CDK-regulated dimerization of M18BP1 on a Mis18 hexamer is necessary for CENP-A loading
Figures
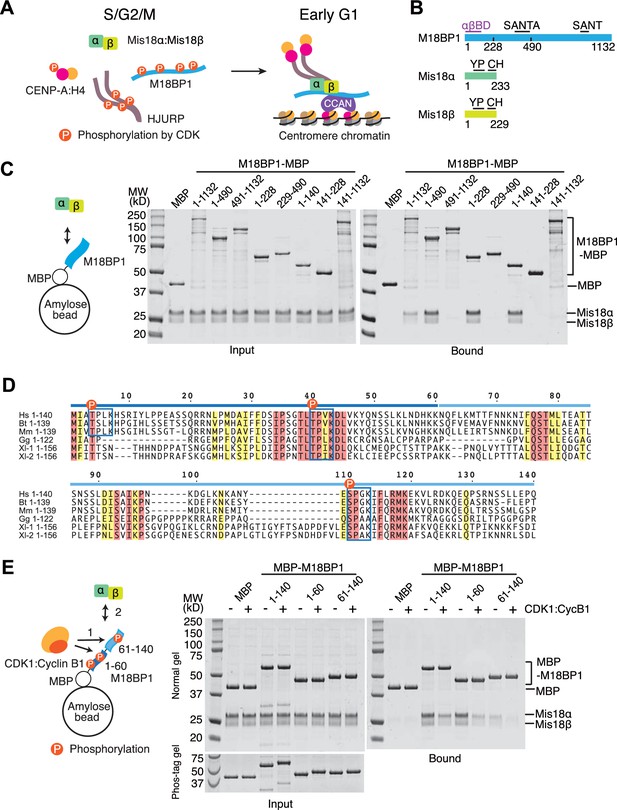
CDK1 regulates binding of M18BP11-140 and Mis18α:Mis18β.
(A) Simplified current model of the cell-cycle-dependent assembly of the CENP-A deposition machinery. CDKs phosphorylate CENP-A, HJURP, and M18BP1 to prevent the assembly of the protein complex during the S, G2, and M phases of the cell cycle. Reduced CDK activity in early G1 phase allows the CENP-A deposition machinery to interact and localize to centromeres through the interaction with subunits of CCAN. (B) Schematic diagrams of domain structures of M18BP1, Mis18α, and Mis18β. YP, Yippee domain; CH, C-terminal helix; αβBD, Mis18α:Mis18β binding domain. (C) Amylose-resin pull-down assays to identify the Mis18α:Mis18β-binding domain of M18BP1. M18BP1-MBP variants and MBP were produced in bacteria and Mis18α:Mis18β was produced using baculovirus co-expression system. Shown is a representative gel of pull-down assays that were repeated at least three times. The same applies to all other pull-down assays in this paper. (D) Sequence alignment of M18BP11-140. Hs, Homo sapiens; Mm, Mus musculus; Bt, Bos taurus; Gg, Gallus gallus; Xl-1, Xenopus laevis isoform 1; Xl-2, Xenopus laevis isoform 2. Blue-boxed regions indicate putative CDK phosphorylation motifs. Residues that are identical in all sequences are shaded red, and residues that only have conserved substitutions are shaded yellow. (E) Amylose-resin pull-down assays examining how phosphorylation affects complex formation. MBP-M18BP1 variants were incubated with or without CDK1:Cyclin B1 at 30°C for 2 hr before mixing with Mis18α:Mis18β. The Phos-tag containing acrylamide gel was used to detect the mobility shift caused by phosphorylation. Gels in panels C and E were stained with CBB.
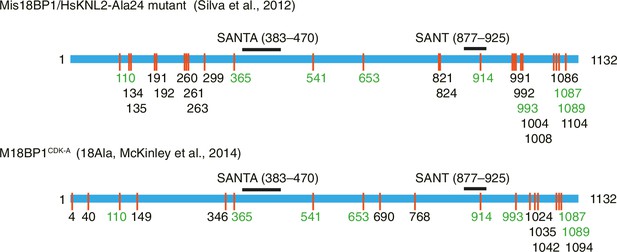
Previously identified phosphorylation sites on M18BP1.
https://doi.org/10.7554/eLife.23352.003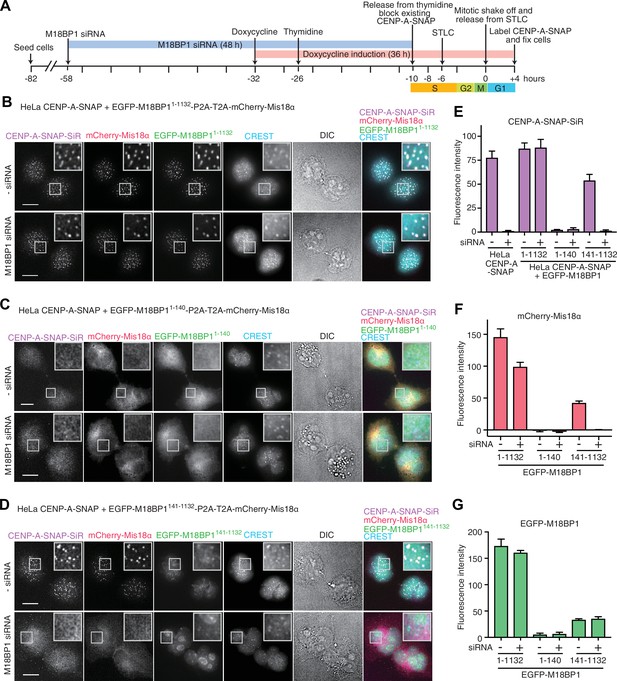
Functional analysis of M18BP11–140 and M18BP1141–1132 in CENP-A recruitment.
(A) Schematic description of the experimental procedure for testing the functionality of M18BP1 variants for new CENP-A deposition and the recruitment of Mis18α to centromeres. SNAP-Cell Block was used to block existing CENP-A-SNAP proteins at the time point −10 hr, and SNAP-Cell 647-SiR was used to label newly produced CENP-A-SNAP at the time point +4 hr. (B–D) Representative images showing the fluorescence of CENP-A-SNAP labeled with SNAP-Cell 647-SiR (CENP-A-SNAP-SiR), mCherry-Mis18α and EGFP-M18BP1 variants in fixed HeLa cells treated as described in panel A. Centromeres were visualized with CREST sera. Control cells were treated with transfection reagent (Lipofectamine RNAiMAX) in the absence of M18BP1 siRNA. Scale bars represent 10 µm. DIC, differential interference contrast. All cell biological experiments in this paper were repeated at least three times. (E) Quantification of the fluorescence intensity of CENP-A-SNAP-SiR on each centromere. Centromere spots were detected using images of CREST channel with the software Fiji (Schindelin et al., 2012). Mean intensity of CENP-A-SNAP-SiR fluorescence on centromeres was obtained from every experiment (>100 centromere spots from 10–12 early G1 cells). The bar graph presents mean values from three independent experiments. Error bars indicate SEM. (F, G) Quantification of the fluorescence intensity of mCherry-Mis18α and EGFP-M18BP1 variants on each centromere.
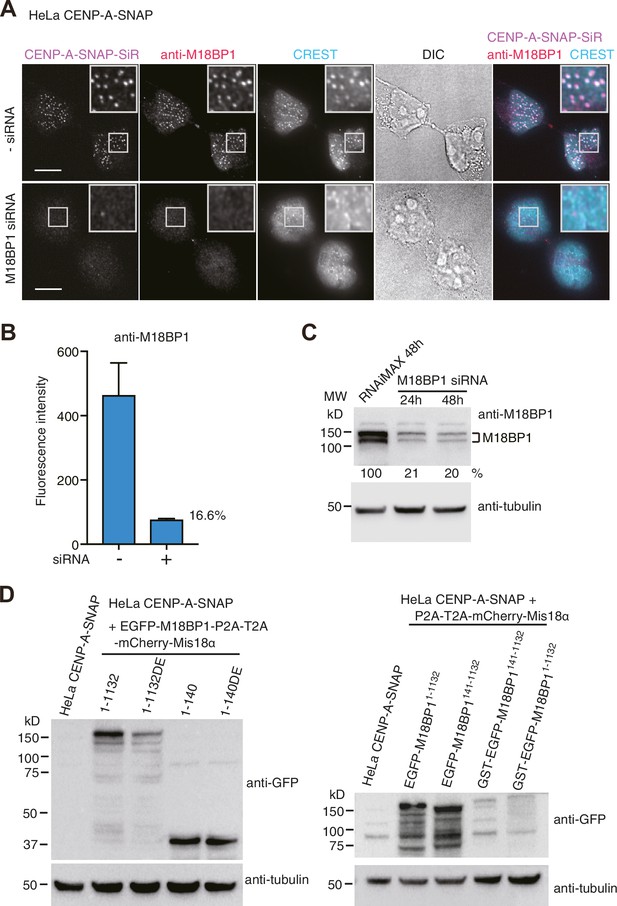
HeLa CENP-A-SNAP cell lines.
(A) Representative images showing the fluorescence of CENP-A-SNAP-SiR and the immunoflurorescence of M18BP1 in fixed HeLa cells treated as described in Figure 2A. Centromeres were visualized with CREST sera. Scale bars represent 10 µm. (B) Quantification of the immunofluorescence intensity of M18BP1 on each centromere. The mean intensity value of M18BP1 immunofluorescence of centromere spots from three independent experiments is presented (>100 centromeres from 10 early G1 cells were quantified in one experiment). Error bars represent SEM (standard error of the mean). Centromeric M18BP1 level of the siRNA-treated cells was reduced to 16.6% of the level of the untreated cells. (C) Cell lysate (containing 50 µg protein) of HeLa CENP-A-SNAP cells treated with M18BP1 siRNA (Fujita et al., 2007) for 24 or 48 hr and the lysate of the control cells were analyzed using Western blotting with anti-M18BP1 antibody (Bethyl, A302-825A), anti-alpha-tubulin antibody (Sigma, T9026), and HRP-conjugated secondary antibodies (Amersham NXA931-1ML, NXA934-1ML). The intensity of protein bands corresponding to M18BP1 and tubulin were quantified from 16-bit image using the software Fiji. The comparison of the intensity ratio of M18BP1/tubulin are shown as percentages. Cellular M18BP1 level of the siRNA-treated cells was reduced to 20% of the level of the untreated cells after 48 hr siRNA treatment. (D) Cell lysate (containing 100 μg protei) of each HeLa cell line treated with 50 ng/ml doxycycline for 36 hr was analyzed using Western blotting with anti-GFP antibody (Abcam, AB6556), anti-alpha-tubulin antibody (Sigma, T9026), and HRP-conjugated secondary antibodies (Amersham NXA931-1ML, NXA934-1ML). DE, T40D/S110E.
Tandem 2A-peptide system enables efficient co-expression of multiple proteins in HeLa cells.
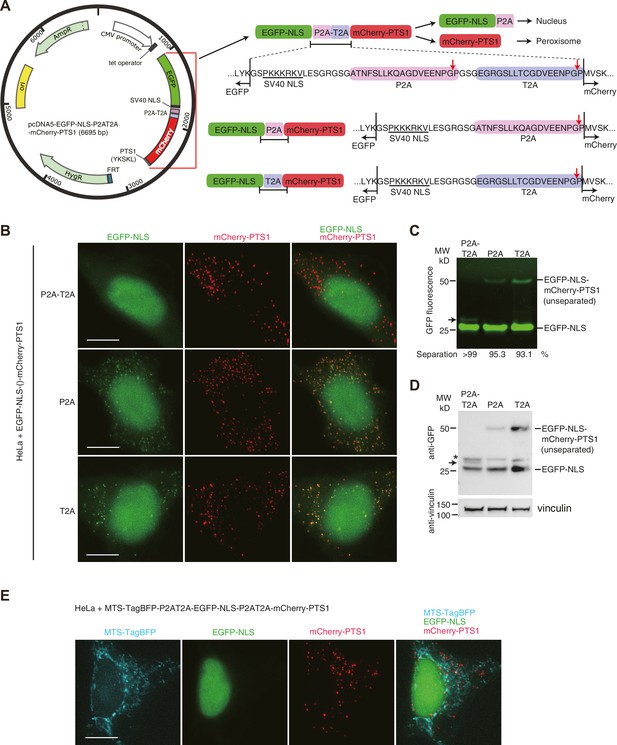
(A) Schematic presentation showing the design of the co-expression vectors using 2A-peptides. The protein expression from the single ORF with the continuous coding sequence of EGFP-NLS (nuclear localization signal, PKKKRKV) (Fontes et al., 2000), P2A-T2A (or P2A, T2A alone), and mCherry-PTS1 (peroxisomal targeting signal–1, YKSKL) (Maynard et al., 2004) is regulated by CMV promoter and tetracycline operator. On translating the ORF sequence, ribosomes skip the synthesis of the glycyl-prolyl peptide bond at the C-terminus of a 2A peptide (indicated by red arrows) (de Felipe et al., 2010; Donnelly et al., 2001), which results in the separation of EGFP-NLS and mCherry-PTS1. Eighteen amino acids from P2A remain at the C-terminus of EGFP-NLS. Separated EGFP-NLS is expected to accumulate in the nucleus and mCherry-PTS1 in peroxisomes. (B) Representative images showing the fluorescence of EGFP-NLS and mCherry-PTS1 in fixed HeLa cells. HeLa cells were grown in DMEM supplemented with 10% tetracycline-free FBS and 2 mM L-glutamine at 37°C and treated with 50 ng/ml doxycycline for 24 hr for inducing protein expression. Green fluorescence of EGFP-NLS in peroxisomes indicates the presence of unseparated EGFP-NLS-mCherry-PTS1 product. (C) Cell lysates (containing 50 µg protein) of the HeLa cells expressing EGFP-NLS-P2AT2A-mCherry-PTS1 (or single P2A, T2A variants) were analyzed using Tricine–SDS-PAGE and imaged by EGFP fluorescence. Unseparated EGFP-NLS-mCherry-PTS1 product was detected only in the sample with single 2A-peptides. The 16-bit image was analyzed using the software Fiji to quantify the intensity of the bands. Separation efficiency = 100 × separated form/(separated form + unseparated form). (D) Western blot analyses of the same samples used in the experiment of panel C. The membrane with transferred proteins was cut into two pieces and immunoblotted using anti-GFP antibody (Abcam, AB6556), anti-vinculin antibody (Sigma, V9131), and HRP-conjugated secondary antibodies (Amersham NXA931-1ML, NXA934-1ML). The asterisk indicates unknown protein bands detected by the anti-GFP antibody. Black arrows in (C) and (D) indicate putative product of EGFP-NLS-P2A-T2A arisen from failure of separation at the cleavage site in P2A but not in T2A. (E) Representative images showing the fluorescence of MTS-TagBFP, EGFP-NLS and mCherry-PTS1 in the fixed HeLa cells. Twenty-nine-amino-acid mitochondrial targeting signal (MTS) from cytochrome c oxidase subunit 8A (Rizzuto et al., 1995) was fused to TagBFP and the fusion protein MTS-TagBFP was co-expressed with EGFP-NLS and mCherry-PTS1 using two P2A-T2A tandem peptides. Each fluorescent protein distributed in different cellular compartments without mislocalization.
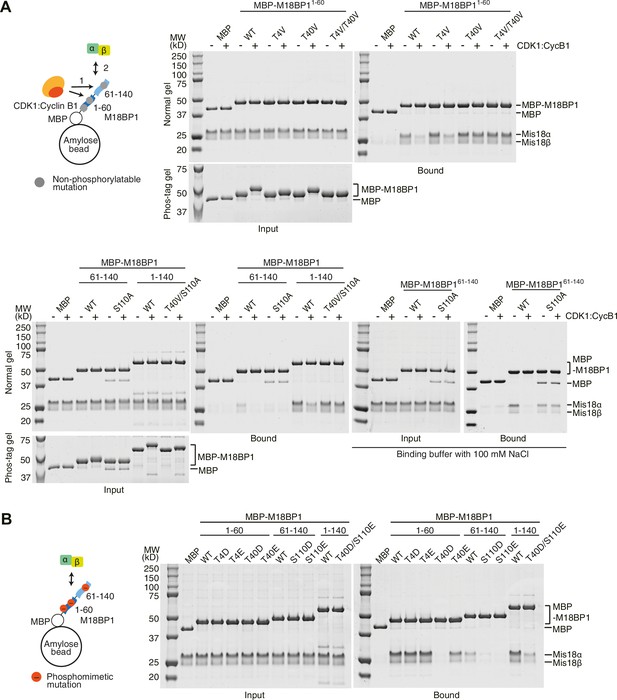
Phosphorylation on M18BP1 Thr40 and Ser110 reduces affinity for Mis18α:Mis18β.
(A, B) Amylose-resin pull-down assays examining complex formation of Mis18α:Mis18β with non-phosphorylatable (panel A) or phosphomimetic (panel B) M18BP1 mutants. The experiments were performed as in Figure 1C,E. Binding buffer with 300 mM NaCl was used, unless indicated. WT, wild type.
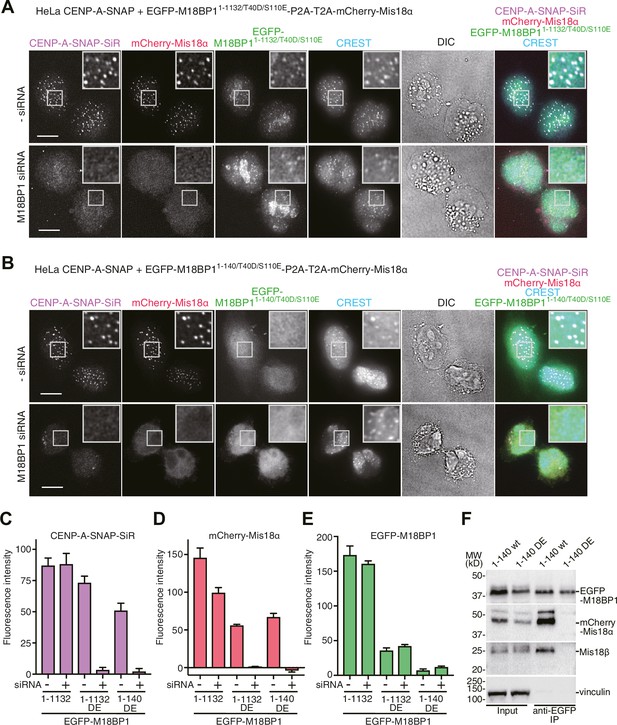
Functional analyses of M18BP1 T40D/S110E mutants in CENP-A recruitment.
(A, B) Representative images showing the fluorescence of CENP-A-SNAP-SiR, mCherry-Mis18α, and EGFP-M18BP1 variants in fixed HeLa cells treated as described in Figure 2A. Centromeres were visualized with CREST sera. Scale bars represent 10 µm. (C–E) Quantification of the fluorescence intensity of CENP-A-SNAP-SiR (panel C), mCherry-Mis18α (panel D), or EGFP-M18BP1 variants (panel E) on each centromere. The quantification was performed and is presented in the same way described in the legend of Figure 2E. DE, T40D/S110E. (F) Western blots of co-immunoprecipitation experiments using GFP-Trap_A beads. HeLa CENP-A-SNAP + EGFP-M18BP11-140-P2A-T2A-mCherry-Mis18α or EGFP-M18BP11-140/T40D/S110E-P2A-T2A-mCherry-Mis18α were analyzed.
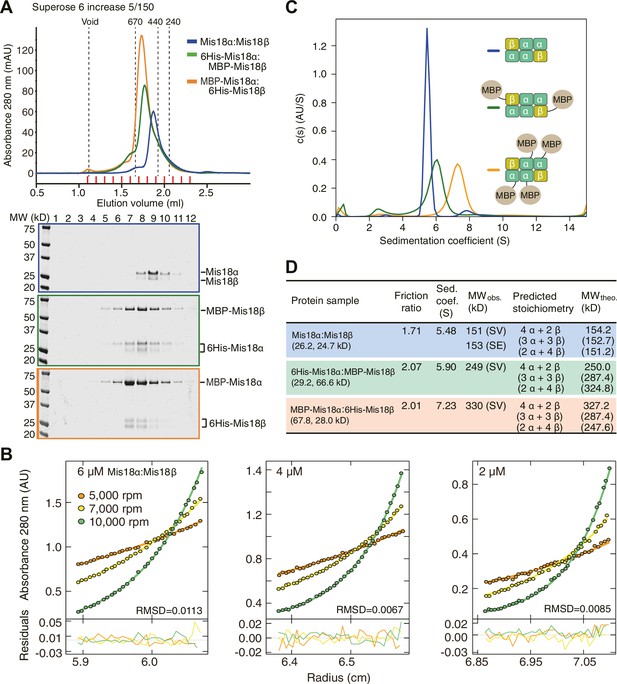
Determination of the stoichiometry of Mis18α:Mis18β complex.
(A) Analytical SEC results of Mis18α:Mis18β (blue), 6His-Mis18α:MBP-Mis18β (green), MBP-Mis18α:6His-Mis18β (orange). The elution volumes of thyroglobulin (670 kD), ferritin (440 kD), and catalase (240 kD) are shown as standards. Red lines indicate fractions collected for Tricine–SDS-PAGE analyses. Gels were stained with CBB. (B) Sedimentation equilibrium AUC analysis of Mis18α:Mis18β. Results of data fitting of three datasets obtained at different protein concentrations are shown. Using the global fitting method of SEDPHAT together with four additional datasets (Figure 5—figure supplement 1A–B), we obtained the experimental MW of 153 kD. The values of root-mean-square deviation (RMSD) for data fitting are shown. (C) Sedimentation velocity AUC results of the same samples used in the analytical SEC experiments (panel A). The best-fit size distributions are shown with the colors indicated in panel A. Data profiles used for curve-fitting analyses are shown in Figure 5—figure supplement 1C. (D) Summary table of the results obtained from the AUC experiments of panel B and C. Sed. coef., sedimentation coefficient; MWobs., observed molecular weight; MWtheo., theoretical molecular weight; SV, sedimentation velocity; SE, sedimentation equilibrium.
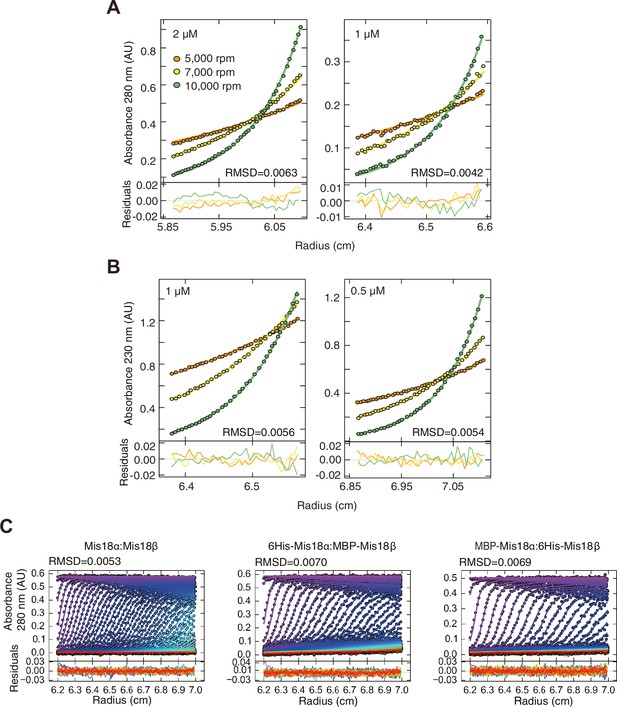
Data profiles for AUC experiments.
(A) Sedimentation equilibrium AUC data of Mis18α:Mis18β measured at 280 nm at the concentrations of 2 and 1 µM. (B) Sedimentation equilibrium AUC data of Mis18α:Mis18β measured at 230 nm at the concentrations of 1 and 0.5 µM. (C) Best-fitting results of the sedimentation velocity AUC data of Mis18α:Mis18β, 6His-Mis18α:MBP-Mis18β and MBP-Mis18α:6His-Mis18β. Residuals represent the deviation of the continuous c(s) distribution model from the observed signals. The values of root-mean-square deviation (RMSD) for data fitting are shown.
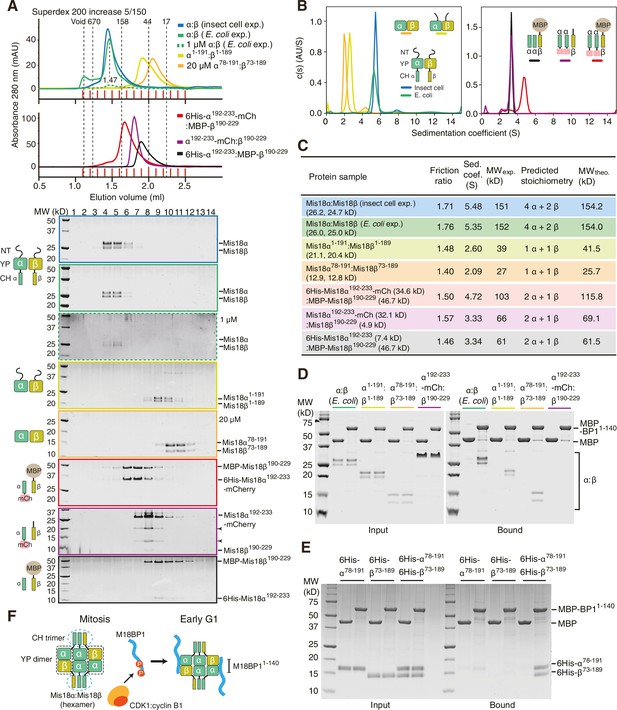
Assembly mechanism of Mis18α:Mis18β-hexamer.
(A) Analytical SEC results of insect-cell-expressed Mis18α:Mis18β (blue), E. coli-expressed Mis18α:Mis18β (green), Mis18α1-191:Mis18β1-189 (yellow-green), Mis18α78-191:Mis18β73-189 (orange), 6His-Mis18α192-233-mCherry:Mis18β190-229 (red), Mis18α192-233-mCherry:Mis18β190-229 (purple), 6His-Mis18α192-233:Mis18β190-229-MBP (black). SEC experiments were carried out with 10 µM protein (loading concentration), unless indicated in the figure. The elution volumes of thyroglobulin (670 kD), aldolase (158 kD), ovalbumin (44 kD), and myoglobin (17 kD) are shown as standards. Red lines indicate fractions collected for Tricine–SDS-PAGE analyses. Gels were stained with CBB, except the gel for 1 µM Mis18α:Mis18β (dashed green line), which was stained with SYPRO Ruby. Left-pointing arrowheads indicate degradation products of Mis18α192-233-mCherry. NT, N-terminal tail; YP, Yippee domain; CH, C-terminal helix; mCh, mCherry. (B) Sedimentation velocity AUC results of the same samples used in the analytical SEC experiments (panel A). The best-fit size distributions are shown with the colors indicated in panel A. Data profiles used for curve-fitting analyses are shown in Figure 6—figure supplement 1. (C) Summary table of the results obtained from the AUC experiments of panel B. Sed. coef., sedimentation coefficient; MWobs., observed molecular weight; MWtheo., theoretical molecular weight. (D, E) Amylose-resin pull-down assays to examine the interaction of Mis18α:Mis18β variants with M18BP11-140. Incubation of amylose beads and proteins (at 5 µM concentration) were performed using a binding buffer containing 30 mM HEPES pH 7.5, 100 mM NaCl, and 1 mM TCEP. Gels were stained with CBB. (F) Hypothetical assembly mechanism of the Mis18α:Mis18β-hexamer in mitosis and the octameric Mis18 complex in the early G1 phase.
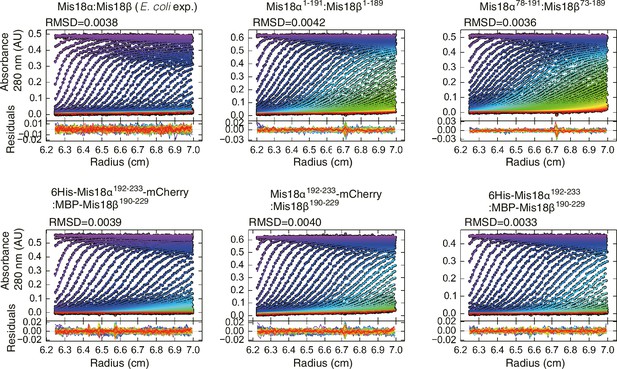
Data profiles for AUC experiments.
Best-fitting results of the sedimentation velocity AUC data of E. coli-expressed Mis18α:Mis18β, Mis18α1–191:Mis18β1–189, Mis18α78–191:Mis18β73–189, 6His-Mis18α192–233-mCherry:MBP-Mis18β190–229, Mis18α192–233-mCherry:Mis18β190–229, Mis18α192–233:MBP-Mis18β190–229. Residuals represent the deviation of the continuous c(s) distribution model from the observed signals. The values of RMSD for data fitting are shown.
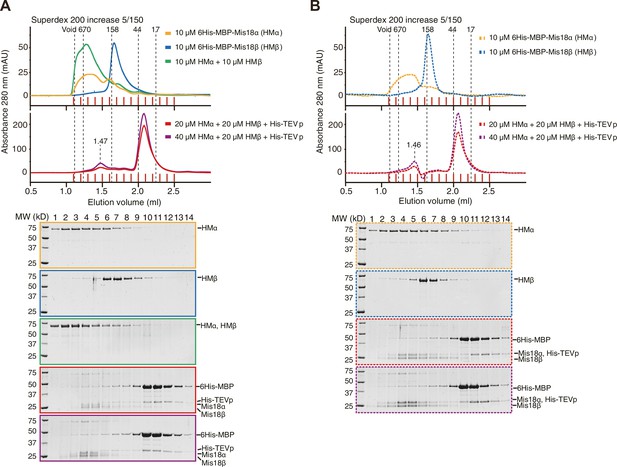
Generation of Mis18α:Mis18β complex by mixing 6His-MBP-Mis18α and 6His-MBP-Mis18β.
(A) Analytical SEC results of 6His-MBP-Mis18α (orange), 6His-MBP-Mis18β (blue), 6His-MBP-Mis18α + 6His-MBP-Mis18β (green), and 6His-MBP-Mis18α + 6His-MBP-Mis18β + His-tagged TEV protease (His-TEVp) incubated at 4°C for 16 hr (red and purple). Experiments were done using the protein concentrations indicated in the figure with a SEC buffer containing 30 mM HEPES pH 7.5, 300 mM NaCl, and 1 mM TCEP. The elution volumes of thyroglobulin (670 kD), aldolase (158 kD), ovalbumin (44 kD), and myoglobin (17 kD) are shown as standards. Red vertical lines indicate fractions collected for Tricine–SDS-PAGE analyses. Gels were stained with CBB. (B) Analytical SEC results of repeated experiments in (A) using the SEC buffer from previous report (Nardi et al., 2016) containing 50 mM Tris-HCl pH 7.5, 350 mM NaCl, 2.5% glycerol, 0.05% NP-40, and 5 mM β-mercaptoethanol.
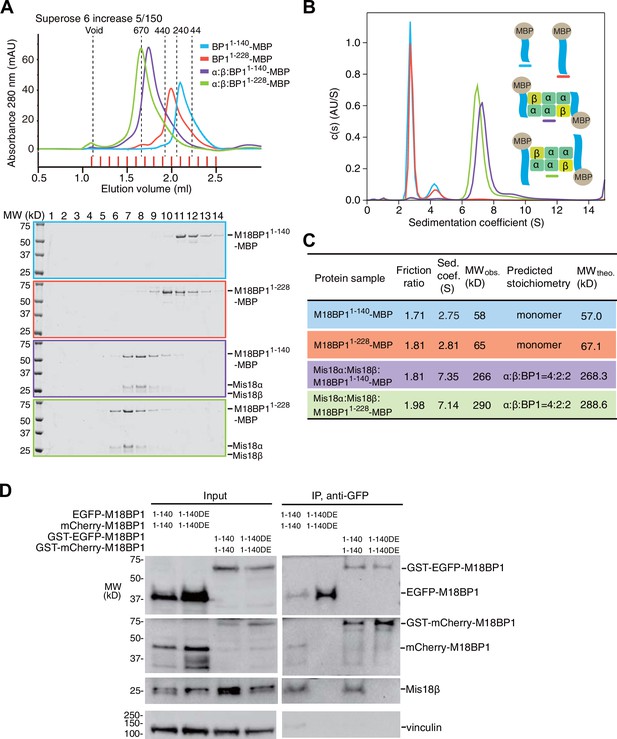
Mis18α:Mis18β-hexamer mediates dimerization of M18BP1.
(A) Analytical SEC results of M18BP11–140-MBP (cyan), M18BP11–228-MBP (red), Mis18α:Mis18β:M18BP11–140-MBP (purple), Mis18α:Mis18β:M18BP11–228-MBP (green). The elution volumes of thyroglobulin (670 kD), ferritin (440 kD), catalase (240 kD) and ovalbumin (44 kD) are shown as standards. Red lines indicate fractions collected for Tricine–SDS-PAGE analyses. Gels were stained with CBB. (B) Sedimentation velocity AUC results of the same samples used in the analytical SEC experiments (panel A). The best-fit size distributions are shown with the colors indicated in panel A. Data profiles used for curve-fitting analyses are shown in Figure 7—figure supplement 1. (C) Summary table of the results obtained from the AUC experiments of panel B. Sed. coef., sedimentation coefficient; MWobs., observed molecular weight; MWtheo., theoretical molecular weight. (D) Western blot results of co-immunoprecipitation experiments using GFP-Trap_A beads. HeLa CENP-A-SNAP + EGFP-M18BP11–140-P2A-T2A-mCherry-M18BP11–140, EGFP-M18BP11–140/T40D/S110E-P2A-T2A-mCherry-M18BP11–140/T40D/S110E, GST-EGFP-M18BP11–140-P2A-T2A-GST-mCherry-M18BP11–140, or GST-EGFP-M18BP11–140/T40D/S110E-P2A-T2A-GST-mCherry-M18BP11–140/T40D/S110E were analyzed.
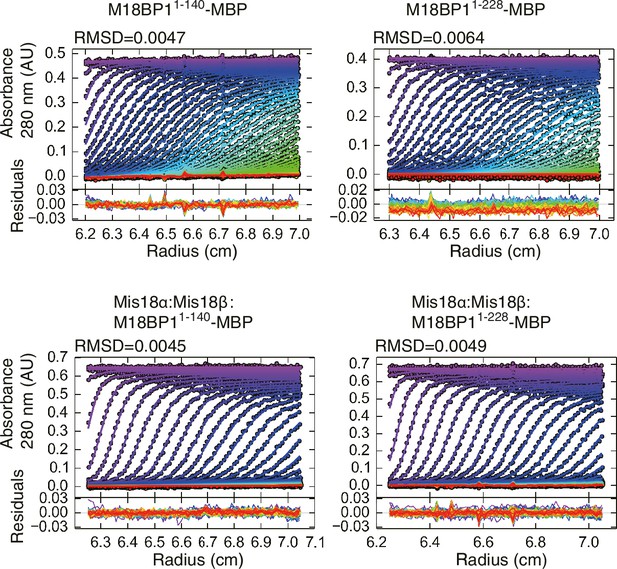
Data profiles for AUC experiments.
Best-fitting results of the sedimentation velocity AUC data of M18BP11–140-MBP, M18BP11–228-MBP, Mis18α:Mis18β:M18BP11–140-MBP, and Mis18α:Mis18β:M18BP11–228-MBP. Residuals represent the deviation of the continuous c(s) distribution model from the observed signals. The values of RMSD for data fitting are shown.
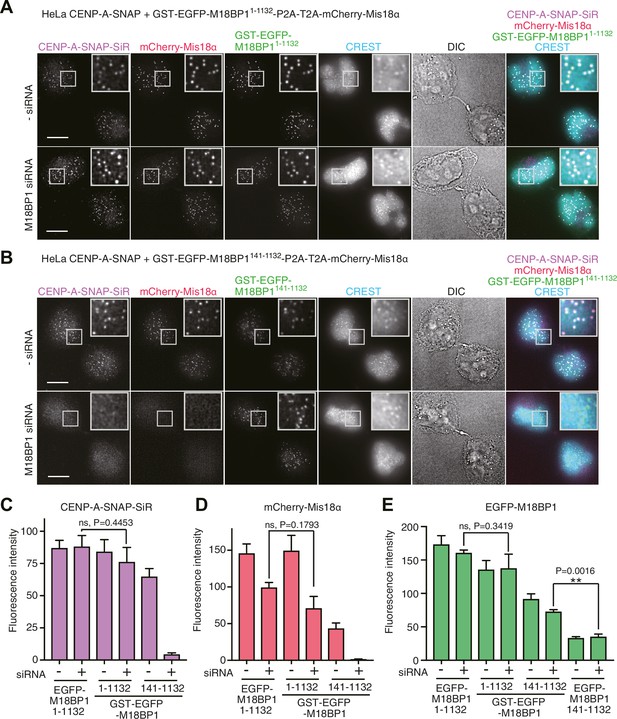
Oligomerization of Mis18 complex promotes robust centromere localization.
(A, B) Representative images showing the fluorescence of CENP-A-SNAP-SiR, mCherry-Mis18α and GST-EGFP-M18BP1 in the fixed HeLa cells treated as described in Figure 2A. Centromeres were visualized with CREST sera. Scale bars represent 10 µm. (C–E) Quantification of the fluorescence intensity of CENP-A-SNAP-SiR (panel C), mCherry-Mis18α (panel D), or EGFP-M18BP1 variants (panel E) on each centromere. The quantification was performed and is presented in the same way described in the legend of Figure 2. P-Values are indicated on the graph (Student’s t test).
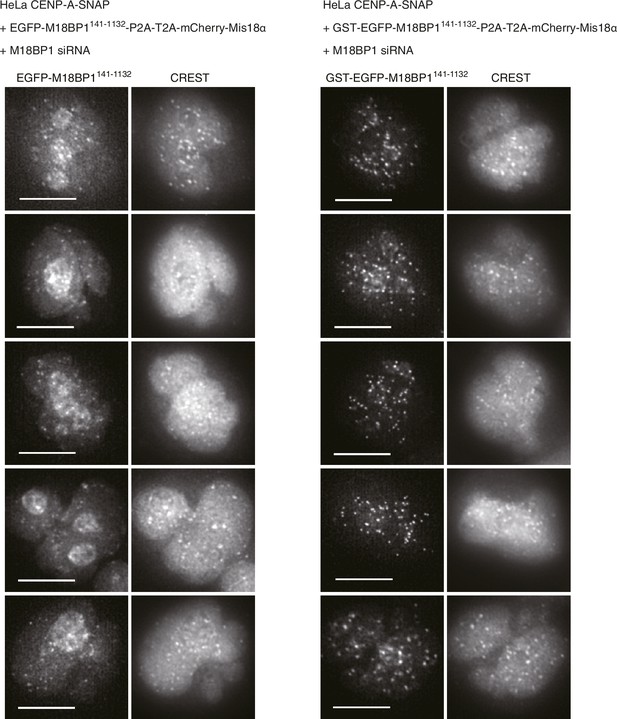
GST-EGFP-M18BP1141-1132 localizes more specifically to centromeres than EGFP-M18BP1141-1132.
Representative images showing EGFP-M18BP1 fluorescence and the immunofluorescence of CREST sera of indicated HeLa cell lines treated as described in Figure 2A. Scale bars represent 10 µm.
Additional files
-
Supplementary file 1
Plasmid vectors used in this study.
- https://doi.org/10.7554/eLife.23352.018
-
Supplementary file 2
HeLa cell lines used in this study.
- https://doi.org/10.7554/eLife.23352.019
-
Supplementary file 3
Fluorescence intensity data.
- https://doi.org/10.7554/eLife.23352.020





