Neuroblasts: Maintaining neural stem cell identity in the brain
Stem cells are cells that have the ability to divide and produce both new stem cells (a process called self-renewal) and specific types of cell (a process called differentiation). There are many different types of stem cell. The brain of the fruit fly Drosophila, for example, contains two different types of neural stem cell. Type I neuroblasts divide to produce a new type I neuroblast and a cell called a ganglion mother cell. The new neuroblast can undergo several more rounds of division, but the ganglion mother cell can only divide once, to produce specific cell types (either neurons or glia cells). Type II neuroblasts, on the other hand, divide to produce a new type II neuroblast and a cell called an intermediate neural progenitor. Each intermediate neural progenitor cell can undergo several rounds of proliferation, each of which produces a ganglion mother cell (Homem and Knoblich, 2012). This ‘amplifies’ the number of neurons and glia cells that are produced.
Controlling the proliferation and differentiation of intermediate neural progenitors as the brain develops is crucial because these cells can also undergo a developmental reversal that results in them becoming neuroblast-like cells. These can then overproliferate and form malignant tumours in the brain. Recent studies in Drosophila have identified several regulatory proteins that prevent this developmental reversal and restrict the proliferation of intermediate neural progenitors. These include the post-transcriptional regulator Brain tumor (Bello et al., 2006; Betschinger et al., 2006; Lee et al., 2006), the Notch signaling pathway component Numb (Wang et al., 2007), the Drosophila SWI/SNF chromatin remodeling complex (Eroglu et al., 2014), and the zinc-finger transcription factor Earmuff (Weng et al., 2010). These proteins also work together to restrict proliferation in type II neuroblasts and the cells that they generate (Figure 1).
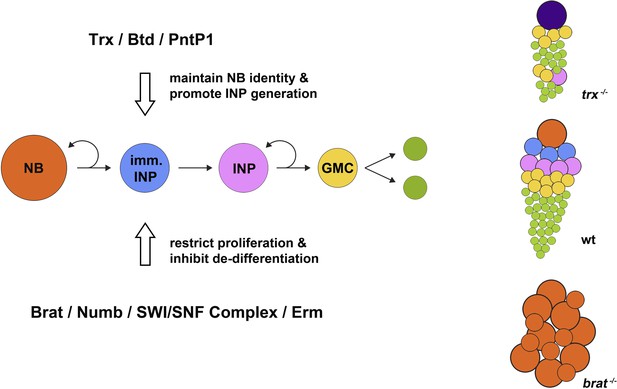
Maintaining neural stem cell identity in the Drosophila brain. A type II neuroblast (NB; orange circles) divides to self-renew and to give rise to an immature intermediate neural progenitor cell (imm. INP; blue circles), which becomes a mature INP (pink) that undergoes several further rounds of self-renewing proliferation. Each round generates a ganglion mother cell (GMC; yellow), each of which gives rise to two differentiated neural cells (green). During neuroblast proliferation, several proteins, including Brat, Numb, SWI/SNF complex, and Erm, restrict the ability of INPs to proliferate and inhibit the de-differentiation of immature INPs. In mutants that lack the brain tumor gene (brat−/−; bottom right), immature INPs revert to neuroblast-like cells that overproliferate and can form brain tumours. On the other hand, as shown by Komori et al. and Xie et al., Trithorax (Trx) and its direct targets Buttonhead (Btd) and Pointed P1 (PntP1) act to maintain the identity of type II neuroblasts and promote the generation of INPs. Hence, in mutants lacking the trithorax gene (trx−/− mutants; top right), INPs are nearly completely lost compared with wild type (wt; centre right), and type II neuroblasts are transformed into type I neuroblasts (purple) that directly generate ganglion mother cells instead of INPs; this ultimately results in fewer neural cells being produced.
It is equally important for type II neuroblasts to maintain their ability to generate self-renewing intermediate neural progenitors so that enough neural cells are produced. The transcription factor Pointed P1 is known to play an important role in this process (Zhu et al., 2011). However, the other proteins that type II neuroblasts might need in order to maintain their identity and generate intermediate neural progenitor cells were not known. Now, in eLife, researchers at the University of Michigan Medical School report that Trithorax, a protein that modifies histones, has a crucial role in these processes in Drosophila (Komori et al., 2014) while, independently, researchers at the State University of New York (SUNY) and Syracuse University report that a transcription factor called Buttonhead is also critically important (Xie et al., 2014).
The fate of cells can be controlled by a number of methods. For example, adding a methyl group to histone H3 at lysine 4 (H3K4) is known to be involved in controlling cell fate during development. Therefore, Cheng-Yu Lee and colleagues at Michigan, including Hideyuki Komori as the first author, tested whether Trithorax—a protein that adds methyl groups to H3K4—might also play a role in maintaining the identity of Drosophila neuroblasts (Komori et al., 2014). By using a combination of antibodies that recognize the different cell markers found on type I and type II neuroblast lineages, Komori et al. showed that Trithorax is necessary for maintaining the functional identity of type II neuroblasts as they proliferate. Thus, a significant reduction in the number of intermediate neural progenitors occurs in type II neuroblast lineages that have a defective version of the trithorax gene. These mutant type II neuroblasts instead transform into type I neuroblasts, which directly generate ganglion mother cells instead of intermediate neural progenitors (Figure 1).
To work in type II neuroblasts, Komori et al. found that Trithorax must have the ability to add methyl groups to histones. However, preventing Trithorax from working did not affect the overall H3K4 methylation pattern in type II neuroblasts, which suggests that Trithorax only regulates a few specific genes. Komori et al. identified a small number of genes specifically expressed in type II neuroblasts, including buttonhead and pointed, and using biochemical analysis found that Trithorax indeed binds directly to the transcription start site of both of these genes. In addition, removing working copies of the buttonhead gene from type II neuroblasts reduced the number of mature intermediate neural progenitors in the fly brain, and over-expressing buttonhead restored the ability of type II neuroblasts with mutant (less functional) trithorax genes to produce intermediate neural progenitor cells. Furthermore, mis-expressing buttonhead in type I neuroblasts caused these cells to behave more like type II neuroblasts and generate cells that resemble intermediate neural progenitor cells.
The role of buttonhead in the type II neuroblast lineages was confirmed and investigated further by Sijun Zhu of SUNY and colleagues, including Yonggang Xie as the first author (Xie et al., 2014). They also observed a marked reduction in the number of intermediate neural progenitor cells produced from buttonhead mutant type II neuroblasts, due to the premature differentiation of immature intermediate neural progenitors into ganglion mother cells. Moreover, Xie et al. showed that the Buttonhead protein most likely prevents this premature differentiation by suppressing the expression of a protein called Prospero that inhibits cell self-renewal and promotes cell cycle exit and differentiation. Xie et al. also provide evidence suggesting that in type II neuroblasts, Buttonhead can cooperate with the transcription factor Pointed P1 to specify the neuroblast function and promote the generation of intermediate neural progenitors (Figure 1).
Komori et al. and Xie et al. provide insight into how the identity of neural stem cells is maintained in the Drosophila brain. This work demonstrates that the histone modification protein Trithorax, together with its direct transcription factor targets Buttonhead and Pointed P1, is crucial for type II neuroblasts to maintain their identity and promotes the generation of intermediate neural progenitor cells.
In the future, it will be important to analyze how this identity maintenance program ensures normal brain development in Drosophila by interacting with a previously characterized program that restricts cell proliferation. Moreover, the mammalian versions of Trithorax (called SET/MLL) and Buttonhead (Sp8) play important roles in mammalian brain development. It will therefore also be important to determine if the Trithorax-dependent mechanisms identified in the fly also operate in the mammalian brain to maintain neural stem cell identity.
References
-
Drosophila neuroblasts: a model for stem cell biologyDevelopment 139:4297–4310.https://doi.org/10.1242/dev.080515
-
Ets transcription factor Pointed promotes the generation of intermediate neural progenitors in Drosophila larval brainsProceedings of the National Academy of Sciences of USA 108:20615–20620.https://doi.org/10.1073/pnas.1118595109
Article and author information
Author details
Publication history
Copyright
© 2014, Jiang and Reichert
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 3,959
- views
-
- 175
- downloads
-
- 3
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 3
- citations for umbrella DOI https://doi.org/10.7554/eLife.05000
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Developmental Biology
- Neuroscience
Intermediate neural progenitor cells (INPs) need to avoid differentiation and cell cycle exit while maintaining restricted developmental potential, but mechanisms preventing differentiation and cell cycle exit of INPs are not well understood. In this study, we report that the Drosophila homolog of mammalian Sp8 transcription factor Buttonhead (Btd) prevents premature differentiation and cell cycle exit of INPs in Drosophila larval type II neuroblast (NB) lineages. We show that the loss of Btd leads to elimination of mature INPs due to premature differentiation of INPs into terminally dividing ganglion mother cells. We provide evidence to demonstrate that Btd prevents the premature differentiation by suppressing the expression of the homeodomain protein Prospero in immature INPs. We further show that Btd functions cooperatively with the Ets transcription factor Pointed P1 to promote the generation of INPs. Thus, our work reveals a critical mechanism that prevents premature differentiation and cell cycle exit of Drosophila INPs.





