nuclei, but COH-3/4 are not.
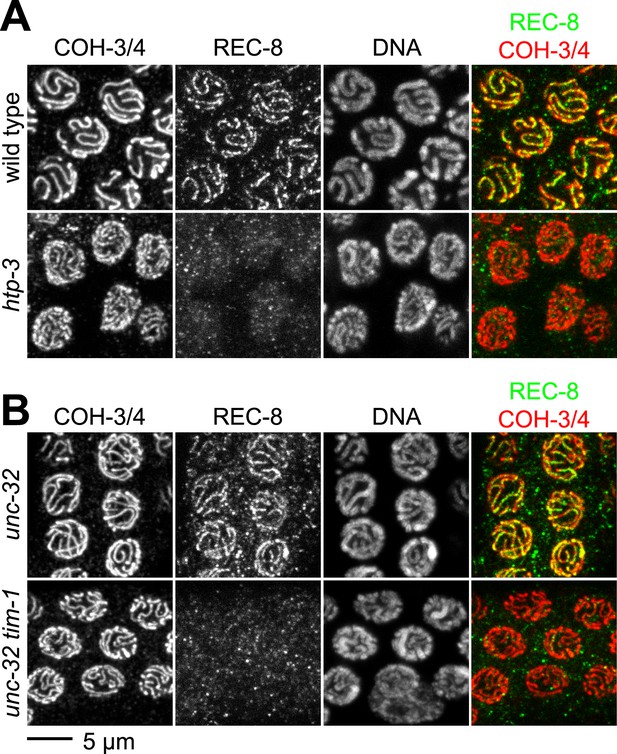
The kleisin subunit determines mechanisms of cohesin loading.
Confocal micrographs of pachytene nuclei reveal that the axial element protein HTP-3 (A) and the Timeless ortholog TIM-1 (B) are both essential for REC-8 cohesin loading, but neither protein is needed for COH-3/4 cohesin loading.
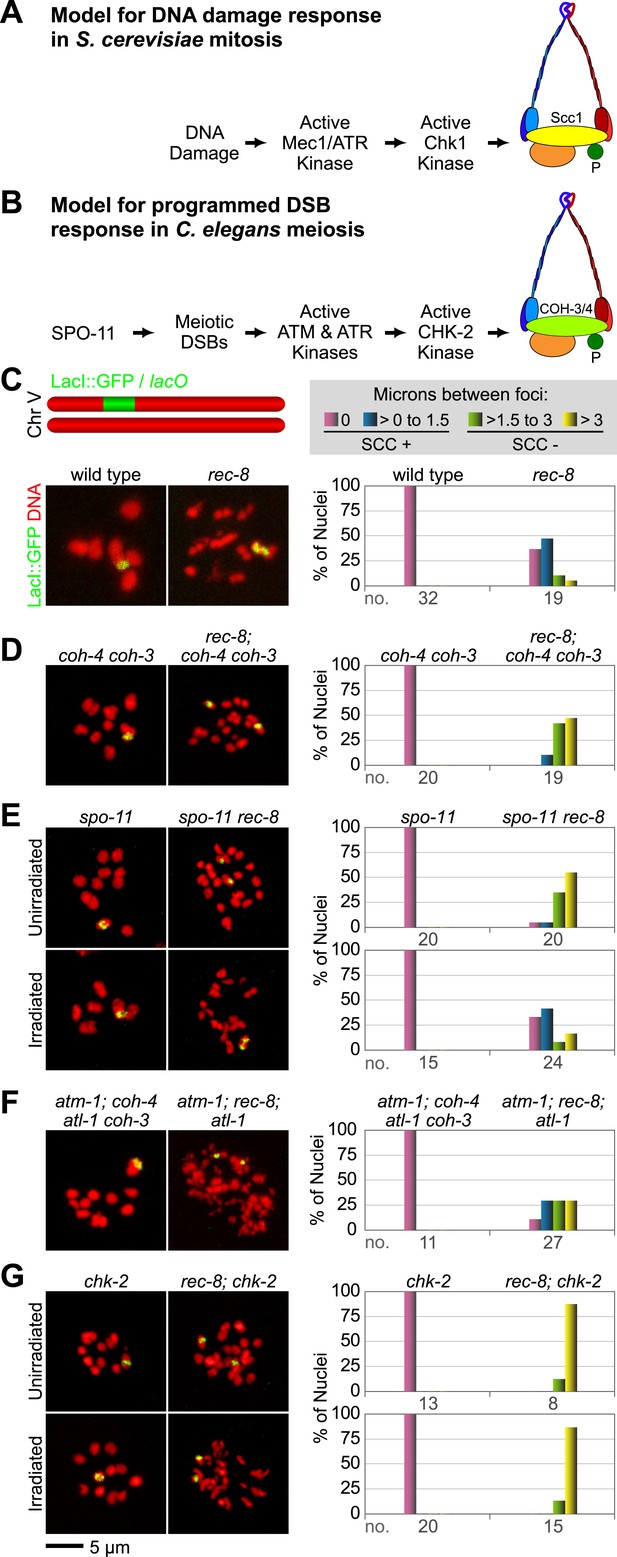
A conserved mechanism initiates SCC in response to programmed DSBs in C. elegans meiosis and exogenous DNA breaks in budding yeast mitosis.
(A) In S. cerevisiae, DNA damage in G2/M activates ATR and Chk1, resulting in Scc1 phosphorylation and SCC establishment. (B) A model for SCC establishment by COH-3/4 cohesin. DSBs created by SPO-11 activate ATM/ATR and CHK-2, leading to COH-3/4 phosphorylation and generation of SCC. (C–G) Data supporting the model in (B). Images on the left show projected Z-sections through entire diakinesis nuclei stained with LacI::GFP (green) and DAPI (red). LacI::GFP bound to a heterozygous lacO array integrated into chromosome V reveals whether sisters are held together by SCC. Charts on the right show quantification of distances between LacI::GFP foci. 0 µm indicates that discrete GFP foci could not be resolved. no. = number of nuclei scored. (C) LacI::GFP labels a single bivalent in wild-type animals, and the two sisters of a single univalent in rec-8 mutants. (D) Sister chromatids are held together by REC-8-dependent SCC in coh-4 coh-3 double mutants, but are apart in kleisin triple mutants. (E) Sister chromatids are held together by SCC in spo-11 mutants, but not in spo-11 rec-8 mutants. DSBs induced by γ-irradiation restore SCC in spo-11 rec-8 mutants. (F) Sisters are apart and extensive chromosomal fragmentation and rearrangement occurs in atm-1; rec-8; atl-1 animals, but not in atm-1; coh-4 atl-1 coh-3 mutants. (G) Cohesion between sisters is not established in rec-8; chk-2 mutants. Irradiation of rec-8; chk-2 mutants does not restore SCC but does induce chromosome fragmentation and rearrangement, demonstrating a role for CHK-2 in SCC establishment that is downstream of DSB formation. CHK-1 is not required for COH-3/4-dependent SCC (Figure 3—figure supplement 3B).
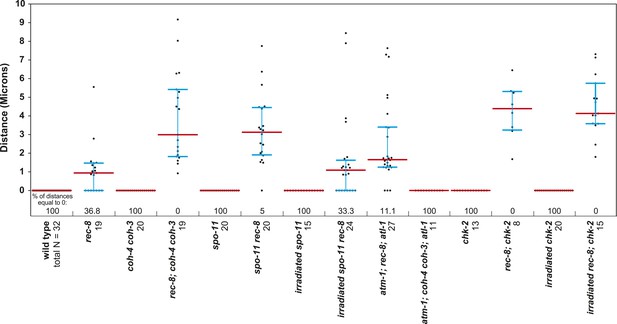
Beeswarm plots show individual distances between LacI::GFP foci in diakinesis nuclei.
Red horizontal lines indicate the median, and blue horizontal lines indicate the interquartile range. A distance of >1.5 microns indicates defective SCC.
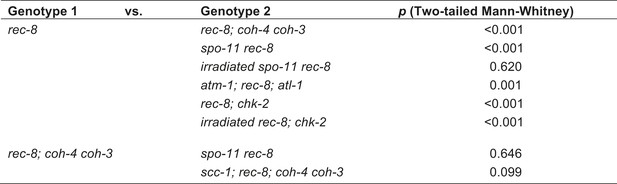

Table of significance values for distances between LacI::GFP foci in diakinesis nuclei.
https://doi.org/10.7554/eLife.03467.008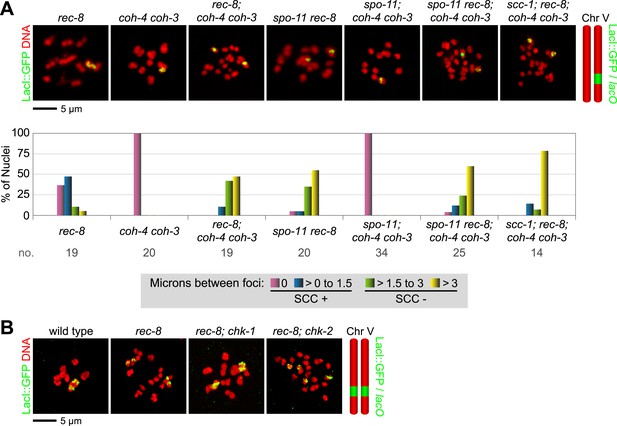
REC-8 and COH-3/4 cohesin tether SCC by different mechanisms.
Shown are projected confocal images of entire diakinesis nuclei stained with DAPI (red) and LacI::GFP (green). SCC was assessed by the distribution of LacI::GFP bound to a lacO array integrated into one (A) or both (B) chromosome V homologs. Bar charts show quantification of distances between LacI::GFP foci. 0 µm indicates that discrete GFP foci could not be resolved. no. = number of nuclei scored. (A) SPO-11-dependent DSBs trigger SCC mediated by COH-3/4 but not REC-8 cohesin. In rec-8 single mutants, the sister chromatids of each homolog are tethered together by COH-3/4-dependent SCC. Similarly, sisters are held together by REC-8-dependent SCC in coh-4 coh-3 double mutants. Sisters are detached in diakinesis nuclei of spo-11 rec-8 double mutants, rec-8; coh-4 coh-3 triple mutants, and spo-11 rec-8; coh-4 coh-3 quadruple mutant animals, and the frequency of detachment and the distance between sisters are similar in all three genotypes. In contrast, disrupting SPO-11 function in coh-4 coh-3 double mutants does not lead to cohesion defects. (B) COH-3/4-dependent SCC requires CHK-2, but not CHK-1. Diakinesis nuclei of rec-8; chk-1(RNAi) animals resemble those of rec-8 single mutants, while sisters are detached in diakinesis nuclei of rec-8; chk-2(RNAi) worms. Thus, CHK-2, but not CHK-1, is required for the establishment of COH-3/4 dependent SCC.
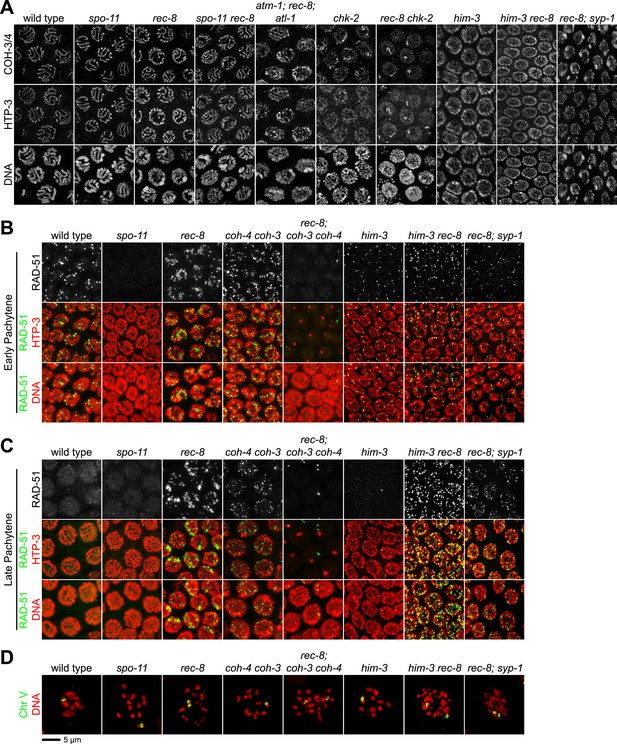
Analysis of COH-3/4 cohesin loading and DSB formation and repair in SCC-defective worms.
(A) Imaging of pachytene nuclei stained with antibodies to COH-3/4 and HTP-3 demonstrated that COH-3/4 associates with meiotic axes in most mutants that fail to establish COH-3/4-dependent SCC. Although COH-3/4 associates with chromosomes of him-3 rec-8 animals, the intensity of COH-3/4 signal is less than that detected in him-3 single mutants, which, in turn, is less than that in wild-type animals (See also Figure 4—figure supplement 2). A reduction in signal is also true of DAPI and HTP-3, which loads onto chromosomes independently of HIM-3 (Goodyer et al., 2008; Severson et al., 2009). Thus, the strong staining of COH-3/4 and HTP-3 observed in wild-type nuclei likely results from the close association of the four chromatid axes via synapsis and SCC, while the reduced staining in him-3 mutants likely results from homolog separation due to defective synapsis, and in him-3 rec-8 mutants from homolog and sister separation due to defective synapsis and SCC. Consistent with this model, a similar reduction in the intensity of COH-3/4 and HTP-3 staining was detected in rec-8 animals also lacking the CR protein SYP-1, which is dispensable for chromosomal loading of all known AE proteins (MacQueen et al., 2002; AFS unpublished data). (B and C) Confocal images of early (B) and late (C) pachytene nuclei stained with DAPI (red) and antibodies to DSB marker RAD-51 (green). Abundant RAD-51 foci are detected in him-3 rec-8 and rec-8; syp-1 mutants, indicating that DSBs are formed. RAD-51 staining persists abnormally late in these mutants, and chromosomal fragmentation and fusions are evident in diakinesis nuclei (D) stained with DAPI (red) and LacI::GFP (green) as in Figure 3C. Thus, establishment of COH-3/4-dependent SCC is essential for homology-directed DSB repair in animals homozygous for a rec-8 deletion. Remarkably, such rearrangements are not detected in kleisin triple mutants. Explaining this finding, few RAD-51 foci form in kleisin triple mutants. Those that do appear in late pachytene, well after DSBs are repaired in wild-type animals.
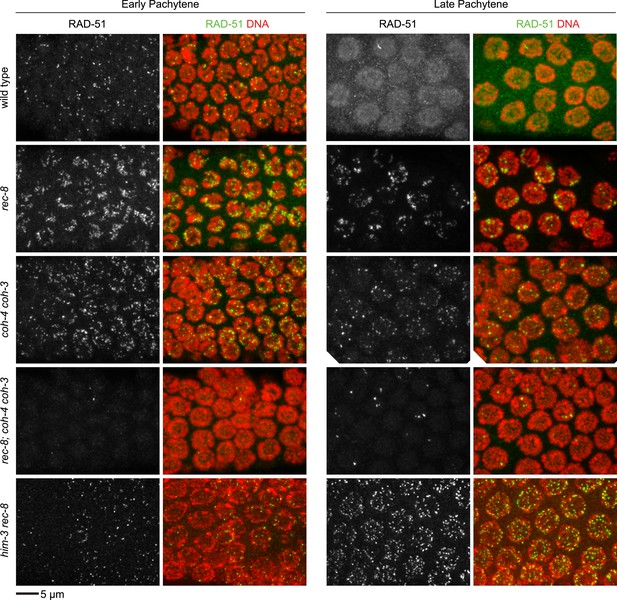
Enlargements of early and late pachytene nuclei of wild-type and mutant animals stained with RAD-51 antibodies.
Abundant RAD-51 foci are present in nuclei of all genotypes, except in rec-8; coh-4 coh-3 mutants, which have a severely reduced number of foci. RAD-51 foci are very rarely present in early pachytene nuclei, and occasional foci appear in late pachytene nuclei.
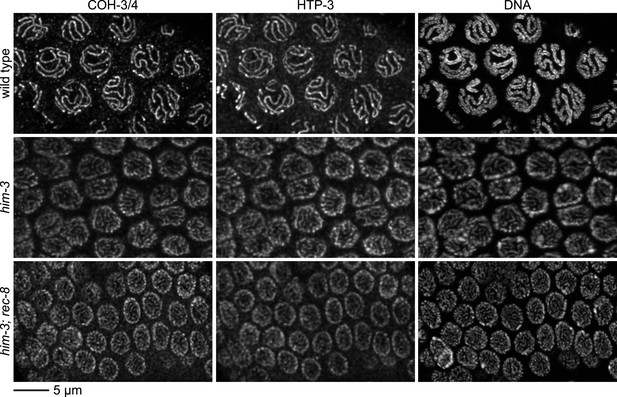
Enlargements of pachytene nuclei from wild-type animals and him-3 or him-3; rec-8 mutants stained with DAPI and antibodies to COH-3/4, HTP-3.
COH-3/4 and HTP-3 associate with pachytene chromosomes but the levels are reduced in mutants relative to wild-type animals, likely reflecting a failure of homolog synapsis in him-3 single mutants and defective synapsis and SCC in him-3 rec-8 mutants.
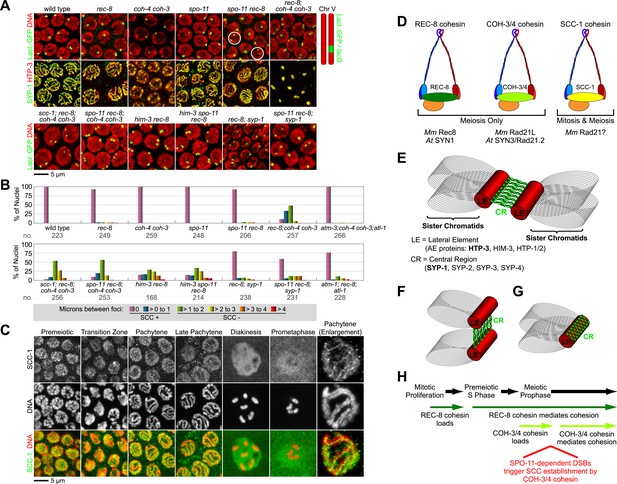
Cohesin-dependent and cohesin-independent SCC holds sisters together in pachytene nuclei.
(A) Projections of confocal Z-sections through entire pachytene nuclei. A single LacI::GFP focus is detected in pachytene nuclei of wild-type, coh-4 coh-3, and spo-11 mutant worms, indicating that sister chromatids are tethered by SCC. In contrast, sisters are separated in most pachytene nuclei of kleisin triple mutants but still remain close together, suggesting that residual SCC persists. Partial depletion of SCC-1 in kleisin triple mutant animals increases both the frequency of sister separation and the distance between sisters, demonstrating a meiotic role for SCC-1. Surprisingly, sisters could be resolved in only ∼10% of nuclei in rec-8 and spo-11 rec-8 worms (white circles). The robust synaptonemal complex (SC) assembly in spo-11 rec-8 worms suggested that SC proteins may tether sister chromatids independently of cohesin. Indeed, disrupting the axial element (AE) protein HIM-3 severely compromised SCC in both rec-8 and spo-11 rec-8 mutants. Disrupting the central region (CR) protein SYP-1 had a lesser effect, suggesting that AE proteins can tether sisters together independently of CR proteins and cohesin. (B) Quantification of sister separation in pachytene nuclei. no. = number of nuclei scored. (C) Z-projected confocal images of wild-type gonads stained with DAPI and antibodies to SCC-1. Similar to REC-8, SCC-1 was detected in premeiotic nuclei and became enriched in axial structures of transition zone and pachytene nuclei. Nucleoplasmic staining obscured any chromosomal signal from pachytene exit until prometaphase; however, SCC-1 was undetectable following nuclear envelope breakdown in prometaphase, indicating that SCC-1 cohesin was removed from chromosomes during diplotene or diakinesis. (D) Similar sets of kleisins function during meiosis in C. elegans, mammals and plants. (E) A schematic of SC structure. Studies in worms have identified four components of the axial/lateral element, or LE (HTP-3, HIM-3, and the functionally redundant proteins HTP-1 and HTP-2) and four components of the CR (SYP-1, SYP-2, SYP-3, and SYP-4). (F and G) Two models of SC-dependent linkages between sisters. (F) CR proteins link AEs formed along each sister. (G) AE proteins hold sisters together independently of CRs. (H) REC-8 cohesin and COH-3/4 cohesin load onto chromosomes at different times and establish SCC by different mechanisms.
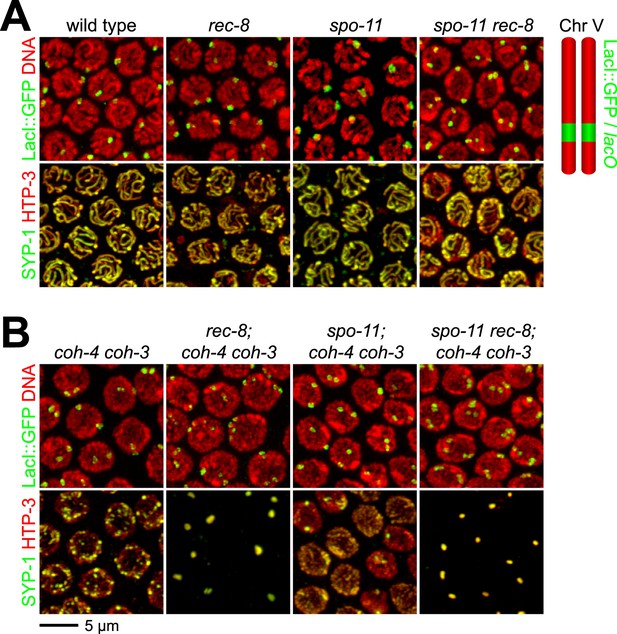
Synaptonemal complex (SC) proteins associate with pachytene chromosomes in rec-8 and spo-11 rec-8 animals, but they do not tether homologous chromosomes.
Shown are confocal images of nuclei stained with LacI::GFP, which labels lacO arrays integrated into both chromosome V homologs (top panels) and antibodies to the axial/lateral element protein HTP-3 and the central region protein SYP-1 (bottom panels) (A) SC proteins associate with pachytene chromosomes of wild-type animals, rec-8 and spo-11 single mutants, and spo-11 rec-8 double mutants. A single focus of LacI::GFP was detected in pachytene nuclei of wild-type and spo-11 worms, as expected since homologs are fully synapsed in these animals. In contrast, two widely separated GFP foci were detected in rec-8 and spo-11 rec-8 mutants, suggesting that synapsis occurs between non-homologous chromosomes or sister chromatids rather than homologs. (B) SC proteins form polycomplexes in pachytene nuclei of rec-8; coh-4 coh-3 triple mutants and spo-11 rec-8; coh-4 coh-3 quadruple mutants. 3–4 LacI::GFP foci can be detected in most nuclei of these animals, consistent with a role for SC proteins in cohesin-independent SCC.
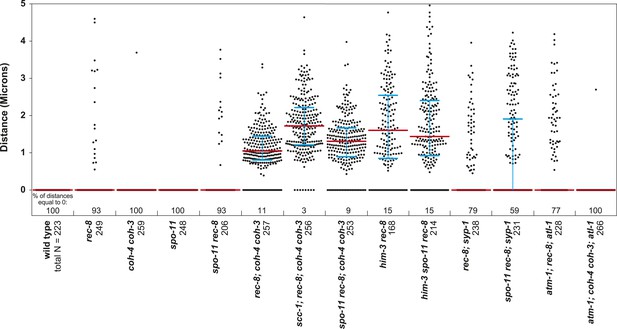
Beeswarm plots show individual distances between LacI::GFP foci in pachytene nuclei.
Red horizontal lines indicate the median, and blue horizontal lines indicate the interquartile range. A distance of >1 micron indicates defective SCC.
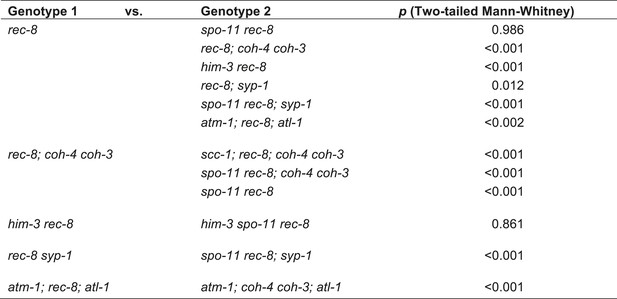
Table of significance values for distances between LacI::GFP foci in pachytene nuclei.
https://doi.org/10.7554/eLife.03467.016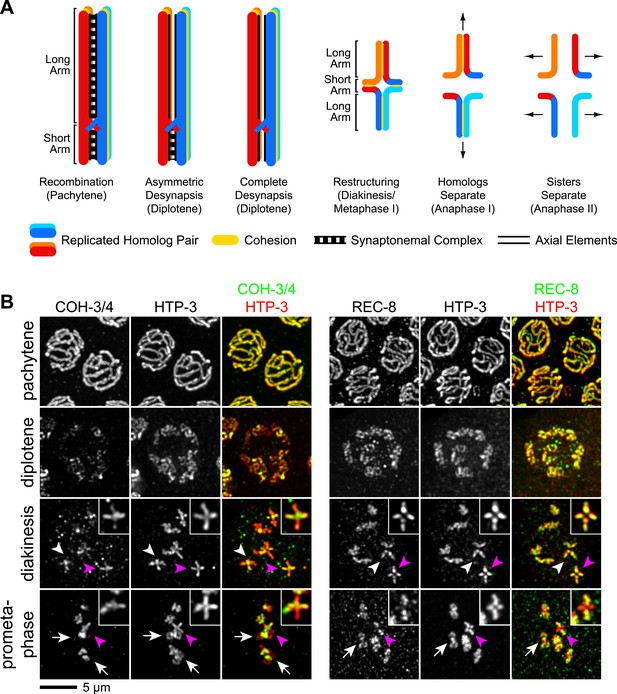
CO recombination triggers removal of REC-8 and COH-3/4 cohesins from reciprocal domains in late prophase/prometaphase of meiosis I.
(A) In worms, CO position determines where SCC will be removed in anaphase I. A single, asymmetrically positioned CO forms between each homolog pair in pachytene, dividing the homologs into long and short arms. In diplotene, each homolog pair is restructured around the CO to form a cruciform bivalent. At anaphase I, SCC is released at the short arm to allow homologs to separate. SCC persists at the long arm until anaphase II. (B) Confocal micrographs showing that REC-8 and COH-3/4 adopt complementary patterns on meiotic chromosomes by metaphase. In pachytene, REC-8 and COH-3/4 overlap with HTP-3 along the entire meiotic axis. In diplotene, HTP-3 and REC-8 persist along the length of the axis, but COH-3/4 staining diminishes at long arms. By diakinesis, COH-3/4 levels are substantially reduced at long arms but not at short arms. In contrast, REC-8 levels usually remain equal at long and short arms until late diakinesis or prometaphase. Diakinesis nuclei shown are from the third oldest oocyte. In prometaphase/metaphase I, REC-8 and COH-3/4 occupy reciprocal domains. REC-8 is reduced or undetectable at short arms, while COH-3/4 is detectable only at short arms. Arrowheads indicate bivalents viewed from the ‘front’, that is with both long and short arms in the image plane. In these bivalents, HTP-3 staining is cruciform and long and short arms can usually be distinguished by their relative lengths. Pink arrowheads indicate the bivalent shown at higher magnification in the inset. Arrows indicate bivalents viewed from the ‘side’, that is with short arms perpendicular to the image plane. In these bivalents, HTP-3 staining resembles a ‘figure 8’, with two loops of uniform staining (the long arms) meeting at a region of more intense staining (the short arms).
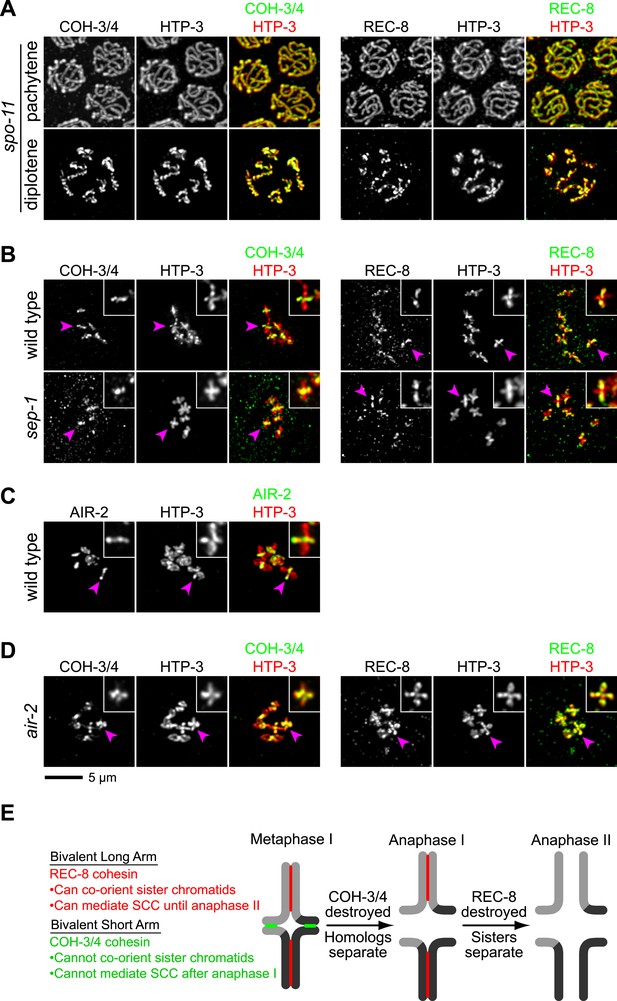
CO recombination triggers separase-independent removal of REC-8 and COH-3/4 from complementary chromosomal territories.
(A–D) Projected images of entire nuclei in pachytene and diplotene (A) or diakinesis (B–D). (A) In spo-11 mutants, CO recombination fails and REC-8 and COH-3/4 are present along the length of meiotic axes in pachytene and diplotene nuclei. In diakinesis, both kleisins are detected at the mid-univalent (Figure 7—figure supplement 1). (B) Depletion of the separase ortholog sep-1 does not impede removal of REC-8 or COH-3/4. (C) AIR-2 associates with short arms of diakinesis bivalents. (D) In air-2(RNAi) animals, REC-8 persists on both long and short arms of prometaphase bivalents, indicating that AIR-2 is required for removal of REC-8 from short arms. COH-3/4 still persists at the midbivalent, indicating that AIR-2 is not required for removal of COH-3/4 from long arms or maintenance of COH-3/4 at short arms. (E) A model demonstrating how establishing reciprocal domains of REC-8 cohesin and COH-3/4 cohesin could facilitate sequential separation of homologs and then sisters. REC-8 cohesin (red) can co-orient sister chromatids and mediate SCC that persists until anaphase II. COH-3/4 cohesin (green) cannot. Restricting REC-8 cohesin to long arms would ensure that co-orientation and persistent SCC occur only in this domain. Co-orientation of long arms would ensure that sister chromatids are pulled to the same spindle pole in anaphase I following proteolytic cleavage of COH-3/4. Proteolysis of REC-8 in meiosis II would allow sisters to separate.
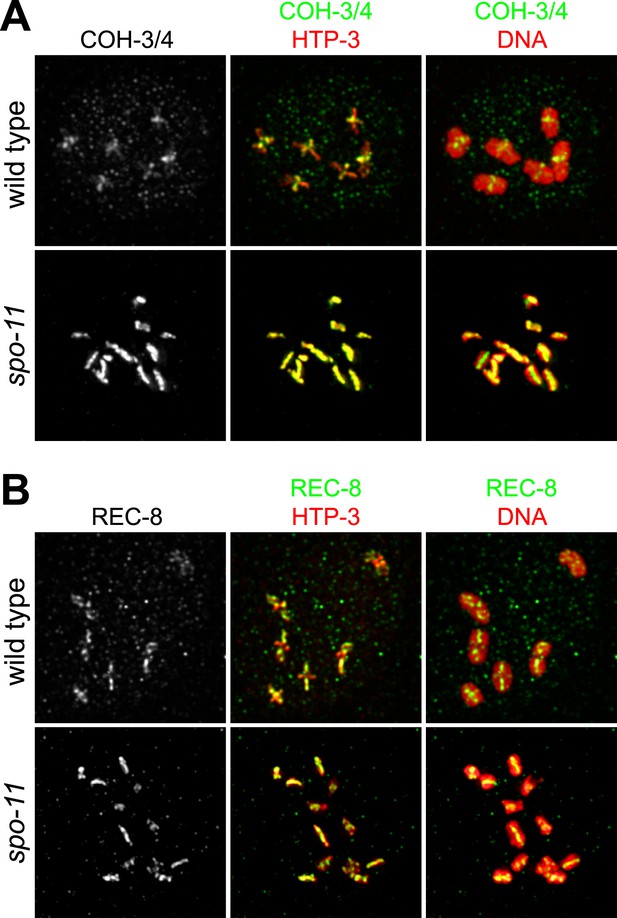
SPO-11-dependent CO recombination triggers the removal of REC-8 and COH-3/4 from complementary domains.
Projected confocal micrographs show the distribution of COH-3/4 and REC-8 on diakinesis bivalents in wild-type worms and univalents of spo-11 mutants. (A) COH-3/4 become substantially reduced or undetectable at the long arms of wild-type bivalents by diakinesis, but persists at high levels at short arms. In contrast, the AE protein HTP-3 is present at uniform levels at both long and short arms. (B) In contrast, REC-8 becomes reduced at short arms but persists at high levels at long arms. CO recombination fails in spo-11 mutants, and homologs remain apart as discrete univalents. COH-3/4 (A) and REC-8 (B) both associate with univalents in diakinesis nuclei of spo-11 mutants.
Additional files
-
Supplementary file 1
Strains used in this study.
- https://doi.org/10.7554/eLife.03467.020
-
Supplementary file 2
Oligos for amplification of dsRNA.
- https://doi.org/10.7554/eLife.03467.021





