Polo-like kinase Cdc5 regulates Spc72 recruitment to spindle pole body in the methylotrophic yeast Ogataea polymorpha
Abstract
Cytoplasmic microtubules (cMT) control mitotic spindle positioning in many organisms, and are therefore pivotal for successful cell division. Despite its importance, the temporal control of cMT formation remains poorly understood. Here we show that unlike the best-studied yeast Saccharomyces cerevisiae, position of pre-anaphase nucleus is not strongly biased toward bud neck in Ogataea polymorpha and the regulation of spindle positioning becomes active only shortly before anaphase. This is likely due to the unstable property of cMTs compared to those in S. cerevisiae. Furthermore, we show that cMT nucleation/anchoring is restricted at the level of recruitment of the γ-tubulin complex receptor, Spc72, to spindle pole body (SPB), which is regulated by the polo-like kinase Cdc5. Additionally, electron microscopy revealed that the cytoplasmic side of SPB is structurally different between G1 and anaphase. Thus, polo-like kinase dependent recruitment of γ-tubulin receptor to SPBs determines the timing of spindle orientation in O. polymorpha.
https://doi.org/10.7554/eLife.24340.001eLife digest
Before a cell divides, it needs to duplicate its genetic material to provide the new daughter cell with a full set of genetic information. To do so, the cell forms a complex of proteins called the spindle apparatus, which is made up of string-like microtubules that divide the chromosomes evenly. In many organisms, the position of the spindle determines where in the cell this separation happens.
However, in baker’s yeast, the location where the cell will divide is determined well before the spindle is formed. Unlike many other eukaryotic cells, these yeast cells divide asymmetrically and create buds that will form the new daughter cells. The position of this bud determines where the spindle should be located and where the chromosomes separate.
The spindle itself is then organised by a structure called the spindle pole body, which connects to microtubules inside the cell nucleus and microtubules in the cell plasma. Several proteins control where and how the spindle forms, including a protein called the spindle pole component 72, or Spc72 for short, and an enzyme called Cdc5. However, until now it was unclear how spindle formation is timed and controlled in other yeast species.
Now, Maekawa et al. have used fluorescent markers and time lapse microscopy to examine how the spindle forms in the yeast species Ogataea polymorpha, an important industrial yeast used to produce medicines and alcohol. The results show that in O. polymorpha, the positioning and orientation of the spindle only occurred very late in the cell cycle and the microtubules in the cell plasma remained unstable until the chromosomes were about to separate. This was linked to changes in the level of Spc72, which increased at the spindle pole body before the chromosomes separated and then dropped again. This was controlled by Cdc5.
Understanding when and where microtubules are formed is an important step in understanding how cells divide. This is the first example of a budding yeast that creates new microtubules in the cell plasma every time the cell divides. Unravelling the molecular differences between yeast species could lead to new ways to optimise the use of industrial yeasts like O. polymorpha, or to combat disease-causing ones.
https://doi.org/10.7554/eLife.24340.002Introduction
Segregation of sister chromatids into two daughter cells is pivotal to the proliferation of eukaryotic cells. Chromosome segregation is followed by cytokinesis, which results in physical separation of two daughter cells. In many organisms, the position of the mitotic spindle dictates the site of cytokinesis, which ensures the inheritance and maintenance of genomic information in the daughter cells. Astral microtubules or cytoplasmic microtubules (cMTs), which emanate from the spindle poles and extend to the cell cortex, have a principle role in positioning and orienting the spindle with respect to the polarity cues of the cell type. Mechanisms governing the spindle positioning/orientation have been studied in a number of systems. However, regulations that determine the timing of establishing the spindle orientation, or the position of the centrosome, the primary MT organizing centre (MTOC), in interphase, are not well understood (Kiyomitsu, 2015; Woyke et al., 2002).
Spindle positioning is of particular importance in the budding yeast Saccharomyces cerevisiae, where the cleavage site is determined at the start of the cell cycle independently of the position of the mitotic spindle. Therefore, cells position the pre-anaphase spindle close to the bud neck and orient it along the mother-bud axis. As the spindle elongates in anaphase, one spindle pole translocates into the bud to accomplish segregation of one set of chromosomes into the daughter cell (Pereira and Yamashita, 2011; Markus et al., 2012; Winey and Bloom, 2012).
In S. cerevisiae, the nuclear positioning and spindle orientation are regulated by two redundant pathways acting on cMTs, the Kar9 and dynein pathways (Li et al., 1993; Miller and Rose, 1998; Winey and Bloom, 2012). Concomitant deletions in components of both pathways result in lethality, whereas loss of one pathway can be compensated by the function of the other with moderate spindle orientation defects (Miller and Rose, 1998). Survival of single deletion mutants largely relies on the function of the spindle orientation checkpoint (SPOC) that retains cells in anaphase until the spindle orientation is corrected (Bardin et al., 2000; Pereira et al., 2000; Caydasi and Pereira, 2012).
Furthermore, MTs in S. cerevisiae are organized exclusively from the spindle pole body (SPB), which is the functional equivalent of animal centrosome. The SPB is a multilayered cylindrical organelle that is embedded in the nuclear envelope (NE) throughout the cell cycle (Byers and Goetsch, 1974; Byers and Goetsch, 1975 )The outer plaque faces the cytoplasm and nucleates cMTs, whereas the inner plaque is inside the nucleus and organizes the nuclear MTs. The central plaque anchors and interconnects the outer and inner plaques (O'Toole et al., 1999; Jaspersen and Winey, 2004). In G1 phase, some fractions of the cMTs are organized from a modified region of the NE associated with one side of the SPB known as the half-bridge (Byers and Goetsch, 1974; Byers and Goetsch, 1975). Spc72, a γ-tubulin complex (γ-TuSC) receptor, is required for nucleating MTs at both the outer plaque and the half-bridge (Chen et al., 1998; Knop and Schiebel, 1998; Wigge et al., 1998; Souès and Adams, 1998). Localisation of Spc72 at the outer plaque is mediated by binding to Nud1, whereas Kar1 serves as a G1 specific binding site of Spc72 at the half-bridge (Pereira et al., 1999; Gruneberg et al., 2000). Spc72 also has a structural role as an integral part of the outer layer and as such localisation of Spc72 to the SPB and the ability to nucleate cMTs persist through the entire cell cycle (Shaw et al., 1997; Pereira et al., 1999; Kosco et al., 2001). Importantly, Spc72, and hence cMTs, is not recruited for the formation of the SPB. New SPB acquires Spc72 and cMTs after the formation of a 1 µm long spindle (Shaw et al., 1997; Segal et al., 2000; Juanes et al., 2013).
In addition to the γ-tubulin complexes, Spc72 exerts a role in recruiting several other proteins to SPBs including Stu2, a microtubule-associated protein (MAP) of the XMAP215/Dis1 family, the SPOC kinase Kin4, as well as polo-like kinase Cdc5 (Chen et al., 1998; Usui et al., 2003; Maekawa et al., 2007; Snead et al., 2007). Cdc5 regulates multiple cellular functions including SPB duplication, progression through G2/M phase, promoting mitotic exit, and cytokinesis (Shirayama et al., 1998; Hu et al., 2001; Song and Lee, 2001; Archambault and Glover, 2009; Elserafy et al., 2014). Cdc5 is also involved in the regulation of spindle orientation in pre-anaphase and migration of the anaphase spindle (Snead et al., 2007; Park et al., 2008). Although Spc72 becomes highly phosphorylated during mitosis in a Cdc5-dependent manner, it is unclear whether this phosphorylation has a regulatory effect on Spc72 and/or cMTs (Maekawa et al., 2007; Snead et al., 2007).
The molecular mechanisms that control spindle orientation in S. cerevisiae have been well established. However, other species that employ the budding mode of cell division may have adopted different strategies. In the pathogenic yeast Candida albicans, the nucleus is located away from the bud neck in pre-anaphase cells (Martin et al., 2004; Finley et al., 2008). C. albicans and probably some of other species in Saccharomycotena may therefore have different mechanisms and regulations in this fundamental biological process.
Ogataea polymorpha (previously Hansenula polymorpha) is extensively used in industrial biotechnology, for the production of various pharmaceuticals in particular for its advantageous characteristics including methylotrophy, nitrate assimilation, availability of strong promoters, and low amount of secreted proteins (Gellissen et al., 2005; Stöckmann et al., 2009 ). Another attractive property of O. polymorpha is its thermotolerant nature (up to approximately 50°C), which may reduce the cost of cooling in, for instance, bioethanol production that requires the treatment of raw materials at high temperature prior to fermentation. However, despite its importance, cell biology research on this organism remains limited. A better understanding of the molecular physiology of O. polymorpha is beneficial towards improving the abilities and characteristics of this yeast for a wide variety of applications.
Here, we describe cMT organization and its regulation during the cell cycle of the methylotrophic yeast O. polymorpha. Unlike S. cerevisiae, the pre-anaphase spindle is not readily positioned and oriented in O. polymorpha owing to the poorly organized cMTs at early cell cycle stages. The bottleneck of cMT nucleation/anchoring at SPBs occurs at the level of Spc72 recruitment to the SPBs, for which the polo-like kinase Cdc5 plays a crucial role. Consistent with the cell cycle dependent activity of cMTs, SPB structure also undergoes cell cycle dependent modification. Thus, our study shed light on the divergent nature of the temporal control of the cMT formation in yeast species.
Results
Nuclear positioning in O. polymorpha differs from that in S. cerevisiae and other budding yeast species
The nucleus is positioned close to the bud neck in large budded pre-anaphase cells of S. cerevisiae (Figure 1A). Similar organization was observed in other budding yeast species including Candida glabrata, Kluyveromyces lactis, Pichia pastoris, and Yarrowia lipolytica (Figure 1—figure supplement 1). Notably in O. polymorpha, however, nuclear position was not biased to the bud neck, although it remained in the mother cell body (Figure 1A, Figure 1—figure supplement 1). The phenotype resembled, but was more exaggerated than, that in C. albicans where the nucleus is located with a distance from the bud neck in pre-anaphase cells (Martin et al., 2004; Finley et al., 2008). A similar phenotype was observed in species closely related to O. polymorpha or C. albicans (Figure 1—figure supplement 1). Close examination of O. polymorpha revealed that the nucleus was located in the cell centre in 76.7% of G2/M cells whereas in the remainder of the cells it was off-centred with no bias towards the bud neck (Figure 1A, Figure 1—figure supplement 2). These results suggest that the nuclear position is not determined before anaphase onset in O. polymorpha. As a consequence, the early stages of anaphase occurred in the mother cell body (Figure 1B). This was further confirmed in cells expressing the α-tubulin gene (Tub1)-GFP and histone H3 gene (Hht1)-mRFP (monomeric red fluorescent protein) to visualize MTs and chromosomal DNA, respectively. Majority of the early anaphase spindles as judged according to their length (<5 µm) as well as a stretched DNA mass were located entirely in the mother cell body (97.4%, Figure 1C, Figure 1—figure supplement 3A). However, 94.7% of the early anaphase spindles were aligned along the bud-mother axis and almost all of the late anaphase spindles with two segregated DNAs were inserted into the bud (Figure 1—figure supplement 3), suggesting immediate and efficient orientation of the spindle during anaphase.
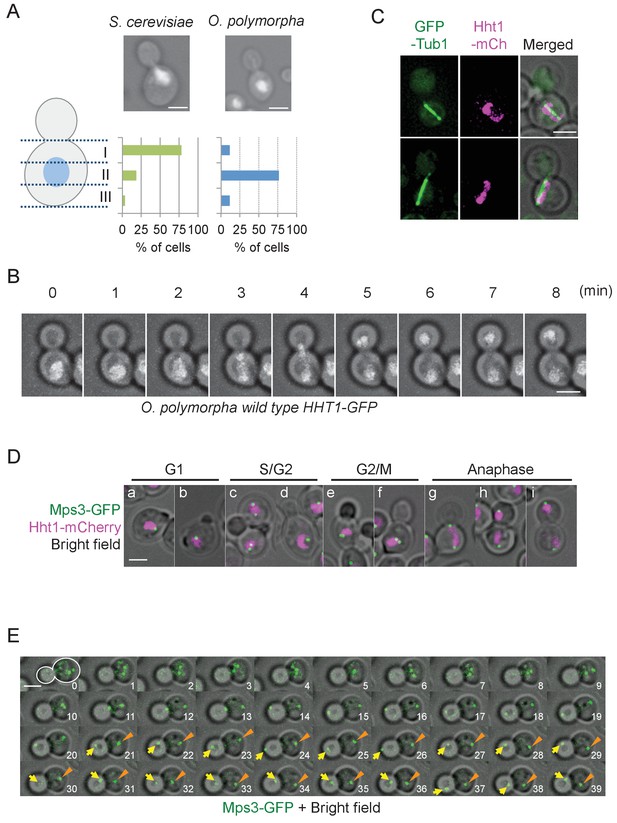
Nuclear positioning in O. polymorpha.
(A) Nucleus is positioned in the cell centre in pre-anaphase cells of O. polymorpha. S. cerevisiae strain YPH499 and O. polymorpha type strain CBS4732 were grown in YPDS at 30°C. DNA was stained with DAPI. The positions of nuclei were as outlined in the cartoon shown on the left side of the subfigure. Scale bar, 2 µm. N = 60 (YPH499), 55 (CBS4732). Result of a similar experiment using HHT1-GFP cells is shown in Figure 1—figure supplement 2. (B) Time-lapse microscopy of histone H3 gene (HHT1)-GFP cells (HPH31). Anaphase onset judged by Hht1-GFP was observed at the 3 min timepoint. Shown are a merged figure of bright field images and deconvolved and projected GFP images. Scale bar, 2 µm. (C) Early anaphase cells (HPH164) with a single DNA mass along the spindle grown in YPDS at 30°C. Microtubules and DNA are visualized by GFP-Tub1 and Hht1-mCherry fluorescence, respectively. Scale bar, 2 µm. (D) Early anaphase cells (HPH1678) grown in YPDS at 30°C. SPB and DNA are visualized by Mps3-GFP and Hht1-mCherry fluorescence, respectively. Scale bar, 2 µm. (E) Time lapse microscopy of cells expressing SPB-GFP maker. MPS3-GFP cells (HPH1681) were grown in SD complete medium at 30°C. Consecutive sections were taken every 60 s. Shown are representative images of cells with an inappropriate angled spindle against the polarity axis. Yellow arrows and orange arrowheads point SPB. Shown are deconvolved and projected GFP images merged with bright field image. Scale bar, 2 µm. Another example is shown in Figure 1—figure supplement 6.
SPB position during the cell cycle was examined to clarify the spindle cycle relative to bud size in cells expressing the SPB marker (Mps3-GFP) (Figure 1D). Two SPB signals appeared in some of small/medium budded cells (Figure 1D, panel c, lower cell), suggesting that SPBs were duplicated at the timing of bud emergence or later, which is similar to that in S. cerevisiae although the precise cell cycle stage should be carefully determined. In the rest of small budded cells, one SPB signal was evident until large budded cells (Figure 1D, panel d). This may be because duplicated SPBs remained in a close proximity and could not be resolved by our fluorescence microscopy. Consistent with this, intensity of Mps3-GFP in medium/large-budded cells with a single SPB was much higher than that in unbudded G1 cells or cells with separated SPBs (Figure 1—figure supplement 4). Moreover, SPB in G1 cells as well as small budded cells was not in the defined position within the mother cell body (Figure 1D). Subsequent time lapse analysis revealed that after spindle assembly, ~1 µm long spindles remained at their central positions and loosely oriented toward the bud neck until shortly before anaphase onset (defined by the rapidly increase of pole-to-pole distance) (Figure 1—figure supplement 5). Anaphase initiated in the mother cell body (Figure 1E, 11 min, Figure 1—figure supplement 6, 3–4 min). These observation defined cells with 2SPBs in a < 2 µm distance as pre-anaphase cells. Spindle alignment was corrected around the time of (or shortly after) spindle elongation, followed by SPB insertion into the bud. After spindle breakdown, the SPB moved vigorously with no relationship to the polarity axis (Figure 1E, Figure 1—figure supplement 6).
O. polymorpha cells contain only fewer cMTs
Lack of nuclear positioning and spindle orientation in pre-anaphase cells may indicate a very low number of cMTs at SPBs during this cell cycle window. To test this notion, we first investigated the MT organization during the cell cycle in GFP-TUB1 HHT1-mCherry cells (Figure 2A and B). Cell cycle stages were judged by the bud size and the number of DNA masses. cMTs were observed in all anaphase cells, whereas less than 50% of G1 and pre-anaphase cells carried cMTs. Furthermore, cMTs that apparently did not associate with the SPB were observed in 13.8% of cells prior to anaphase (Figure 2C). Time lapse analysis revealed that detached cMTs remained in the cytoplasm for only a short period of time before depolymerized (Figure 2—figure supplement 1). This situation is in stark contrast to that in S. cerevisiae where almost all cells exhibit cMTs that are stably associated with the SPB during the cell cycle (Shaw et al., 1997; Kosco et al., 2001).
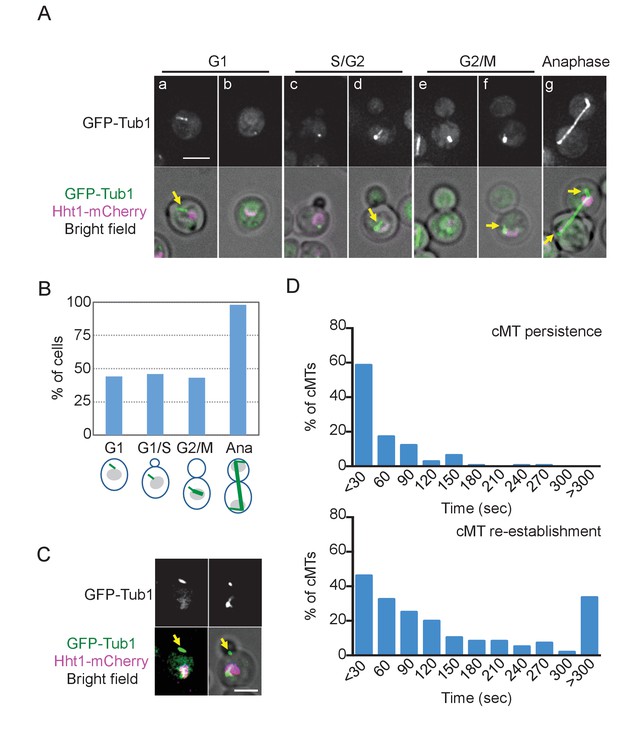
O. polymorpha cells contain fewer cytoplasmic microtubules (cMTs).
(A) Wild-type cells (HPH164) were grown in YPDS medium at 30°C. Microtubules and DNA are visualized by GFP-Tub1 and Hht1-mCherry fluorescence, respectively. cMTs are marked by yellow arrows. a, b: unbudded G1 cell; c, d: preanaphase cell with monopolar nuclear MTs; e, f: preanaphase cell with bipolar spindle; g: anaphase cell. Scale bar, 2 µm. (B) Quantification of (A). G1, S/G2, G2/M, and anaphase represent unbudded cells, small budded cells with single unduplicated SPB, medium budded cells with 2 SPB, and large budded cells with an SPB in both the mother and the bud, and large budded cells with elongated spindle, respectively. n > 50 cells for each category. Ana, anaphase. (C) Cells containing the cMT detached from the SPB in (A). Scale bar, 2 µm. (D) Analysis of the duration of cMT persistence and cMT re-establishment time at SPBs in preanaphase cells with a bipolar spindle by time-lapse microscopy. GFP-TUB1 cells (HPH194) were grown in SD complete medium at 30°C. Consecutive sections were taken every 30 s. Duration of continuous cMT presence was scored as cMT persistance, while the time between the loss of cMT and the acquisition of a new cMT was scored as cMT re-establishment. Total recording time was 38460 s.
The small number of observed cMTs might have arisen because of reduced cMT nucleation. Another and not mutually exclusive possibility considers that cMTs might not be stably anchored to the SPB and thus might not persist over long periods. To test these possibilities, we performed time lapse experiments with cells expressing GFP-TUB1, in which cMTs were observed during 20.6% of the recorded time points although the majority (>80%) did not persist longer than 30 s (Figure 2D, cMT persistence, Figure 2—figure supplement 2). These results suggest that cMTs are short-lived during early stages of the cell cycle. Once cMTs were lost, a relatively long time was required until new MTs appeared at the SPB (Figure 2D, cMT re-establishment; median value 90.0 s, average 173.8 ± 192.4 s). Thus, cMTs are less frequently nucleated and unstable at early stages of the cell cycle. Acquired cMTs efficiently corrected the spindle orientation in pre-anaphase cells, suggesting that the spindle orientation is regulated largely at the level of cMT acquisition (Figure 2—figure supplement 3).
Organization of the SPB structure on the cytoplasmic side is cell cycle dependent
Next, we examined the SPB structure in G1 and anaphase by electron microscopy. An electron dense SPB-like structure was evident in all cells examined, while a half-bridge-like structure—which plays an important role in cMT organization in G1 of S. cerevisiae—was not clearly observed (Figure 3). Anaphase SPBs had an additional thin layer in the cytoplasm that resembled the outer plaque of S. cerevisiae SPB (Figure 3A and B) (Byers and Goetsch, 1974; Byers and Goetsch, 1975). In contrast, there was no detectable outer plaque in G1 SPBs (Figure 3C and D). These results suggested that O. polymorpha SPBs undergo structural cycling in every cell cycle.
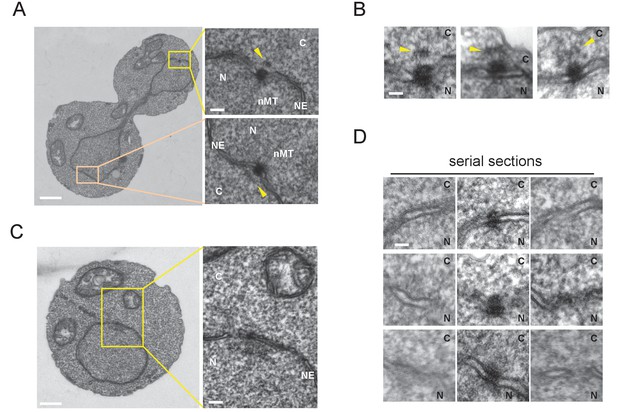
Cytoplasmic structure of SPB is regulated during the cell cycle.
Electron microscopy (EM) of thin serial sections of cells in G1 and anaphase. Wild-type cells (BY4329) were grown to log phase at 30°C in YPDS and then prepared for EM. Indicated are the cytoplasm (C), nucleus (N), nuclear envelope (NE), and nuclear microtubules (nMT). (A, B) Representative SPBs in anaphase cells (n = 10). The SPB in the mother does not appear in the section shown as the main image. The pale orange rectangle in the mother merely indicates the position of the lower inset which is the image of the section containing the SPB in the mother. (C, D) representative SPBs in unbudded G1 cells (n = 10). Consecutive three sections are shown in (D). Scale bars of the main images in (A) and (C) represent 1 µm. Scale bars of the insets in (A) and (C), i.e., (B) and (D), represent 100 nm.
Our attempt to arrest cells in late G1 by introducing cdc28-as allele, which arrest S. cerevisiae cells in late G1 with a single SPB and satellite, was failed probably because of insufficient inhibition of the kinase (Figure 3—figure supplement 1). However, inhibitor addition delayed cell cycle progression leading to the accumulation of cells with unseparated SPBs. This allowed us to examine the structure of side-by-side SPBs by EM (Figure 3—figure supplement 2). All six side-by-side SPBs had outer plaques which were similar to that in nocodazole arrested cells, albeit some of those were somewhat fuzzy. The result suggested that the SPB structure on cytoplasmic side is reconstructed before spindle formation. An additional electron dense cloud was observed on the cytoplasmic side of nuclear envelope between two SPB bodies, which resembled the half-bridge/bridge structure of S. cerevisiae SPB (Figure 3—figure supplement 2A, B and E; orange arrowheads). It was not clear whether this structure was present at other cell cycle stages. A better synchronization method is required to determine the fine structure and the precise timing of emergence/disappearance of the outer plaque during the cell cycle.
Spc72 associates with SPB in a cell cycle-dependent manner
Lack of the outer plaque in G1 prompted us to search for SPB components whose association with SPBs was cell cycle dependent. In S. cerevisiae, core SPB components are found at SPBs throughout the mitotic cell cycle including central plaque components (Spc42, Spc29), outer plaque components (Cnm67, Nud1), half-bridge components (Sfi1, Cdc31, Mps3, and Kar1), and membrane anchors (Ndc1, Nbp1, Mps2, and Bbp1) (Winey et al., 1991; Spang et al., 1995; Bullitt et al., 1997; Brachat et al., 1998; Wigge et al., 1998; Chial et al., 1998; Adams and Kilmartin, 1999; Elliott et al., 1999; Schramm et al., 2000; Kilmartin, 2003; Jaspersen et al., 2002; Araki et al., 2006). The γ-TuSC recruiting factors Spc110 and Spc72 also represent core components of SPB in the inner and outer plaques, respectively (Knop and Schiebel, 1997; Knop and Schiebel, 1998). BLAST and HMMER searches have identified putative orthologues of genes for Mps3, Sfi1, Spc72, Spc110, and Nud1 as well as γ-TuSC components, Tub4, Spc97, and Spc98 in the O. polymorpha genome (Altschul et al., 1990; Sobel and Snyder, 1995; Geissler et al., 1996; Knop and Schiebel, 1997; Eddy, 1998; Maekawa and Kaneko, 2014; Riley et al., 2016). SPB-like localisation was verified by expressing GFP or mRFP-fused version of these proteins. GFP or mRFP dot-like signals of Tub4, Spc98, Mps3, Sfi1, and Nud1 were observed in most of the cells, suggesting that they represent constitutive components of the SPB throughout the cell cycle (Figure 4A, Figure 4—figure supplement 1). In contrast, the Spc72 signal was either weak or absent in cells at early cell cycle stages, whereas all anaphase cells carried two strong SPB signals (Figure 4A, Figure 4—figure supplement 1). Deletion of SPC72 in S. cerevisiae results in severe growth defects or lethality depending on the strain background. To evaluate the effect of SPC72 deletion in O. polymorpha, SPC72/spc72Δ::natNT2 heterozygous diploid cells were subjected to tetrad dissection analysis. Notably, 21 out of 29 tetrads yielded one or two viable colonies, all of which were sensitive to nourseothricin (Figure 4—figure supplement 2). Microscopic inspection revealed that 91.7% of spores with spc72Δ::natNT2 genotype derived from tetrads that gave two viable nourseothricin-sensitive colonies were germinated. These results suggested that SPC72 is essential for growth in O. polymorpha.
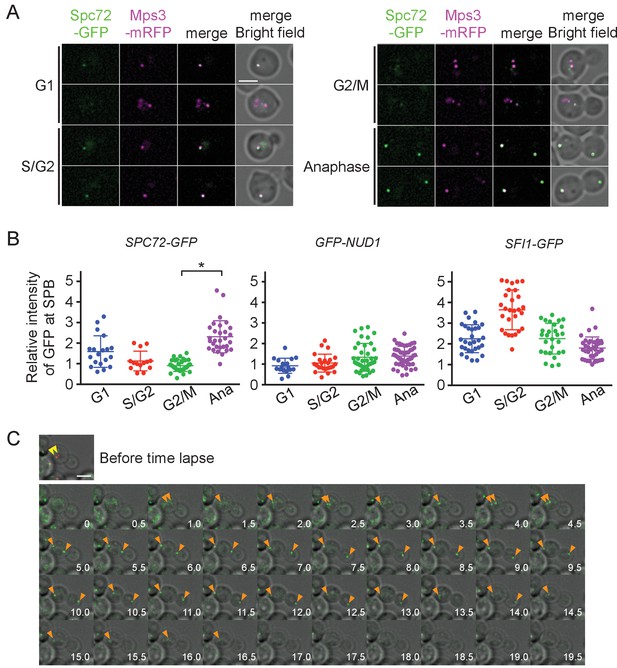
Accumulation of Spc72 at SPBs is cell cycle dependent.
(A) Cell cycle dependent localization of Spc72-GFP. SPC72-GFP MPS3-mRFP cells (HPH1394) were grown in SD complete medium at 30°C. Cell cycle stages are as shown in Figure 2B. Mps3-mRFP is a marker for SPB. Scale bar, 2 µm. (B) Quantification of the Spc72/Nud1/Sfi1 SPB signal of cells at different cell cycle stages. Yeast strains HPH972, HPH1396, and HPH1400 were used. Signal intensities were background-subtracted. Statistical significance of the difference between 2 SPBs and anaphase was determined by the student t-test and is indicated by an asterisk. Error bars indicate SD. n = 95, 127, and 124 for Spc72, Nud1, and Sfi1, respectively. Note that intensity of some of Sc72-GFP signals in G1 was high, because it decreased only gradually at SPB during the end of mitosis and the following G1 as shown in (C). (C) Time-lapse microscopy of SPC72-GFP MPS3-mRFP cells (HPH1394). Images were taken every 30 s. RFP signal was captured only before staring the time-lapse series. Anaphase onset judged by sudden spindle elongation was observed at the 5 min timepoint. Spindle orientation was corrected between 4.5 min and 5 min timepoints. Yellow arrowheads indicate the position of Mps3-mRFP before the image capture. Orange arrowheads indicate Spc72-GFP signals at SPBs. Shown are deconvolved and projected images. Scale bar, 2 µm. Another example is presented in Figure 4—figure supplement 1.
To more precisely evaluate the amount of Spc72 at the SPB, images were obtained in logarithmically growing wild-type cells carrying SPC72-GFP MPS3-mRFP and the GFP intensity at the SPB was quantified (Figure 4B). The GFP signal was 2.5 times weaker in cells with a short spindle than in anaphase cells (p<0.001). In contrast, Nud1, which comprises the putative binding site of Spc72 on the outer plaque as suggested by the direct interaction between orthologues of these proteins in S. cerevisiae, did not show this trend, nor did Sfi1, a half-bridge component (Figure 4B) (Gruneberg et al., 2000; Kilmartin, 2003). High intensity of Sfi1-GFP signal in S/G2 cells most likely arose from SPBs that were duplicated but not yet separated. These results suggest that Spc72 is cell cycle regulated and the incorporation of Spc72 into SPBs may be the key step to stabilize cMTs. To further confirm this notion, time-lapse microscopy was carried out to determine the timing of Spc72 association with SPBs (Figure 4C, Figure 4—figure supplement 3). In all cells that progressed into anaphase, an Spc72-GFP signal became detectable <4 min prior to the initiation of anaphase (average 3.68 ± 1.74 min, n = 14) (Figure 4C, orange arrowheads). Within 5 min after appearance of the Spc72-GFP signal, spindle orientation was corrected when it had not done already (Figure 4—figure supplement 3, average 3.50 ± 1.61 min, n = 12); therefore, one half part of an anaphase nucleus was successfully inserted into the bud. Thus, Spc72 accumulates at SPB in early mitosis, most likely in metaphase, and remains high during anaphase. As cells exit from mitosis and entre the next cell cycle, Spc72-GFP signal was gradually decreased at SPBs with the timing that varied from cell to cell. This difference of timing may explain the relatively high and variable intensity of Spc72-GFP at SPB in G1 cells (Figure 4B). However, in all cases, Spc72-GFP levels reached a minimum well before short spindle was formed (Figure 4, Figure 4—figure supplement 4).
If low abundance of Spc72 at the SPB is the reason underlying cMT instability, higher expression of Spc72 might increase the level of Spc72 at the SPB and consequently raise the number of cMTs, and thereby promote positioning the nucleus close to the bud neck at early stages of the cell cycle. We expressed the SPC72-GFP gene from a strong constitutive TEF1 promoter in cells whose endogenous SPC72 was also fused to GFP and examined the position of the SPB relative to the bud neck in pre-anaphase cells as a readout of cMT function (Kiel et al., 2007). Overexpressed Spc72-GFP was efficiently targeted to SPB because a strong Spc72-GFP signal was observed in cells carrying the PTEF1-SPC72-GFP gene but not in wild-type cells during G1 and G2/M phases (Figure 5A, Figure 5—figure supplement 1). In cells overexpressing SPC72-GFP, SPB was positioned close to the bud neck, which is reminiscent of the SPB position in S. cerevisiae (Figures 1A and 5B), and cMTs were more often observed (Figure 5C and D). Together, these results strongly supported our hypothesis that cMT organization is regulated at the level of repeated Spc72 recruitment to the SPB in every cell cycle.
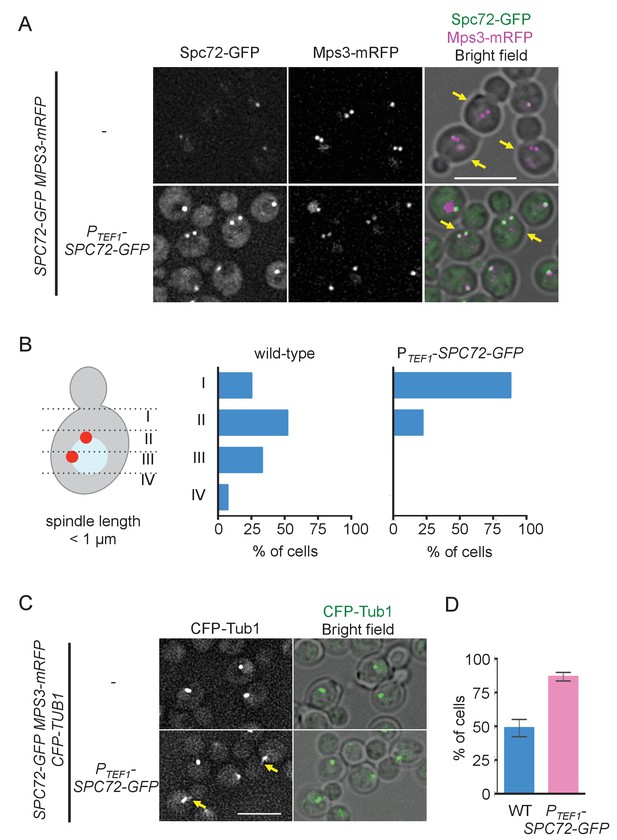
Overexpression of Spc72 converts the O. polymorpha type of nuclear position to the S. cerevisiae type.
(A) Overexpressed Spc72-GFP were recruited to SPB at all stages of the cell cycle. SPC72-GFP was expressed from a strong promoter of the TEF1 gene (HPH1393). Enrichment of Spc72 to SPB was evident in G1 cells and cells with short spindles (SPB distance <1 µm, yellow arrows) compared with images of wild-type cells (HPH1394). Mps3-mRFP marks SPB. Scale bar, 5 µm. (B) SPB is positioned close to the bud neck in G2/M cells carrying PTEF1-SPC72-GFP. The position of the SPB closer to the bud was as outlined in the cartoon shown on the left side of the subfigure. Strains used were HPH1393 (n = 110) and HPH1394 (n = 117). (C) Overexpression of Spc72 stimulated cMT acquisition. MTs were visualized with CFP-TUB1 in wild type and cells overexpressing SPC72-GFP (HPH1653 and HPH1652, respectively) were grown in YPDS medium at 30°C. Images were captured only for CFP and brightfield. Scale bar, 5 µm. (D) Quantification of (C). Presence/absence of cMTs was scored in cells with short spindle. Shown is the average of three independent experiments. Error bars indicate SD. n > 100. Average of three independent experiments.
cMT play important roles in yeast mating and karyogamy, which are initiated in G1. Because mating is triggered by nutrient starvation in O. polymorpha, we examined cMTs and Spc72 in nutrient starved cells. Interestingly, while Spc72 was accumulated at SPBs, cMTs were not observed (Figure 5—figure supplement 2). Thus, specific mechanism may regulate Spc72 and cMT organization under such conditions.
SPB association of Spc72 is dependent on the polo-like kinase Cdc5
Spc72 might be regulated at the level of protein expression. To synchronize cells for monitoring changes in protein levels during the cell cycle, we transferred the recently developed auxin-inducible degradation (AID)-degron system to O. polymorpha (Nishimura and Kanemaki, 2014). CDC5 encodes the only polo-like kinase in yeast. It is thus essential for growth in S. cerevisiae, and its inactivation causes cell cycle arrest in late anaphase (Kitada et al., 1993). Similarly, a single CDC5 orthologue was identified in the O. polymorpha genome (OpCDC5). Logarithmically growing cells carrying a 3mAID-tagged version of CDC5 were arrested as large budded cells by incubation in the presence of auxin and then released into fresh medium without auxin to resume the cell cycle (Figure 6A). Spc72 protein abundance did not fluctuate as cells entered into anaphase and proceeded into the following cell cycle (Figure 6A, Figure 6—figure supplement 1A). Furthermore, the Spc72 band migrated slower in nocodazole-arrested cells than that in asynchronous cells (Figure 6—figure supplement 1B). These results suggested that either post-translational modification of Spc72 or regulation of Spc72 binding proteins might be utilized to achieve cell cycle dependency of SPB localisation.
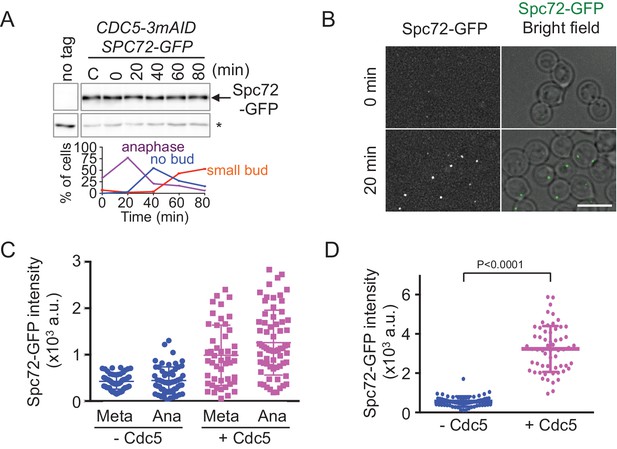
The recruitment of Spc72 to SPBs is dependent on the polo-like Cdc5 kinase.
(A) Abundance of Spc72 does not fluctuate during the cell cycle. Logarithmically growing SPC72-GFP CDC5-3mAID PCDC28-OsTIR cells (HPH1380) were synchronized with IAA followed by release. Samples were analysed by immunoblotting for Spc72-GFP. Comparable intensities of unspecific signal in immunoblotting (asterisk) indicate equal loading of samples. Budding index and mitotic index (DAPI) were determined over time. n > 100 cells per time point. (B) SPC72-GFP CDC5-3mAID PCDC28-OsTIR cells (HPH1380) were synchronized and released as in (A). Images were captured without fixation. Shown are deconvolved and projected images. Time after release is indicated on the left. Scale bar, 5 µm. (C) Quantification of Spc72-GFP intensity at SPBs in (B). Signal intensities were background-subtracted. Statistical significance of the difference between 2 SPBs and anaphase was determined by the student t-test. – Cdc5: before the release;+Cdc5: 20 min after the release. Error bars indicate SD. n > 50 cells per time point. (D) Accumulation of Spc72 at SPBs in metaphase depends on Cdc5 function. SPC72-GFP CDC5-3mAID PCDC28-OsTIR cells (HPH1380) were arrested with nocodazole in the presence (− Cdc5) or absence (+Cdc5) of IAA and Spc72-GFP signal at SPBs was quantified. Signal intensities were background-subtracted. Error bars indicate SD. n > 50 cells.
Furthermore, we noticed that the GFP intensity of Spc72-GFP at the SPB was significantly lower in cells arrested by Cdc5-depletion than that in cells after re-accumulation of Cdc5 in both metaphase (SPB distance <2 µm, p<0.0001) and anaphase (SPB distance >4 µm, p<0.0001) (Figure 6B and C). Strong dependency of SPB binding of Spc72 on Cdc5 was further confirmed in metaphase-arrested cells by nocodazole (Figure 6D, Figure 6—figure supplement 2). These results suggested that the stable association of Spc72 requires Cdc5 kinase, the activity of which is likely cell cycle-regulated.
Spc72 is phosphorylated by Cdc5 kinase in S. cerevisiae (Maekawa et al., 2007; Snead et al., 2007), which prompted us to investigate whether Spc72 is subjected to a Cdc5-dependent phosphorylation in O. polymorpha. Our gel electrophoresis analyses of nocodazole-arrested cells suggested that Spc72 of O. polymorpha is subjected to post-translational modifications in a Cdc5-dependent manner because the Spc72 band was more smeared and migrated slightly slower in wild type and CDC5-overexpressing cells as compared to that in Cdc5-depleted cells (Figure 6—figure supplement 3A). Although this difference was largely lost during preparation of cell extract, the λ-phosphatase treatment revealed in vivo phosphorylation of Spc72 that was independent of Cdc5 (Figure 6—figure supplement 3B), suggesting that Cdc5 contributes to a subset of phosphorylations of Spc72. Thus, Spc72 is phosphorylated at multiple sites, only some of which depend on Cdc5.
Cdc5 localises to NE and SPB during mitosis
We next examined the localisation of Cdc5 during the cell cycle. Distinctive localization of Cdc5-GFP became apparent after S phase and was lost prior to or during the following G1 phase (Figure 7A and B, Figure 7—figure supplement 1). Nuclear and NE localisation appeared at early stages of the cell cycle and persisted until the end of mitosis. In addition, a fraction of the GFP signals appeared to overlap with those of SPBs during mitosis. Neither NE nor SPB-associated GFP signals were detected in unbudded or small budded cells. Notably, SPB signal may arise from the SPB itself, NE surrounding the SPB, or kinetochores that cluster close to the SPB during interphase and anaphase in yeast (Jin et al., 1998). Therefore, in order to clarify on which side of SPB the Cdc5-GFP signal resided, we employed structured illumination microscopy (SIM). Localisation was investigated in metaphase-arrested cells with nocodazole where Cdc5-GFP signal was observed at SPB as well as in nucleus (Figure 7—figure supplements 1B and 2), and Spc72, and Spc110 were used as references for the cytoplasmic and nuclear side of SPB, respectively. Spc72-GFP and Spc110-tdTomato signals were clearly distinguished in 58% of the cells, which verified that our method could discriminate signals in the cytoplasmic and the nuclear side of SPB in >50% of cells (Figure 7C). The resolution of both signals probably depends on the orientation of the SPB (top versus side view). Only the SPB side view will resolve Spc72-GFP and Spc110-tdTomato signals at SPBs. Cdc5-GFP overlapped with Spc110-tdTomato in 53% of cells, which is similar to the degree of co-localisation observed between Spc72 and Spc110. In contrast, the Cdc5-GFP signal overlapped with Spc72-tdTomato in 87% of cells (Figure 7D). These results suggest that Cdc5-GFP locates at the position on SPBs closer to Spc72 than to Spc110, indicating that Cdc5-GFP signal arises from the cytoplasmic side of SPB. Thus, Cdc5 likely becomes first localised to the nucleus and the NE in G2, and then in mitosis to the cytoplasmic side of SPBs. The timing of Cdc5 binding to SPBs coincides well with the recruitment of Spc72 to SPBs.
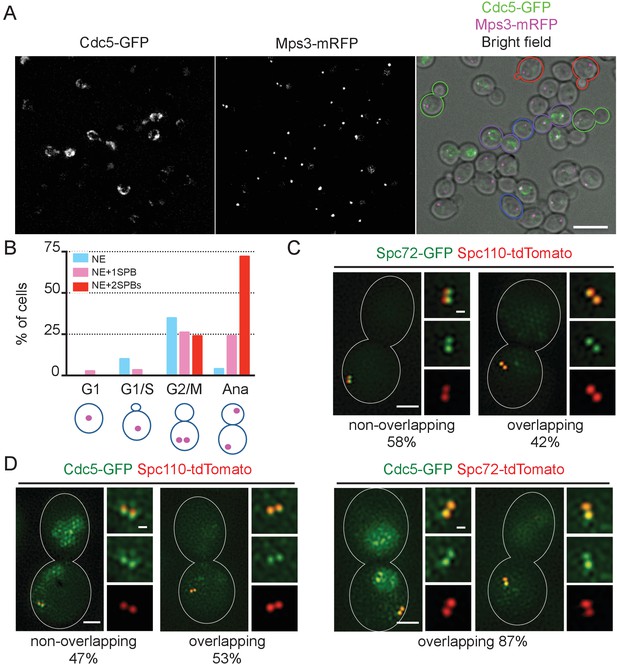
Localization of polo-like Cdc5 kinase in the nucleus, the nuclear envelope, and at the SPB is cell cycle dependent.
(A) CDC5-GFP MPS3-mRFP cells (HPH1562) were grown in YPDS at 30°C. Blue, red, green, and purple cell contours mark G1, S/G2, G2/M, and anaphase cells, respectively. Shown is a projected image after deconvolution. Scale bar, 5 µm. (B) Quantification of Cdc5-GFP localization in (A). The position of SPB (magenta) was as outlined in the cartoon shown at the bottom of the subfigure. n > 30 cells for each cell cycle stage. Ana, anaphase. (C) SIM images of Spc72-GFP and Spc110-tdTomato in nocodazole-arrested cells. SPC72-GFP SPC110-tdTomato cells (HPH1581) grown in SC medium at 30°C were arrested in metaphase with nocodazole. Scale bars, 1 µm and 0.2 µm in the large and small images, respectively. n = 78. (D) SIM images of Cdc5-GFP together with either Spc110-tdTomato or Spc72-tdTomato in nocodazole-arrested cells. CDC5-GFP SPC110-tdTomato and CDC5-GFP SPC72-tdTomato cells grown in SC medium at 30°C were arrested in metaphase with nocodazole. Scale bars, 1 µm and 0.2 µm in the large and small images, respectively. Strains used were HPH1583 (n = 75) and HPH1575 (n = 55). Diffused nuclear signal of Cdc5-GFP were also observed in all cells (Figure 7—figure supplement 2).
CDC5 overexpression accelerates the Spc72 recruitment to SPB
To further confirm the significance of Cdc5 kinase in the recruitment of Spc72 to SPBs, we constitutively expressed the CDC5 gene at high level. While Cdc5 expression showed no effect on the protein level of Spc72-GFP (Figure 8—figure supplement 1), Spc72-GFP intensity at the SPB was higher in metaphase-arrested cells following ectopic expression of CDC5 from the TEF1 promoter than in wild-type cells (p<0.0001) (Figure 8A and B). In the similar analysis performed in asynchronously growing cells, accumulation of Spc72-GFP at SPB was significantly higher at all stages of the cell cycle in cells overexpressing CDC5 than in wild type cells (p<0.01 for G1 cells; p<0.0001 for S/G2, G2/M, and anaphase cells), with the strongest effect observed in G2/M phase (Figure 8C and D), and cMTs were more prevalent (Figure 8E and F). As a consequence, the SPB positioned closer to the bud neck (Figure 8G) and the spindle was at an angle within 30° with respect to the mother-bud axis in 83% of pre-anaphase cells overexpressing CDC5 compared with 48% in wild type cells (Figure 8—figure supplement 2). These results suggest that the stable SPB association of Spc72 is restricted to the time period where Cdc5 kinase activity is sufficiently high.
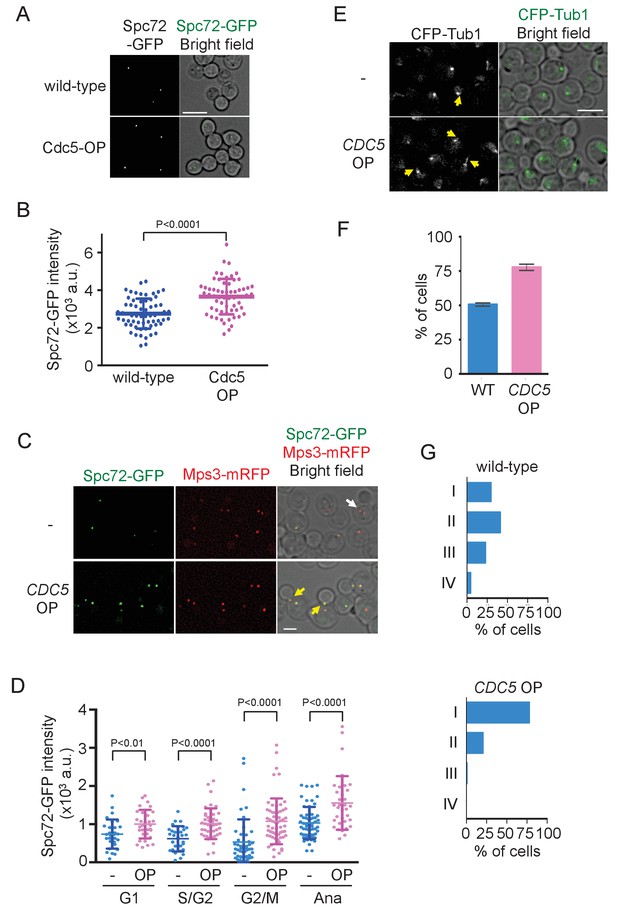
Overexpression of Cdc5 kinase promotes early association of Spc72 to SPBs.
(A) Overexpression of CDC5 enhances SPB binding of Spc72. SPC72-GFP MPS3-mRFP (HPH1394) and SPC72-GFP MPS3-mRFP PTEF1-CDC5∆53 cells (HPH1542) were arrested in metaphase with nocodazole. Deconvolved and projected images are shown. Scale bar, 5 µm. (B) Spc72-GFP signal at SPBs in (A) was quantified. Signal intensities were background-subtracted. Error bars indicate SD. n > 50 cells. (C) Spc72-GFP was recruited to SPB at earlier stages of the cell cycle when CDC5 gene is overexpressed. SPC72-GFP MPS3-mRFP (HPH1394, -) and SPC72-GFP MPS3-mRFP PTEF1-CDC5∆53 cells (HPH1542, CDC5 OP) were grown in YPDS medium. Enrichment of Spc72 to SPB and alignment of the spindle along the mother-bud axis were evident in preanaphase cells (yellow arrows) overexpressing CDC5, compared with wild-type cells (white arrow). Shown are deconvolved and projected images. Mps3-mRFP marks SPB. Scale bar, 2 µm. (D) Quantification of Spc72-GFP intensity at SPBs in (C). –: wild-type; OP: CDC5 overexpression. Signal intensities were background-subtracted. Statistical significances of the difference between wild-type (-) and CDC5 overexpressing cells (CDC5 OP) were determined by the student t-test. Error bars indicate SD. (E) Overexpression of Spc72 stimulated cMT acquisition. MTs were visualized with CFP-TUB1 in wild type and CDC5 overexpressing cells (HPH1680 and HPH1673, respectively) were grown in YPDS medium at 30°C. Images were captured only for CFP and brightfield. Scale bar, 5 µm. (F) Quantification of (E). Presence/absence of cMTs was scored in cells with short spindle. Shown is the average of three independent experiments. Error bars indicate SD. n > 100. (G) SPB is positioned close to the bud neck in G2/M cells overexpressing CDC5. The position of the SPB closer to the bud was as outlined in the cartoon shown in Figure 5B. Strains used were HPH1394 (n = 125) and HPH1542 (n = 101).
Discussion
The mode of cell division by budding represents a type of asymmetric cell division. The mechanism to achieve a high-fidelity of chromosome segregation in such a situation has been a focus of interest and has been investigated intensively in S. cerevisiae. These process was recently studied in the other ascomycetous yeast C. albicans as well as in the yeasts Cryptococcus neoformans and Ustilago maydis in another phylum of fungi, Basidiomycota (McCully and Robinow, 1972a; McCully and Robinow, 1972b; Kopecká et al., 2001; Steinberg et al., 2001; Woyke et al., 2002; Straube et al., 2003; Martin et al., 2004; Finley et al., 2008). Although the movement of the nucleus during the cell cycle differs between ascomycetous and basidiomycetous yeasts, it is commonly positioned close to the bud neck in both phyla prior to chromosome segregation. We report here that the ascomycetous yeast O. polymorpha does not follow the same strategy. In O. polymorpha, the nucleus generally locates centrally within the mother cell body and the spindle is not aligned properly along mother-bud axis until anaphase onset. Consequently, spindle elongation in early anaphase occurs entirely in the mother often with an inappropriate angle against the polarity axis. Despite this potential complication, one nucleus penetrates successfully into the bud during anaphase, which may largely rely on an immediate correction of the orientation of the spindle and on SPOC activity. Those SPB movements are in contrast to S. cerevisiae in which spindle is aligned during metaphase and therefore SPB translocation into the bud coincides with spindle elongation. Currently molecular mechanism(s) that regulate spindle orientation is unknown. However, although the timing of spindle orientation relative to cell cycle progression appears to be different from that of other yeasts, two redundant molecular mechanisms of spindle orientation, one requiring dynein and the other Kar9, may be conserved in O. polymorpha, because putative orthologs of KAR9 and dynein were identified in O. polymorpha genome sequences (Li et al., 1993; Miller and Rose, 1998; Maekawa and Kaneko, 2014; Nordberg et al., 2014).
In S. cerevisiae, Spc72 is stably incorporated into SPBs once it is recruited and organise cMTs throughout the cell cycle. In O. polymorpha, the strong SPB association of OpSpc72 in anaphase becomes weakened as cells enter into the following G1 phase, whereas they re-accumulate later in the cell cycle. The timing of OpSpc72 recruitment to SPBs during early mitosis appears to primarily dictate the organization of cMTs and hence nuclear position (Figure 9). As the polo-like kinase Cdc5 protein plays an important role in this regulation, a question arises regarding the substrates of Cdc5 kinase in this process (Archambault and Glover, 2009). Because ScSpc72 binds to and is phosphorylated by Cdc5 in S. cerevisiae, OpSpc72 represents an obvious candidate (Maekawa et al., 2007; Snead et al., 2007). Our electrophoresis analyses indeed detected Cdc5-dependent phosphorylation of Spc72. However, OpCdc5 failed to phosphorylate recombinant Spc72 in vitro. Further analyses are required to verify whether Cdc5 directly phosphorylates Spc72 and the potential effects of such modifications on the regulation of cMT. Additionally, it may also be important to identify other kinase(s) that are responsible for the Cdc5-independent phosphorylation of Spc72 observed in our analyses. Cdc5 may have another important substrates other than Spc72. Notably, a polo box binding site present in ScSpc72 is missing in OpSpc72. Cdc5 might therefore phosphorylate other SPB proteins such as Nud1, which has one site matching the consensus sequences of polo box binding site (S-Sp/Tp-P), and thereby indirectly influence the affinity of Spc72 towards the SPB.
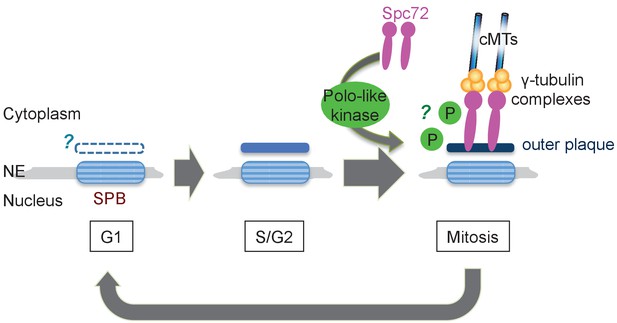
Model of the SPB cycle in O. polymorpha.
See Discussion for details.
It is unclear what brought highly expressed Spc72-GFP to SPBs at early cell cycle stages when Cdc5 activity was low. Weak Spc72-GFP signal in the absence of Cdc5 may be because of low affinity or unstable association to SPBs. Increased Spc72 protein levels may simply result in the increased number of Spc72 protein at SPBs at any given time. Alternatively, overexpression may overcome a negative regulation that normally maintains the low level of Spc72 at SPBs during early stages of the cell cycle (Figure 4). Such Cdc5-independent regulation is consistent with the observation that Spc72 was gradually lost from SPBs at early stages of the cell cycle. Examining properties of Spc72 protein when overexpressed such as post-translational modifications, protein stability, and molecular dynamics at SPBs would clarify this point.
In S. cerevisiae, SPB duplication initiates in late G1 phase by forming a satellite at the distal end of an extended half-bridge of the pre-existing ‘old’ SPB, which is then inserted into the nuclear envelope. In contrast to the old SPB which maintains cMTs from the previous mitosis, this ‘new’ SPB acquires the MT nucleation activity in the inner plaque prior to spindle assembly while it is still connected to the old SPB (side-by-side SPBs). On the other hand, a recent report suggested that the acquisition of ScSpc72 to the outer plaque, and hence of cMTs, occurs only after SPB separation and spindle assembly (Juanes et al., 2013). In O. polymorpha, the cMTs acquisition is also regulated at the level of Spc72 recruitment to SPBs. OpSpc72 dissociate from SPBs at the end of mitosis and recruited to both old and new SPBs shortly before anaphase of the following cell cycle. This suggests that the cell cycle regulation of Spc72 recruitment may be applied to both old and new SPBs in O. polymorpha. Even though the Spc72 recruitment is cell cycle regulated in both species, its timing seems to be different: while it occurs in early G2 phase in S. cerevisiae, it does in metaphase in O. polymorpha. Moreover, their regulatory mechanisms are likely different because their acquisition is Cdc5-dependent in O. polymorpha, but not in S. cerevisiae (Juanes et al., 2013). Inhibition of Cdc5 kinase in S. cerevisiae causes the misaligned spindle phenotype, which indicates a role of Cdc5 in cMT functions (Snead et al., 2007). However, it is unlikely that Cdc5 acts at the level of Spc72 recruitment since both SPBs of the misaligned spindle caused by Cdc5 inhibition carried cMTs (Snead et al., 2007). Alternatively, it is possible that the Cdc5-dependent regulation of ScSpc72 function on cMT nucleation/anchoring has been overlooked. Growth suppression of cnm67∆ cells by CDC5 overexpression might point towards this possibility (Park et al., 2004). It might also be because of the redundancy with the SPB-anchored Stu2 function (Usui et al., 2003). The Stu2 binding domain of Spc72 (aa176–230 in ScSpc72, Figure 4—figure supplement 5) is conserved in ascomycetous yeasts including O. polymorpha but it differs somewhat in OpSpc72. In particular, the detached cMTs observed in approximately 14% of O. polymorpha pre-anaphase cells are reminiscent of the spc72∆Stu2 phenotype in S. cerevisiae (Usui et al., 2003). Thus, the potential binding of Stu2 to Spc72 in O. polymorpha should be investigated.
In O. polymorpha, structure of the cytoplasmic side of SPB, along with cMT nucleation competence, is modified as the cell cycle progresses (Figure 9). Notably, our BLAST search failed to identify orthologs of several SPB core components identified in S. cerevisiae in genomes of outside of Saccharomycetaceae: the central plaque components Spc42 and Spc29, the membrane anchors Nbp1 and Bbp1, and the half-bridge component Kar1. The failure of identification may be due to the highly divergent nature of amino acid sequences of these coiled-coil proteins, or alternatively the function of these SPB components is not required in O. polymorpha. The absence of these proteins could be one of the reasons underlying the poor appearance of the central plaque and the half-bridge in our electron microscopy anlayses. Our observation that the outer plaque in the cytoplasm was evident in anaphase SPBs but not in G1 SPBs may suggest the outer plaque from the previous mitosis is removed in the following G1 phase. However, given that Nud1, a putative outer plaque component, is present at the SPB in G1, it is more likely that the outer plaque is only partially disassembled. Furthermore, the appearance of the outer plaque in EM analysis proceeded that of Spc72 in fluorescent microscopy (Figure 9). Electron microscopy analysis using another fixation method that better preserves the SPB structure might clarify this point. Equally important is the analysis of the half-bridge which organizes cMTs in G1 in S. cerevisiae. If the half-bridge plays the same role in O. polymorpha SPB as that in S. cerevisiae, the loss of Spc72 from outer plaque alone should not cause the loss of cMTs. In addition, in S. cerevisiae, SPBs are segregated in a defined mode where the old SPB normally migrates into the daughter cell while the new SPB remains in the mother cell (Pereira et al., 2001). The SPB history in the outer plaque was proposed to primarily determine the destination of SPBs and to bias spindle asymmetry via Nud1 (Hotz et al., 2012; Juanes et al., 2013). Whether the mode of SPB inheritance is conserved in O. polymorpha and other yeast species is also worthy of further study.
It is unlikely that the mode of nuclear positioning in O. polymorpha is ancestral, given that Y. lipolytica and P. pastoris, who are diverted from the common ancestor earlier than O. polymorph, share the same nuclear organization with S. cerevisiae. However, several yeast species relatively close to O. polymorpha or C. albicans exhibited a similar nuclear position in pre-anaphase cells (Figure 1—figure supplement 1). Thus, the Spc72 recruitment mechanism described in O. polymorpha may be widely utilized in several Clades in Saccharomycotina, including Ogataea, Ambrosiozyma, and Nakazawaea.
Materials and methods
Yeast strains and plasmids
Request a detailed protocolYeast strains and plasmids used in this study are listed in Table 1. Unless otherwise indicated, all O. polymorpha strains were derived from NCYC 495 and were generated by PCR-based methods (Lu et al., 2000; Janke et al., 2004; Saraya et al., 2012). GFP, mRFP, and 5flag tagged alleles were generated in ku80Δ or ku70Δ cells and then crossed with auxotrophic wild-type strains to obtain KU80+ or KU70+ cells carrying the tagged allele (Maekawa and Kaneko, 2014). O. polymorpha cells were transformed by electroporation (Faber et al., 1994). The 500 bp sequences up- and downstream of the OpTEF1 open reading frame (ORF) were used as the OpTEF1 promoter and terminator, respectively and those of the OpCDC28 ORF were used as the OpCDC28 promoter and terminator (Kiel et al., 2007). For overexpression of CDC5, we expressed an N-terminal-truncated version of CDC5 (CDC5∆53) that is equivalent to the ScCDC5∆N70 allele which is resistant to APC-dependent ubiquitination (Shirayama et al., 1998). The CDC5∆53 ORF was amplified by PCR and inserted into pHM949. The resulting plasmid pHM950 was digested and a zeocin resistance marker was inserted (pHM956). To obtain the dyn1∆ strain, tetrad dissection plates were incubated at room temperature for >5 days until colonies were formed because the dyn1∆ cells grew extremely slowly. Glycerol stocks were prepared from the initial master plate of tetrad analysis. YPDS liquid medium was inoculated with either the initial colonies from tetrad dissection or glycerol stocks, and the resulting cells were subjected to analyses.
Yeast strains and plasmids
https://doi.org/10.7554/eLife.24340.037| Name | Genotype/species/construction | Source or reference |
|---|---|---|
| O. polymorpha strains | ||
| BY4329 | leu1-1 | NBRP |
| BY21401 | CBS4732 Type strain | NBRP |
| HPH31 | HHT1::pHM713 ura3-1 | this study |
| HPH41 | ura3-1 pHM719 | this study |
| HPH164 | HHT1::pHM726 TUB1:: pHM737 leu1-1 | this study |
| HPH194 | TUB1::pHM737 leu1-1 | this study |
| HPH206 | Δdyn1::natNT2 leu1-1 | this study |
| HPH207 | Δdyn1::natNT2 leu1-1 | this study |
| HPH221 | wild type | this study |
| HPH222 | Δbub2::hphNT1 | this study |
| HPH223 | Δkar9::natNT2 | this study |
| HPH224 | Δkar9::natNT2 Δbub2::hphNT1 | this study |
| HPH225 | leu1-1 | this study |
| HPH399 | SPC72-GFP-hphNT1 | this study |
| HPH449 | SPC72-mRFP-hphNT1 TUB1:: pHM737 leu1-1 | this study |
| HPH466 | wild type | this study |
| HPH475 | hphNT1-PSPC98-GFP-SPC98 | this study |
| HPH972 | SPC72-GFP-hphNT1 | this study |
| HPH1150 | Δcdc28::natNT2::pHM878 ura3-1 | this study |
| HPH1210 | Δcdc28::natNT2::pHM878 TUB1:: pHM737 HHT1::pHM713 ura3-1 | this study |
| HPH1380 | SPC72-GFP-hphNT1 CDC5-3mAID-natNT2 ura3-1::pHM922 | this study |
| HPH1393 | SPC72-GFP-hphNT1 MPS3-mRFP-kanMX6 ura3-1::pHM859 | this study |
| HPH1394 | SPC72-GFP-hphNT1 MPS3-mRFP-kanMX6 | this study |
| HPH1396 | hphNT1-PNUD1-GFP-NUD1 | this study |
| HPH1400 | SFI1-GFP-hphNT1 | this study |
| HPH1405 | SPC110-GFP-hphNT1 | this study |
| HPH1430 | SPC72-5flag-hphNT1 CDC5-3mAID-natNT2 ura3-1::pHM922 | this study |
| HPH1542 | SPC72-GFP-hphNT1 MPS3-mRFP-kanMX6 TEF1::pHM950::pHM956 | this study |
| HPH1562 | CDC5-GFP-hphNT1 MPS3-mRFP-kanMX6 | this study |
| HPH1564 | CDC5-GFP-hphNT1 MPS3-mRFP-kanMX6 HHT1::pHM726 | this study |
| HPH1575 | CDC5-GFP-hphNT1 SPC72-tdTomato-hphMX | this study |
| HPH1581 | SPC72-tdTomato-hphMX SPC110-GFP-hphNT1 | this study |
| HPH1583 | CDC5-GFP-hphNT1 SPC110-tdTomato-natNT2 | this study |
| HPH1678 | MPS3-GFP-hphNT1 HHT1::pHM713 | this study |
| other yeast strains | ||
| BY21467 | S. cerevisiae YPH499 | NBRP |
| BY21165 | Kluyveromyces lactis NH27 | NBRP |
| BY21167 | Yarrowia lipolytica T22 | NBRP |
| BY23876 | Candida glabrata YAT3377 | NBRP |
| BY5243 | Ogataea parapolymorpha DL-1 | NBRP |
| JCM9829 | Candida peltata | JCM |
| JCM 10237 | Ogataea methanolica | JCM |
| JCM15019 | Ambrosiozyma kashinagacola | JCM |
| plasmids | ||
| pHM713 | pREMI-Z carrying HHT1-GFP and HHF1(histoneH4) | this study |
| pHM719 | pKS144 carrying TUB4-GFP | this study |
| pHM726 | pREMI-Z carrying HHT1-mCherry and HHF1(histoneH4) | this study |
| pHM737 | pRS305 carrying PTUB1-GFP-TUB1 | this study |
| pHM859 | pBSII carring HpURA3 and PTEF1-HpSPC72-GFP | this study |
| pHM878 | pBSII carring HpURA3 and cdc28-as | this study |
| pHM922 | pBSII carring HpURA3 and PCDC28-OsTIR | this study |
| pHM950 | pFA6a-natNT2 carrying PTEF1-HpCDC5Δ53 | this study |
| pHM956 | pFA6a-natNT2 carrying PTEF1-HpCDC5Δ53 and zeo | this study |
Yeast growth conditions and general methods
Request a detailed protocolYeast strains were grown either in YPD medium containing 200 mg/l adenine, leucine, and uracil (YPDS) or in synthetic/defined (SD) medium supplemented with appropriate amino acids and nucleotides (Sherman, 1991). Cells were grown at 30°C unless otherwise indicated. To depolymerize MTs, cells were incubated in either YPDS medium or SD medium containing 1.5 μg/ml nocodazole at 30°C for 1.5 hr.
Microscopy
Request a detailed protocolYeast cells carrying GFP-TUB1, HHT1-mCherry, SPC72-GFP, GFP-NUD1, SFI1-GFP, SPC98-GFP, TUB4-GFP, SPC110-GFP, CDC5-GFP, or MPS3-mRFP were immediately analysed by fluorescence microscopy without washing or fixation in Figures 1A, 2, 4, 5, 6B–E, 7 and 8A, Figure 4—figure supplement 1, Figure 5—figure supplement 1. For the visualisation of DNA with 4'6,-diamidino-2-phenylindole (DAPI), cells were fixed with 70% ethanol, washed with phosphate buffered saline (PBS), and incubated in PBS containing 1 µg/ml DAPI.
Z-series images of 0.4 μm steps were captured with DeltaVision (Applied Precision, Issaquah, WA, USA) equipped with GFP and TRITC filters (Chroma Technology Corp., Bellows Falls, VT, USA), a 100 × NA 1.4 UPlanSApo oil immersion objective (IX71; Olympus, Tokyo, Japan), and a camera (CoolSNAP HQ; Roper Scientific, Trenton, NJ, USA) and were quantified/processed with SoftWoRx 3.5.0 (Applied Precision, Issaquah, WA, USA) or Prism4.3.0 software (Chen et al., 1992; Chen et al., 1996). Deconvolved and projected images are shown. The fluorescence intensity of Spc72-GFP was measured on a plane that has an SPB in focus. Time-lapse experiments of Figures 1A and 2D and that of Figure 4C were carried out in YPDS and SD complete medium respectively on a glass-bottom dish (MatTek, Ashland, MA, USA) coated with concanavalin A (037–08771, Wako, Japan) at room temperature. Z series at 0.4 μm steps were acquired every 3 min for Figures 1A and 4C, or every 30 s for Figure 2D.
For Figure 1B, cells were fixed with 70% ethanol, washed with PBS, and incubated in PBS containing 1 mg/ml DAPI to visualize the DNA (Maekawa and Kaneko, 2014). ImageJ 1.47 (NIH, Bethesda, MD, USA) and Photoshop (Adobe Systems, San Jose, CA, USA) were used to mount the images and to produce merged colour images. No manipulations other than contrast and brightness adjustments were used.
To exclude cells that were non-proliferating from the GFP intensity measurements in Figure 4B, cells were first incubated in YPDS containing Alexa 594 conjugated concanavalin A (Thermo Fisher Scientific, Waltham, MA, USA) until all cells were labelled and then washed once with YPDS and incubated in YPDS for 1 hr prior to image capture. Cells that had lost the label or had a bud with no label were subject to the analyses.
Electron microscopy
Request a detailed protocolCells were mounted on a glass-bottom dish (MatTek) coated with concanavalin A and covered with fixative [2% glutaraldehyde in 0.1 M sodium phosphate buffer (pH 7.2)]. After 1 min, cells were further fixed with fresh fixative for 2 hr at 4°C. After washing with buffer, low melting agarose was applied onto the cells to prevent loss of cells during subsequent procedures. Zymolyase solution (0.4 mg⁄ml zymolyase 100T, Seikagaku Co., Tokyo, Japan) was applied on top of the agarose for 60 min at 37°C, postfixed with 2% OsO4 for 2 hr at room temperature, stained with 1% uranyl acetate for 1 hr, dehydrated with acetone in an ascending series from 50% to 100%, and embedded in epoxy resin. Serial sections of 80 nm thicknesses were obtained, poststained with uranyl acetate and lead citrate, and analysed using a Zeiss EM900 Transmission Electron Microscope at Central Unit Electron Microscopy in the German Cancer Research Center (DKFZ) (ZEISS, Oberkochen, Germany) or a Hitachi H-7500 Transmission Electron Microscope at Research Centre for Ultra-High Voltage Electron Microscopy at Osaka University (Hitachi, Tokyo, Japan).
Cell cycle analysis and growth conditions
Request a detailed protocolFor synchronization, CDC5-3mAID cells were incubated in YPDS containing indole-3-acetic acid (IAA) (45533, Sigma-Aldrich, St. Louis, MO, USA)) for 2.5 hr at 30°C until >80% of cells had a large sized bud to deplete Cdc5 (Nishimura and Kanemaki, 2014). Cells were then washed with pre-warmed growth medium to remove IAA and re-suspended in YPAD medium at 30°C.
Yeast cell extract and immunoblotting
Request a detailed protocolWhole cell extracts were prepared for SDS-PAGE and immunoblotting (Knop et al., 1999; Janke et al., 2004; Meitinger et al., 2016). Samples representing 1–2 OD600 of liquid culture were resuspended in 950 μl 0.29 M NaOH and incubated on ice for 10 min. Then, 150 μl 55% (w/v) trichloroacetic acid was added and the solutions were mixed and incubated for 10 min on ice. After centrifugation the supernatant was removed. The protein pellet was resuspended in high urea buffer (8 M urea, 5% SDS, 200 mM NaPO3 pH 6.8, 0.1 mM EDTA, 100 mM dithiothreitol, and bromophenol blue) and heated at 65°C for 10 min. A sample comprising one-fifth of the total sample amount was loaded for SDS-PAGE (Figure 6A) and western blotting was performed using a standard protocol. For immunoprecipitation, total cell extracts were prepared from logarithmically growing cells in immunoprecipitation buffer (100 mM Tris, pH 8.0, 10 mM EDTA, 150 mM NaCl, 5% glycerol, 0.2 mM NaVO3100 mM β-glycerophosphate, 50 mM NaF, 1 mM PMSF, 1 mM DTT, 1% NP-40, and Complete EDTA-free protease inhibitor cocktail [Roche]). 10 mg of total cell extract was incubated with M2-bound magnetic beads (M8823, Sigma) for 2 hr at 4°C. The beads were washed three times with immunoprecipitation buffer. The bound proteins were subjected to λ phosphatase treatment and then eluted in 30 μl of SDS-PAGE sample buffer by incubated at 37°C for 30 min. 3 μl of eluates were loaded on a Mini-PROTEAN TGX Precast Gels (4561021, BIO-RAD Laboratories, Hercules, CA, USA) and western blotting was performed using a standard protocol. Monoclonal antibodies JL-8 (632381, TaKaRa Bio Clontech, Shiga, Japan) and M2 (F1804, Sigma) were used to detect GFP- and flag-tagged proteins respectively. Plot profile function of ImageJ was used to plot intensity value across a line in Figure 6—figure supplement 2.
Structured illumination microscopy (SIM)
Request a detailed protocolCells were arrested for 2.5 hr with 1.5 µg/ml nocadazole and fixed for 15 min in 4% paraformaldehyde/2% sucrose in phosphate-buffered saline (PBS) solution followed by extensive washing in PBS. The cells were immobilized on a concanavalin A (Sigma-Aldrich, MO, USA)- coated 35 mm glass bottom dish (MatTek, P35G-1.5–14C) and maintained in PBS for the duration of the imaging process in PBS. The samples were imaged in the 2D-SIM mode on a Nikon N-SIM system (Tokyo, Japan) equipped with a TIRF Apochromat 100x/1.49 NA oil immersion objective and a single photon detection EM-CCD camera (Andor iXon3 DU-897E; Belfast, UK). The 488 nm and 561 nm laser lines were used for excitation of yeGFP and tdTomato, respectively, combined with emission band pass filter 520/45 and 610/60. Images were taken sequentially within a small z-stack and in consideration of imaging SPBs close to the coverslip to minimise spherical aberrations. Subsequently the reconstruction and channel alignment was performed using the NIS imaging and image analysis software (Nikon). For the xyz chromatic shift correction we used in a reference sample tetraspeck beads in a reference sample. All images show a single stack of the z-slices.
Data availability
-
Opolymorpha_4329Submitted to the DNA Data Bank of Japan and publicly available under BioProject ID PRJDB3035.
-
Ogataea polymorpha NCYC 495 leu1.1 v2.0The U.S. Department of Energy is committed to making its electronic and information technologies accessible to individuals with disabilities in accordance with Section 508 of the Rehabilitation Act (29 U.S.C. 794d), as amended in 1998.
References
-
Localization of core spindle pole body (SPB) components during SPB duplication in Saccharomyces cerevisiaeThe Journal of Cell Biology 145:809–823.https://doi.org/10.1083/jcb.145.4.809
-
Basic local alignment search toolJournal of molecular biology 215:403–410.https://doi.org/10.1016/S0022-2836(05)80360-2
-
Polo-like kinases: conservation and divergence in their functions and regulationNature reviews. Molecular cell biology 10:265–275.https://doi.org/10.1038/nrm2653
-
Duplication of spindle plaques and integration of the yeast cell cycleCold Spring Harbor Symposia on Quantitative Biology 38:123–131.https://doi.org/10.1101/SQB.1974.038.01.016
-
Behavior of spindles and spindle plaques in the cell cycle and conjugation of Saccharomyces cerevisiaeJournal of Bacteriology 124:511–523.
-
SPOC alert--when chromosomes get the wrong directionExperimental cell research 318:1421–1427.https://doi.org/10.1016/j.yexcr.2012.03.031
-
ConferencePRIISM: an integrated system for display and analysis of 3-D microscope imagesSPIE/IS&T 1992 Symposium on Electronic Imaging: Science and Technology. pp. 784–790.
-
IVE (Image Visualization Environment): a software platform for all three-dimensional microscopy applicationsJournal of Structural Biology 116:56–60.https://doi.org/10.1006/jsbi.1996.0010
-
The yeast spindle pole body component Spc72p interacts with Stu2p and is required for proper microtubule assemblyThe Journal of Cell Biology 141:1169–1179.https://doi.org/10.1083/jcb.141.5.1169
-
Saccharomyces cerevisiae Ndc1p is a shared component of nuclear pore complexes and spindle pole bodiesThe Journal of Cell Biology 143:1789–1800.https://doi.org/10.1083/jcb.143.7.1789
-
Regulation of mitotic spindle orientation: an integrated viewEMBO Reports 17:1106–1130.https://doi.org/10.15252/embr.201642292
-
Profile hidden Markov modelsBioinformatics 14:755–763.https://doi.org/10.1093/bioinformatics/14.9.755
-
Highly-efficient electrotransformation of the yeast Hansenula polymorphaCurrent Genetics 25:305–310.https://doi.org/10.1007/BF00351482
-
The spindle pole body component Spc98p interacts with the gamma-tubulin-like Tub4p of Saccharomyces cerevisiae at the sites of microtubule attachmentThe EMBO Journal 15:3899–3911.
-
Mps3p is a novel component of the yeast spindle pole body that interacts with the yeast centrin homologue Cdc31pThe Journal of Cell Biology 159:945–956.https://doi.org/10.1083/jcb.200208169
-
The budding yeast spindle pole body: structure, duplication, and functionAnnual Review of Cell and Developmental Biology 20:1–28.https://doi.org/10.1146/annurev.cellbio.20.022003.114106
-
Sfi1p has conserved centrin-binding sites and an essential function in budding yeast spindle pole body duplicationThe Journal of Cell Biology 162:1211–1221.https://doi.org/10.1083/jcb.200307064
-
Mechanisms of daughter cell-size control during cell divisionTrends in Cell Biology 25:286–295.https://doi.org/10.1016/j.tcb.2014.12.003
-
Microtubules and actin cytoskeleton in Cryptococcus neoformans compared with ascomycetous budding and fission yeastsEuropean Journal of Cell Biology 80:303–311.https://doi.org/10.1078/0171-9335-00157
-
Control of microtubule dynamics by Stu2p is essential for spindle orientation and metaphase chromosome alignment in yeastMolecular Biology of the Cell 12:2870–2880.https://doi.org/10.1091/mbc.12.9.2870
-
Disruption of mitotic spindle orientation in a yeast dynein mutantPNAS 90:10096–10100.
-
Cloning, sequencing, and functional analysis of H-OLE1 gene encoding delta9-fatty acid desaturase in Hansenula polymorphaApplied Microbiology and Biotechnology 54:499–509.
-
The yeast centrosome translates the positional information of the anaphase spindle into a cell cycle signalThe Journal of Cell Biology 179:423–436.https://doi.org/10.1083/jcb.200705197
-
Astral microtubule asymmetry provides directional cues for spindle positioning in budding yeastExperimental Cell Research 318:1400–1406.https://doi.org/10.1016/j.yexcr.2012.04.006
-
Mitosis in Heterobasidiomycetous Yeasts. 1. Leucosporidium-Scottii (Candida-Scottii)Journal of Cell Science 10:857–881.
-
Mitosis in Heterobasidiomycetous Yeasts. 2. Rhodosporidium Sp (Rhodotorula-Gultinis) and Aessosporon-Salmonicolor (Sporobolomyces-Salmonicolor)Journal of Cell Science 11:1–31.
-
Detection of phosphorylation status of cytokinetic componentsMethods in Molecular Biology 1369:219–237.https://doi.org/10.1007/978-1-4939-3145-3_16
-
Kar9p is a novel cortical protein required for cytoplasmic microtubule orientation in yeastThe Journal of Cell Biology 140:377–390.https://doi.org/10.1083/jcb.140.2.377
-
Rapid depletion of budding yeast proteins via the fusion of an auxin-inducible degron (AID)Current Protocols in Cell Biology pp. 20.9.1–20.920.https://doi.org/10.1002/0471143030.cb2009s64
-
The genome portal of the department of energy joint genome institute: 2014 updatesNucleic Acids Research 42:D26–D31.https://doi.org/10.1093/nar/gkt1069
-
High-voltage electron tomography of spindle pole bodies and early mitotic spindles in the yeast Saccharomyces cerevisiaeMolecular Biology of the Cell 10:2017–2031.https://doi.org/10.1091/mbc.10.6.2017
-
Novel functional dissection of the localization-specific roles of budding yeast polo kinase Cdc5pMolecular and Cellular Biology 24:9873–9886.https://doi.org/10.1128/MCB.24.22.9873-9886.2004
-
Fly meets yeast: checking the correct orientation of cell divisionTrends in Cell Biology 21:526–533.https://doi.org/10.1016/j.tcb.2011.05.004
-
Novel genetic tools for Hansenula polymorphaFEMS Yeast Research 12:271–278.https://doi.org/10.1111/j.1567-1364.2011.00772.x
-
Coordinated spindle assembly and orientation requires Clb5p-dependent kinase in budding yeastThe Journal of Cell Biology 148:441–452.https://doi.org/10.1083/jcb.148.3.441
-
A novel function of Saccharomyces cerevisiae CDC5 in cytokinesisThe Journal of Cell Biology 152:451–469.
-
SPC72: a spindle pole component required for spindle orientation in the yeast Saccharomyces cerevisiaeJournal of Cell Science 111:2809–2818.
-
The Cdc31p-binding protein Kar1p is a component of the half bridge of the yeast spindle pole bodyThe Journal of Cell Biology 128:863–877.https://doi.org/10.1083/jcb.128.5.863
-
Microtubules in the fungal pathogen Ustilago maydis are highly dynamic and determine cell polarityJournal of Cell Science 114:609–622.
-
Analysis of the Saccharomyces spindle pole by matrix-assisted laser desorption/ionization (MALDI) mass spectrometryThe Journal of Cell Biology 141:967–977.https://doi.org/10.1083/jcb.141.4.967
-
Mitotic spindle form and functionGenetics 190:1197–1224.https://doi.org/10.1534/genetics.111.128710
-
MPS1 and MPS2: novel yeast genes defining distinct steps of spindle pole body duplicationThe Journal of Cell Biology 114:745–754.https://doi.org/10.1083/jcb.114.4.745
-
Effect of Auristatin PHE on Microtubule Integrity and Nuclear Localization in Cryptococcus neoformansAntimicrobial Agents and Chemotherapy 46:3802–3808.https://doi.org/10.1128/AAC.46.12.3802-3808.2002
Article and author information
Author details
Funding
Japan Society for the Promotion of Science (JP24570214)
- Hiromi Maekawa
Deutsche Forschungsgemeinschaft (Schi 295/4-3)
- Elmar Schiebel
Deutsche Forschungsgemeinschaft (PE1883)
- Gislene Pereira
Deutsche Forschungsgemeinschaft (SFB873)
- Gislene Pereira
Deutsche Forschungsgemeinschaft (SFB1036)
- Gislene Pereira
Institute for Fermentation, Osaka (the Endowed Chair Program)
- Yoshinobu Kaneko
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
We thank Dr. K Richter and the Central Unit Electron Microscopy at the German Cancer Research Center (DKFZ) for their kind support for the electron microscopy (EM) analyses. The Nikon Imaging Facility Heidelberg is acknowledged for the SIM analyses. We thank Research Center for Ultra-High Voltage Electron Microscopy at Osaka University for supporting our EM analyses. We thank Japan Collection of Microorganisms (JCM) and Dr T Endo for yeast strains, information, and advice on phylogenetic relationships among yeast species. We thank Dr. T Lin for assistance on bioinformatics. Dr. H Takuma, Dr. Yamaguchi, and Integrated Imaging Research Support (IISR) Japan are acknowledged for technical advice and the primary EM analysis, respectively. HM is grateful to Dr. Takegawa for generously allowing access to facilities in the lab. This work was supported by the Endowed Chair Program of the Institute for Fermentation, Osaka (IFO), Japan (YK) and JSPS KAKENHI Grant Number JP24570214 (HM). GP acknowledges funding from the German Research Council (DFG, PE1883) and DFG collaborative programs SFB873 and SFB1036. ES acknowledges funding from the German Research Council (DFG, Schi 295/4-3).
Copyright
© 2017, Maekawa et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 1,203
- views
-
- 181
- downloads
-
- 11
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 11
- citations for umbrella DOI https://doi.org/10.7554/eLife.24340






