A conserved quality-control pathway that mediates degradation of unassembled ribosomal proteins
Figures
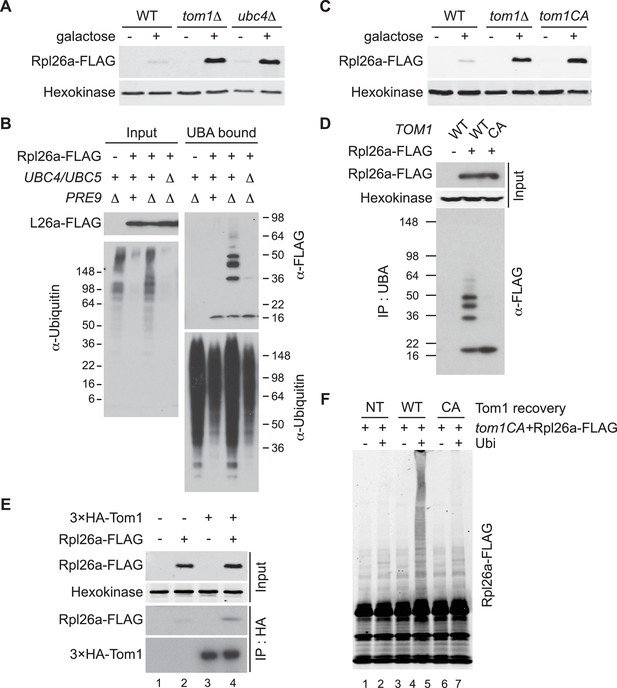
Ubc4/5 and Tom1 are the E2 and E3 enzymes responsible for ERISQ.
(A) Rpl26aFLAG accumulates in tom1∆ and ubc4∆. Accumulation of Rpl26aFLAG upon galactose induction in WT, tom1∆ and ubc4∆ cells was evaluated by SDS-PAGE and immunoblotting with the indicated antibodies. n = 3 biological replicates. (B) Rpl26aFLAG ubiquitination depends on Ubc4/Ubc5. Rpl26aFLAG was induced in cells of the indicated genotypes and cell lysates were prepared and subjected to pull-down with UBA domain resin. Input and bound proteins were evaluated as in (A). n = 3 biological replicates. (C) Rpl26aFLAG accumulates in tom1CA cells. As in (A) except that the Tom1 ligase-dead (tom1CA) mutant was used. n = 3 biological replicates. (D) Rpl26aFLAG ubiquitination depends on Tom1. As in (B) except that cells expressing WT Tom1 or Tom1CA were treated with bortezomib for 45 min after addition of galactose. n = 3 biological replicates. (E) Rpl26aFLAG binds 3xHATom1. Anti-HA immunoprecipitates from cells expressing 3×HATom1 and Rpl26aFLAG were immunoblotted with antibodies to HA, FLAG, and hexokinase. n = 3 biological replicates. (F) In vitro ubiquitination of Rpl26aFLAG by Tom1. Rpl26aFLAG retrieved in 3xHATom1CA immunoprecipitates was supplemented or not with E1/E2/ubiquitin/ATP (Ubi) and Tom1 retrieved from untagged (NT), 3xHATOM1 (WT), or 3xHATOM1CA (CA) cells, as indicated. See detailed methods in Material and methods. n = 3 biological replicates.
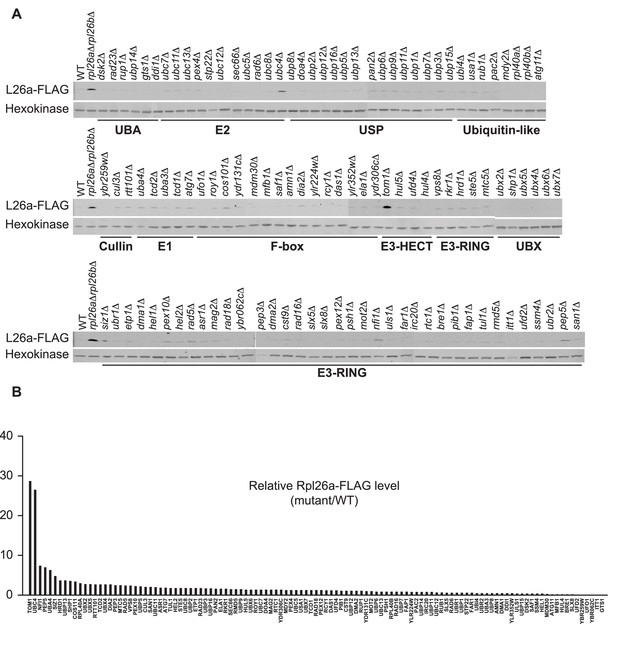
Identification of ERISQ defect in tom1∆ and ubc4∆.
(A) One hundred fifteen different knockout mutant strains each containing a plasmid that expressed Rpl26aFLAG from the GAL10 promoter were used. Rpl26aFLAG induced in rpl26a∆rpl26b∆ cells was used as a positive control. n = 1 biological replicate. (B) Quantification of data in (A).
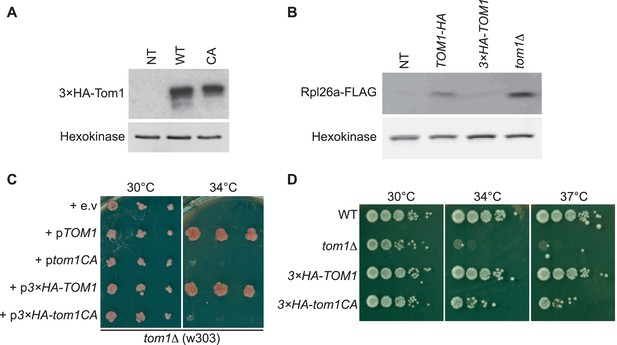
Characterization of tagged and ligase-dead Tom1.
(A) Protein level of 3×HATom1 in cells expressing Tom1 (NT), 3×HATom1 (WT) and 3×HATom1CA (CA). n = 2 biological replicates. (B) Protein level of overexpressed Rpl26aFLAG induced in cells expressing Tom1 (NT), Tom1HA, 3×HATom1 and tom1∆. Note that a C-terminal tag on Tom1 compromises function, allowing for greater accumulation of galactose-induced Rpl26aFLAG. n = 2 biological replicates. (C) Cells of the indicated genotypes were spotted on SC-TRP and incubated at 30°C or 34°C for 2 days. n = 2 biological replicates. (D) As in (C), except that cells of the indicated genotypes were spotted on YPD and incubated at the indicated temperatures for 2 days. n = 2 biological replicates.
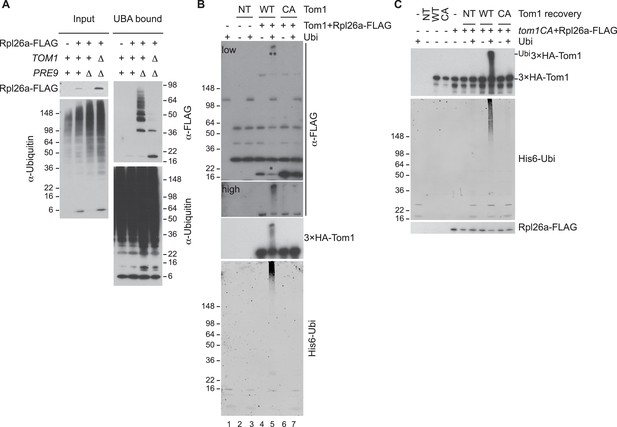
Tom1 mediates ubiquitination of overexpressed Rpl26a.
(A) Polyubiquitination of Rpl26aFLAG. Rpl26aFLAG was induced in cells of the indicated genotypes and cell lysates were prepared and subjected to pull-down with UBA domain resin. Input and bound proteins were fractionated by SDS-PAGE and detected by immunoblot with the indicated antibodies. n = 2 biological replicates. (B) In vitro ubiquitination of Rpl26aFLAG by Tom1. Lysates of cells expressing Rpl26aFLAG and the indicated allele of 3xHATom1 (NT is untagged control) were subjected to pull-down with anti-HA followed by addition of E1/E2/ubiquitin/ATP (Ubi) and incubation at 30°C for 1 hr. Reaction products were evaluated by immunoblot with the indicated antibodies. *indicates unmodified Rpl26aFLAG. **indicate ubiquitinated Rpl26aFLAG. Note the increase in ubiquitinated Rpl26aFLAG (**) and the loss of unmodified Rpl26aFLAG (*). n = 2 biological replicates. (C) Additional immunoblots of samples in Figure 1F. n = 3 biological replicates.
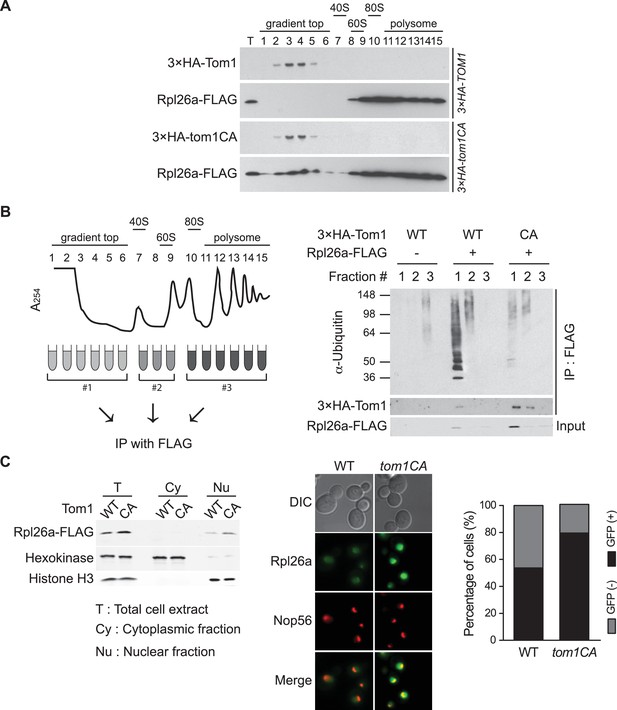
Tom1 functions in non-ribosomal fractions.
(A) Sucrose gradient fractionation behavior of 3xHATom1 and Rpl26aFLAG upon galactose induction of Rpl26aFLAG in 3×HATOM1 or 3×HATOM1CA cells. T indicates total extract. n = 2 biological replicates. (B) Tom1 is required for ubiquitination of unassembled Rpl26aFLAG. Left: experimental scheme. Right: cells were treated with bortezomib for 30 min after induction of Rpl26aFLAG with galactose and then lysed and fractionated as in panel A prior to being processed as depicted in panel B. n = 2 biological replicates. (C) Rpl26a accumulates in the nucleus of tom1CA cells. Left: Subcellular fractionation of Rpl26aFLAG induced in WT and tom1CA cells. Histone H3 and Hexokinase were used as nuclear and cytoplasmic markers, respectively. CA refers to Tom1-Cys3235Ala. Right: Fluorescence microscopy of Rpl26aGFP induced in WT and tom1CA cells. Nop56-RFP marks nucleoli. Shown at far right is the percentage of GFP positive cells. n = 2 biological replicates.
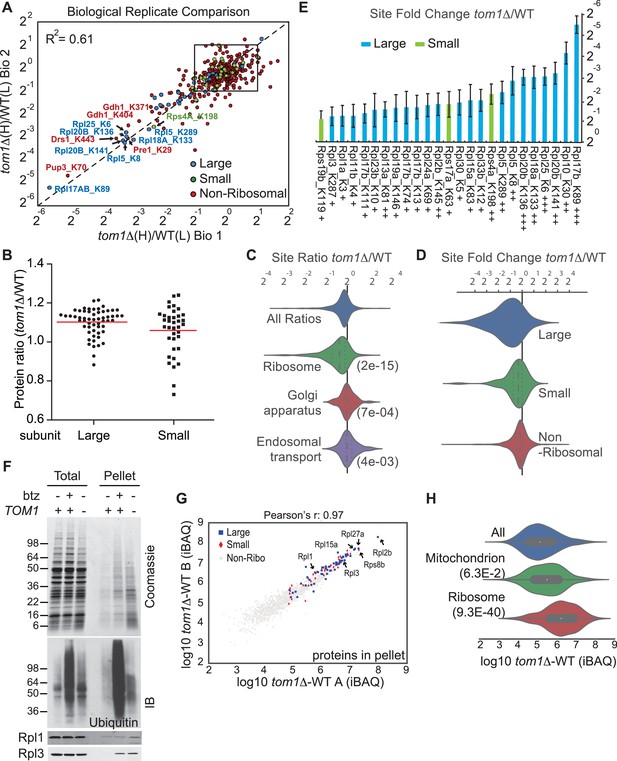
Diminished ubiquitination and accumulation of insoluble ribosomal proteins in tom1 cells.
(A) Diminished ubiquitination of ribosomal proteins in tom1∆. Scatter plot of the SILAC ratios (tom1∆/WT) for GlyGly-modified peptides identified in biological replicate 1 versus 2. Sites with the largest decrease in ubiquitination are annotated. The other pairwise comparisons are in Figure 3—figure supplement 2A. n = 3 biological replicates. (B) Column scatter plot representing the distribution of ratios (tom1∆/WT) for proteins of the large (circles) and small (square) ribosome subunits. A red bar indicates the mean. (C) Violin plot of gene ontology analysis of ubiquitinated proteins that had one or more ubiquitination site that decreased by ≥ 2-fold. The most strongly affected categories are shown. The number in parentheses refers to the disproportionate enrichment for the category in the top 10% of identifications and is the Benjamini and Hochberg corrected p-value from a Fisher Exact test. (D) Violin plot representing the distribution of ubiquitin site occupancy ratios (tom1∆/WT) for the large (blue) and small (green) ribosome subunits, and non-ribosomal proteins (red). (E) The 25 ribosomal ubiquitination sites with the largest decrease in ubiquitin occupancy in tom1∆. +++p<0.001; ++p<0.01; +p<0.05. Each site was observed in at least two of the three biological replicates. The error bars represent 95% confidence intervals. Note that ubiquitination at K37 and K69 in Rpl26b was decreased by 2.4-fold and 1.6-fold in tom1∆, respectively (Supplementary file 3B). (F, G) Insoluble ribosomal proteins accumulate in tom1∆. (F) Detergent-insoluble pellet fractions isolated from lysate (Total) of indicated cells were analyzed by SDS-PAGE and staining with Coomassie blue or immunoblotting with the indicated antibodies. The pellet fraction is overloaded 20-fold compared to the total and supernatant fractions. n = 2 biological replicates. (G) Scatter plot representing ∆iBAQ of biological replicate B vs. A for insoluble proteins in tom1∆ mutants. Ribosomal proteins with the largest increase in the pellet fraction upon TOM1 deletion, and Rpl1 and Rpl3 are annotated. Pearson’s r-value is indicated on top of the plot. The other pairwise comparisons are in Figure 3—figure supplement 3B. n = 3 biological replicates. (H) Gene ontology analysis of proteins exhibiting increased insolubility in tom1∆. Analysis is the same as for panel C.
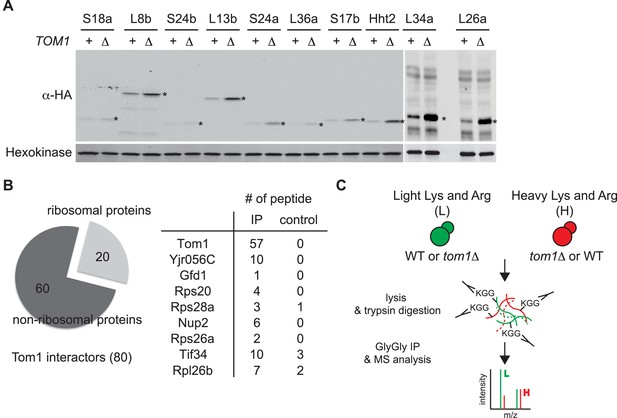
Tom1 targets a broad range of overexpressed and endogenous ribosomal proteins.
(A) Relative levels of the transiently overexpressed, indicated ribosomal proteins (all tagged with a His6-HA-protein A ZZ domain epitope) in WT and tom1∆ mutants. n = 2 biological replicates. *indicates the expected size of protein. (B) Left: Cells expressing untagged-Tom1 or 3×HATom1 were treated with bortezomib for 1 hr. Total cell extracts were adsorbed to HA resin and the bound fractions were analyzed by mass spectrometry. 80 proteins that increased >1.5 fold in 3×HATom1 samples versus untagged samples were categorized into ribosomal or non-ribosomal proteins. Right: The number of peptides derived from the 9 proteins with the highest fold changes in 3×HATom1 vs. untagged samples are shown. n = 1 biological replicate. (C) Schematic diagram of SILAC-based quantitative K-ε-GlyGly mass spectrometry (MS) strategy to identify Tom1-depdendent ubiquitination sites.
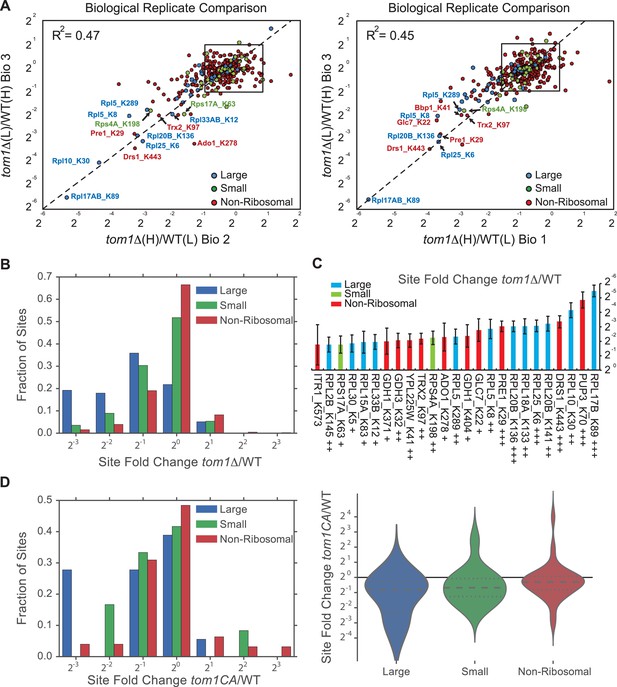
Quantitative GlyGly proteomic analyses of tom1 mutants.
(A) Scatter plots of the SILAC ratios (tom1∆/WT) of biological replicate 3 vs. 2 (left) and biological replicate 3 vs. 1 (right) for GlyGly-modified peptides in tom1∆ and WT cells. Sites with the largest decrease in ubiquitination are annotated. These data accompany Figure 3A. n = 3 biological replicates. (B) Histogram of tom1∆/WT ubiquitination site ratios for ribosomal proteins of the large and small subunits, and non-ribosomal proteins. For each protein category, the fraction of total with a given ratio is plotted. (C) The 25 ubiquitination sites with the largest decrease in ubiquitin occupancy in tom1∆. +++p<0.001; ++p<0.01: +p<0.05. Each site was observed in at least two of the three biological replicates. The error bars represent 95% confidence intervals. (D) Left: same as (B), except that a tom1CA mutant was used instead of tom1∆. Right: violin plot representing the distribution of ubiquitin site occupancy ratios for ribosomal proteins of the large (blue) and small (green) subunit, and non-ribosomal proteins (red). n = 3 biological replicates.
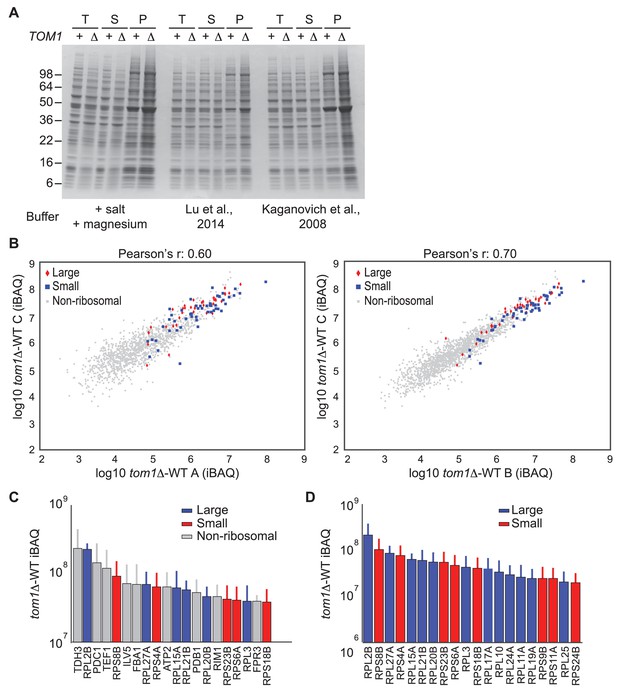
Endogenous ribosomal proteins accumulate as insoluble species in tom1∆ mutants.
(A) Accumulation of insoluble proteins in tom1 mutant cells is independent of lysis method or buffer. Cells of the indicated genotypes were lysed with glass beads in the presence of 3 different lysis buffers as indicated below gel image, and fractionated into detergent-soluble and insoluble fractions. Samples were separated by SDS-PAGE and stained with Coomassie Blue. The pellet fraction is overloaded 10-fold compared to the total and supernatant fractions. n = 2 biological replicates. T, S and P indicates total, soluble and pellet fractions, respectively. Note that results shown here are qualitatively similar to results in Figures 3F and 6D–G, even though the method employed to generate those figures employed lysis of spheroplasts in an EDTA-containing buffer. The reason for the higher background in this panel relative to the others is that the pellet fractions were not washed prior to analysis. (B) Scatter plots representing the ∆iBAQ of biological replicate C vs. A (left) and biological replicate C vs. B (right) for insoluble proteins in tom1∆ mutants. Pearson’s r-value is indicated on top of the plot. These data accompany Figure 3G. n = 3 biological replicates. (C) The 20 proteins with the largest increase in the pellet fraction upon TOM1 deletion. Bars represent the average ∆iBAQ values with error bars indicating the standard error of the mean (SEM). (D) Same as (C), except the top 20 ribosomal proteins are shown.
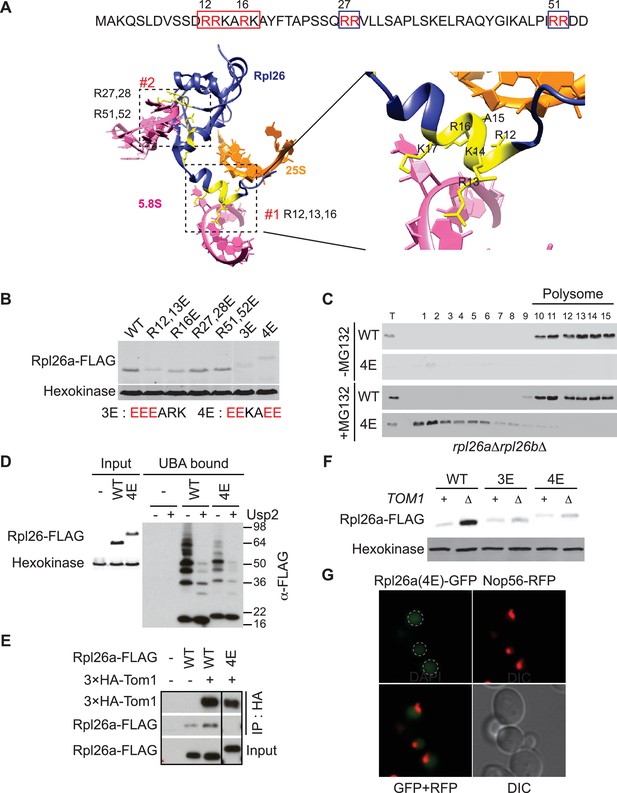
A short stretch of positively-charged residues in Rpl26a that mediates rRNA binding promotes association with Tom1.
(A) Top: The first 54 amino acids of Rpl26a. Sequences adjacent to rRNA are boxed. Arginine residues targeted for mutation are in red. Bottom: Relative positions of arginines and rRNA based on the atomic model of the yeast 80S ribosome (PDB files 3U5D and 3U5E). Orange and pink ribbons correspond to 25S and 5.8S rRNA, respectively. Blue ribbon corresponds to Rpl26. Predicted regions (#1 and #2) for rRNA binding are highlighted in yellow and boxed. (B) Differential accumulation of WT and mutant Rpl26aFLAG upon galactose induction. n = 2 biological replicates. (C) Top: Ribosome assembly of WT Rpl26aFLAG or Rpl26a-4EFLAG induced in rpl26a∆rpl26b∆ cells. Bottom: Same as above except that MG132 was added 30 min after addition of galactose. T indicates total extract. n = 2 biological replicates. (D) Polyubiquitination of Rpl26a-4EFLAG. Assay was performed as described for Figure 1D. Samples in ‘+’ lanes were treated with deubiquitinating enzyme Usp2 prior to processing for SDS-PAGE, to demonstrate that high MW species were modified with ubiquitin. n = 2 biological replicates. (E) The Rpl26-4E mutation disrupts binding to Tom1. Lysates from cells of the indicated genotypes were subjected to pull-down with anti-HA followed by SDS-PAGE and immunoblotting for the indicated proteins. n = 2 biological replicates. (F) Protein level of Rpl26aFLAG mutants upon galactose induction in WT and tom1∆ cells. n = 2 biological replicates. (G) Fluorescence images of Rpl26a4E-GFP induced in WT cells. Nop56-RFP marks nucleoli. Dashed circles indicate nuclear region as judged by DAPI staining. n = 2 biological replicates.
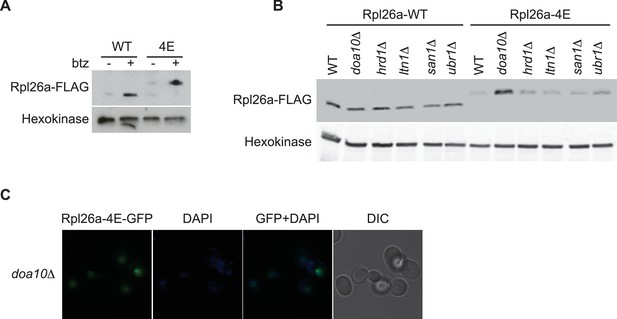
Rpl26a-4E mutant is unstable and degraded by Doa10 in the nucleus/nucleolus.
(A) Similar accumulation of Rpl26aFLAG and Rpl26a-4EFLAG upon galactose induction in the presence of bortezomib. Total cell lysates were evaluated by SDS-PAGE and immunoblotting with the indicated antibodies. n = 1 biological replicate. (B) Differential accumulation of WT and Rpl26a-4E upon galactose induction in WT and known PQC mutants was evaluated by SDS-PAGE and immunoblotting with the indicated antibodies. n = 1 biological replicate. (C) Fluorescence images of Rpl26a-4EGFP induced in doa10∆ cells. n = 2 biological replicates.
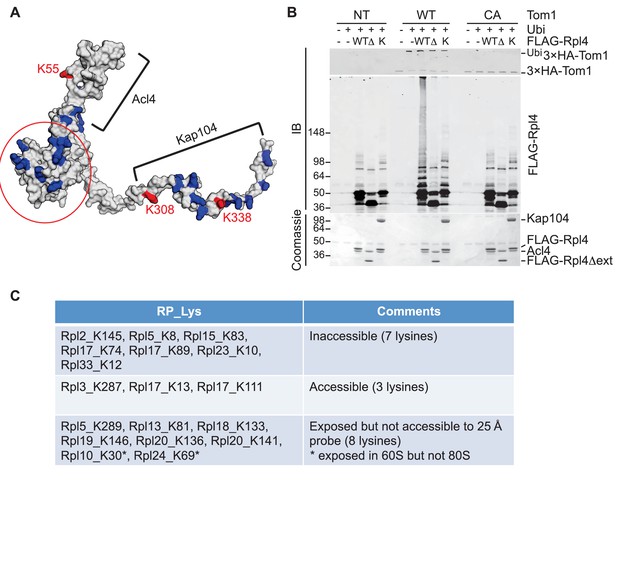
Tom1 acts through residues that are normally inaccessible in the structure of the mature ribosome.
(A) Structure of Rpl4 within the mature ribosome (PDB ID 4V88). Lysine residues are colored blue, with K55, K308, and K338 colored red. Areas involved in binding Acl4 and ctKap104 are indicated. The exact boundaries of the Kap104 binding site are not known. The globular central domain, which is fully exposed in the ternary Acl4–Rpl4–Kap104 complex but is not ubiquitinated, is circled in red. (B) Ubiquitination of Acl4–Rpl4 by 3xHATom1. Anti-HA immunoprecipitates from untagged (NT), 3xHATOM1 (WT), and 3xHAtom1CA (CA) cells were supplemented or not with E1/E2/ubiquitin/ATP (Ubi) and purified Acl4–FLAGRpl4, Acl4-FLAGRpl4∆ext and Acl4–FLAGRpl4–ctKap104 proteins. Samples were analyzed by SDS-PAGE and staining with Coomassie blue or immunoblotting with the indicated antibodies. WT, ∆, and K refer to Acl4–FLAGRpl4, Acl4-FLAGRpl4∆ext and Acl4–FLAGRpl4–ctKap104, respectively. See detailed methods in Material and methods. n = 3 biological replicates. (C) Tom1 preferentially targets lysines that are inaccessible in mature ribosomes. Lysine residues shown are those from large subunit ribosomal proteins in Figure 3E that are incorporated in the model for the structure of the yeast ribosome (pdb 4V88). The structure of a HECT domain–donor ubiquitin complex (pdb 4LCD) predicts that a gap of radius 25 Å must be present for Tom1 to access a lysine for ubiquitination. Two of the sites (Rpl10 K30 and Rpl24 K69) are accessible in the 60S large subunit but become inaccessible upon formation of the 80S ribosome.
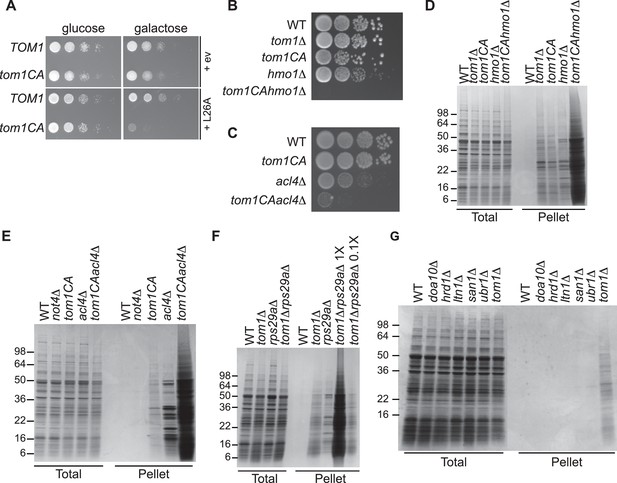
Defective ribosome assembly homeostasis and proteostatic collapse in tom1 mutant cells.
(A–C) Hypersensitivity of tom1CA cells to imbalances in ribosome components. (A) Cells of the indicated genotypes were spotted in serial 10-fold dilutions on glucose or galactose medium and incubated at 30°C for 2 days. ev refers to empty vector. n = 2 biological replicates. (B, C) As in (A) except that cells of the indicated genotypes were spotted on YPD. n = 2 biological replicates. (D–G) Massive accumulation of insoluble proteins in tom1 mutant cells. Cells of the indicated genotypes were lysed and fractionated into detergent-soluble and insoluble fractions, which were separated by SDS-PAGE and stained with Coomassie Blue. The pellet fraction is overloaded 20-fold compared to the total and supernatant fractions. n = 2 biological replicates.
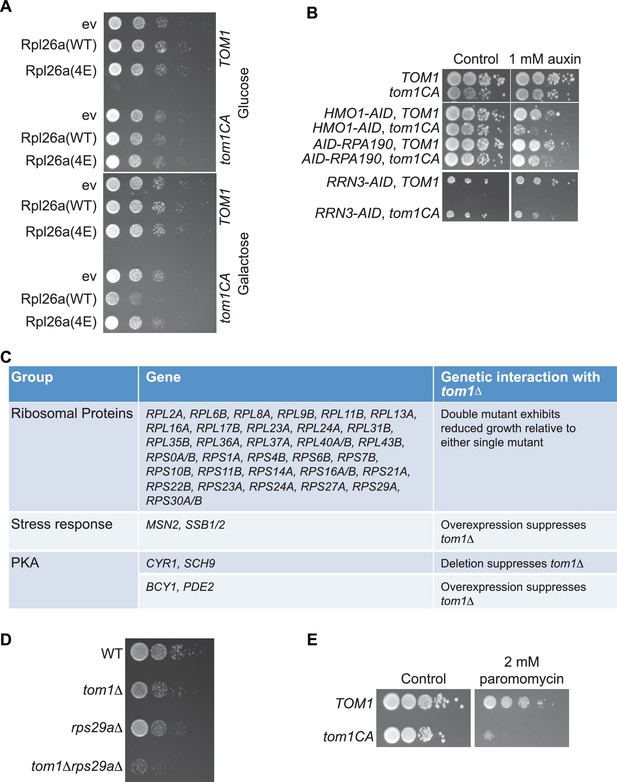
Tom1 is required for maintaining proteostasis.
(A) Cells of the indicated genotypes were spotted in serial 10-fold dilutions on glucose or galactose medium and incubated at 30°C for 2 days. ev refers to empty vector. n = 2 biological replicates. (B) Cells of the indicated genotypes were spotted as serial 10-fold dilutions on YPD with or without 1 mM auxin and incubated at 30°C for 2 days. AID refers to auxin-inducible degron and CA refers to the Cys3235Ala mutation in TOM1. n = 2 biological replicates. (C) List of genetic interactions with tom1∆ as reported in the Saccharomyces Genome Database (www.yeastgenome.org). (D) Synthetic growth defects of rps29a∆tom1∆ double mutants. Cells of the indicated genotypes were spotted in serial 10-fold dilutions on YPD and incubated at 30°C for 2 days. n = 2 biological replicates. (E) same as panel B except medium was supplemented or not with 2 mM paromomycin.
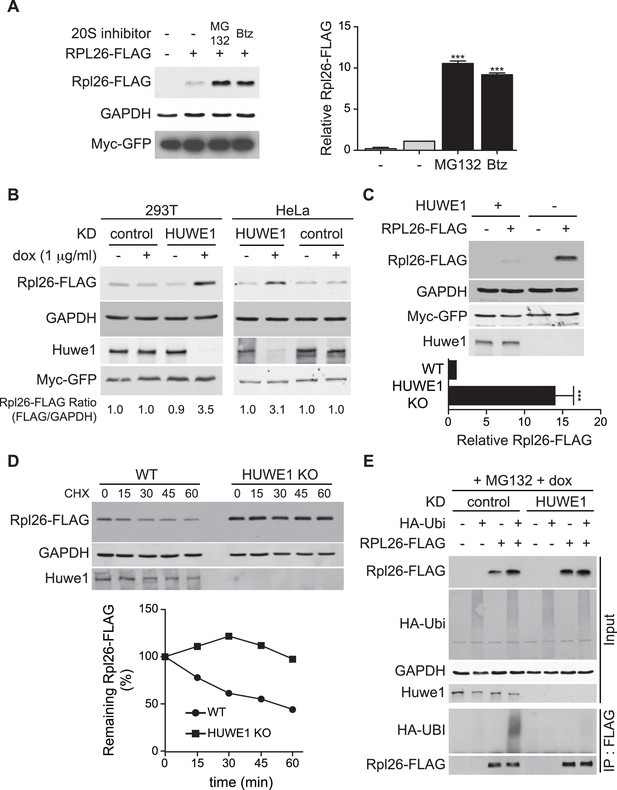
ERISQ is conserved in human cells.
(A) Proteasome inhibition enables overexpression of human Rpl26. Left: transiently expressed hRpl26FLAG in T-REx-293 cells treated with 10 µM MG132 or 1 µM bortezomib (btz) for 3 hr. Right: quantification of blots. Values are the mean of three independent experiments and error bars indicate standard deviations. Asterisks indicate significant differences (two-tailed student's t-test, ***p<0.0001, compared with DMSO treatment). n = 3 biological replicates. Source data are available in Figure 7—source data 1. (B) Depletion of HUWE1 enables overexpression of human Rpl26. As in (A) except that T-REx-293 (left) and HeLa (right) cells were induced with doxycycline for 3 days to express stably integrated shControl or shHUWE1. The relative ratio of hRpl26FLAG/GAPDH is shown below each lane. n = 3 biological replicates. Source data are available in Figure 7—source data 2. (C) Knockout of HUWE1 enables overexpression of human Rpl26. Upper: as in (A) except that wild type and HUWE1 knockout HEK293T cells were used. Bottom: quantification of blots. Values are the mean of three independent experiments and error bars indicate standard deviations. Asterisks indicate significant differences (two-tailed student's t-test, ***p<0.0001, compared with WT cells). n = 3 biological replicates. Source data are available in Figure 7—source data 3. (D) Overexpressed human Rpl26 is stable in HUWE1 knockout cells. Upper: wild type and HUWE1 knockout HEK293T cells transiently expressing hRpl26FLAG were treated with 10 μg/ml cycloheximide (CHX). Bottom: quantification of blot. n = 1 biological replicate. Source data are available in Figure 7—source data 4. (E) HUWE1 promotes ubiquitination of overexpressed human Rpl26. As in (B) except that HAubiquitin was co-expressed with Rpl26FLAG and MG132 was added 3 hr prior to cell lysis. Total cell extract prepared under denaturing condition was adsorbed to FLAG resin and the bound fraction was immunoblotted with antibodies against FLAG and HA. n = 2 biological replicates.
-
Figure 7—source data 1
Quantification of hRpl26-FLAG and GAPDH levels from three biological replicates.
- https://doi.org/10.7554/eLife.19105.018
-
Figure 7—source data 2
Quantification of hRpl26-FLAG and GAPDH levels from one biological replicate.
- https://doi.org/10.7554/eLife.19105.019
-
Figure 7—source data 3
Quantification of hRpl26-FLAG and GAPDH levels from three biological replicates.
- https://doi.org/10.7554/eLife.19105.020
-
Figure 7—source data 4
Quantification of hRpl26-FLAG and GAPDH levels from one biological replicate.
- https://doi.org/10.7554/eLife.19105.021
Additional files
-
Supplementary file 1
Yeast strains used in this study.
- https://doi.org/10.7554/eLife.19105.022
-
Supplementary file 2
Plasmids used in this study.
- https://doi.org/10.7554/eLife.19105.023
-
Supplementary file 3
(A) Tom1 interactors identified by MS. (B) Dataset from SILAC-GlyGly analysis. (C) Insoluble proteins identified in tom1 mutants.
- https://doi.org/10.7554/eLife.19105.024





