Allosteric communication in class A β-lactamases occurs via cooperative coupling of loop dynamics
Figures
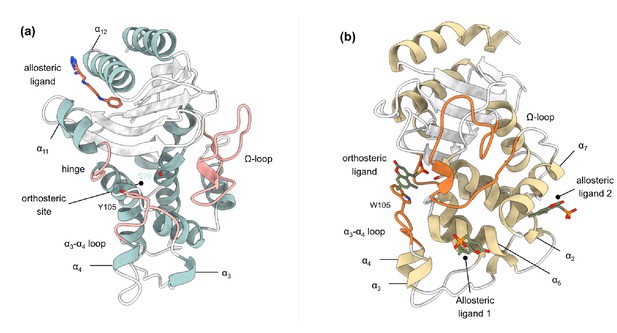
Crystal structures of (a) TEM-1 (PDB id 1PZP) and (b) KPC-2 (PDB id 6D18) β-lactamases in complex with ligands bound to allosteric and the orthosteric sites.
The helices around the allosteric binding sites and the loops that define the orthosteric binding site are highlighted. In case of KPC-2, allosteric ligand 2 is the site investigated here. See Table S1 for structural nomenclature.
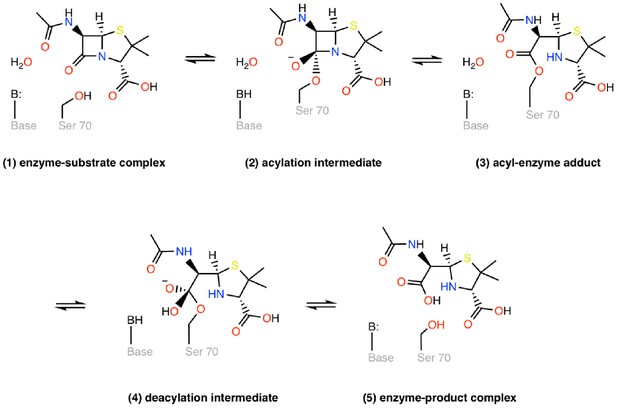
Catalytic cycle of a class A β-lactamase illustrated on the core structure of penicillins.
Class A β-lactamases use an active site serine nucleophile to cleave the β-lactam bond of the substrate in a two-step acylation-deacylation reaction cycle that leads to overall hydrolysis. (1) The acylation reaction initiates with the reversible binding of the antibiotic in the active site and the formation of the enzyme-substrate complex. In the next step, a general base-catalyzed nucleophilic attack on the β-lactam carbonyl by the serine hydroxyl takes place through a tetrahedral intermediate (2) to form a transient acyl-enzyme adduct (3). In the deacylation step, the acyl-enzyme adduct (3) undergoes a general base-catalyzed attack by a hydrolytic water molecule to form a second tetrahedral intermediate (4), which then forms a postcovalent product complex (5), from which the hydrolyzed product is released.
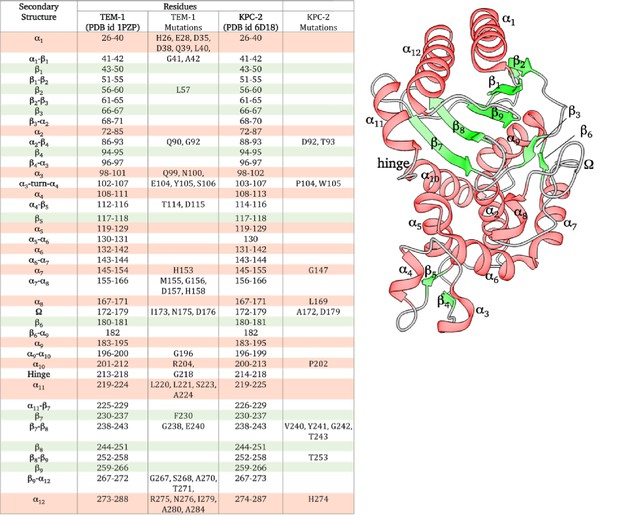
Naming of the loops based on the secondary structure it connects.
The loops are named based on the secondary structure it connects. For example, loop α1-β1 connects α1 helix and β1 sheet. It must be noted that the boundaries of the secondary structure are approximate and may vary by ±2 residues based on the visualization software used. This work employed ChimeraX to define secondary structure. The mutations listed are those that fall on the communication pathway.
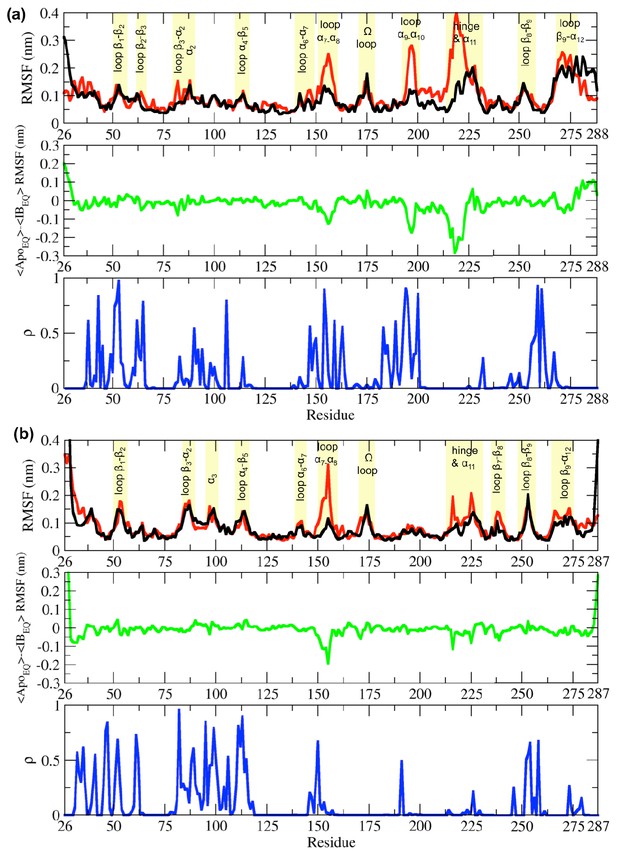
Root-mean-square fluctuation (RMSF) differences between the ApoEQ and IBEQ states of the (a) TEM-1 and (b) KPC-2 systems.
The average change in RMSF in the ApoEQ (black), the IBEQ (red), the difference ApoEQ-IBEQ (green), and the associated ρ value (blue) is illustrated. The ρ values were obtained by conducting a Student’s t-test to compare ApoEQ and IBEQ systems and to assess the significance of the differences.
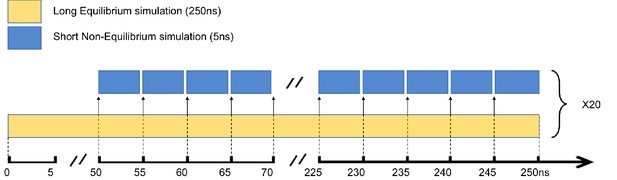
Schematic description of the long equilibrium (EQ) and short nonequilibrium (NE) simulations.
Twenty replicates of IBEQ simulations were run starting from the minimized crystal structure. From the equilibrated part of each replica (50 ns onward), the final conformation of the protein-ligand complex was extracted at every 5 ns, the perturbation (removal of the ligand) was introduced, and a short ApoNE simulations was run for 5 ns. In total, 800 ApoNE simulations were performed for each system. In addition to these simulations, 20 replicas of the ApoEQ were also simulated.
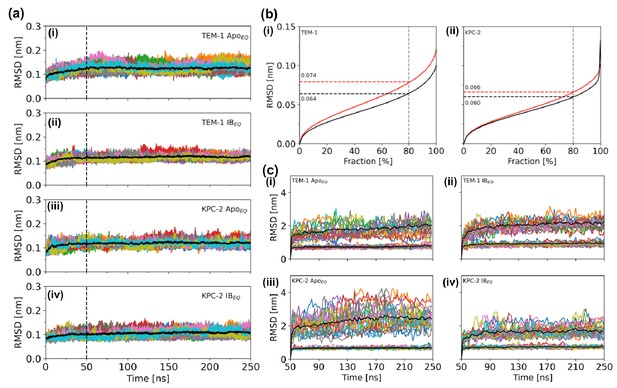
Conformational drift measured by Cα root mean square deviation.
(a) Time series of the Cα root-mean-square deviation (RMSD) of (i) TEM-1 ApoEQ, (ii) TEM-1 IBEQ, (iii) KPC-2 ApoEQ, and (iv) KPC-2 IBEQ systems, measured over the course of the 250 ns of each replica. The black line represents the average of the 20 replicates. (b) RMSD calculated as a function of the fraction of the total Cα atoms considered for structural alignment in (i) TEM-1 and (ii) KPC-2. The plots indicate that 80% of conformations in TEM-1 can be aligned to below 0.064 nm (ApoEQ; black) and 0.074 nm (IBEQ; red). Similarly, in KPC-2, 80% of the conformations could be superimposed to below 0.060 nm (ApoEQ; black) and 0.066 nm (IBEQ; red). This constitutes the core of the enzyme. (c) Fractional Cα RMSD calculated after identification of the core in (i) TEM-1 ApoEQ, (ii) TEM-1 IBEQ, (iii) KPC-2 ApoEQ, and (iv) KPC-2 IBEQ. The bottom black line denotes the stable core and consists of 80% Cα atoms. The top black line is the average of the remainder 20% Cα atoms calculated from all 20 replicates. These constitute the non-core Cα atoms that display deviation in all simulations.
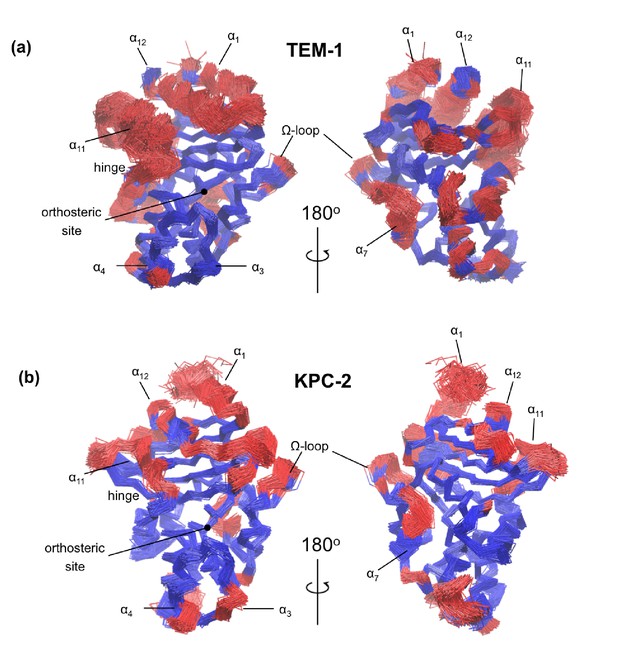
Core Cα root-mean-square deviation (RMSD) superimposition from (a) TEM-1 and (b) KPC-2 IBEQ simulations.
The structural alignment was calculated from the equilibrated section of all IBEQ trajectories and rendered to illustrate 100 uniformly separated frames. The least mobile Cα atoms are colored blue and the most mobile atoms (red) provide the structural basis for the differential RMSDs.
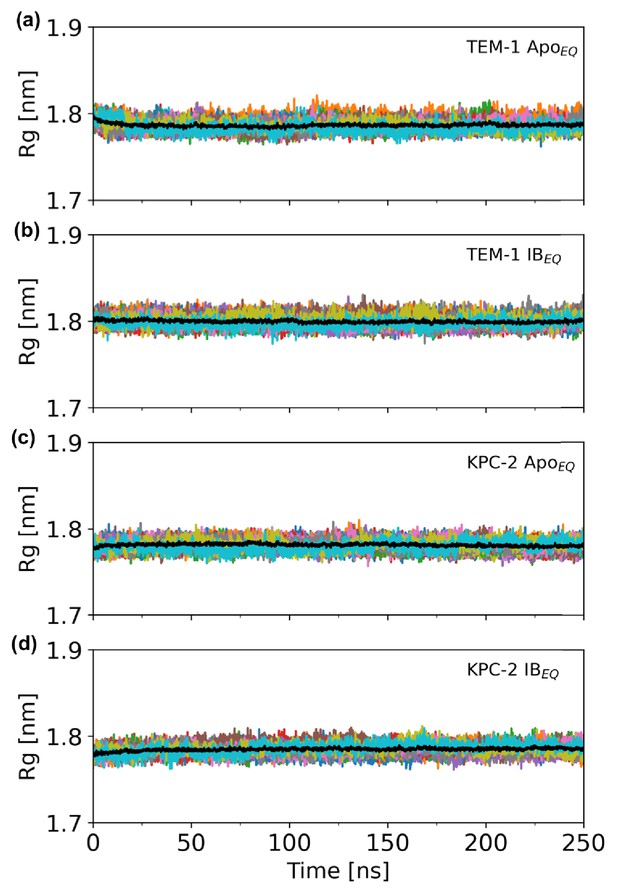
Time evolution of the radius of gyration (Rg) over the course of the 250 ns of each replicate.
Time evolution of the Rg of the complete (a) TEM-1 ApoEQ, (b) TEM-1 IBEQ, (c) KPC-2 ApoEQ, and (d) KPC-2 IBEQ enzymes, measured over the course of the 250 ns of each replicate. The black line represents the average of 20 replicates.
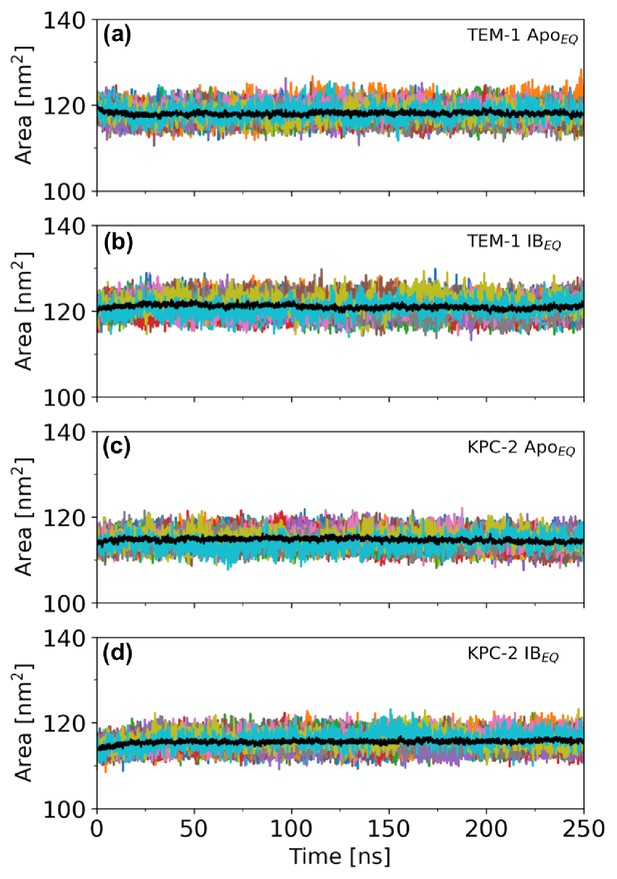
Solvent accessible surface area (SASA) over the course of the 250 ns of each replicate.
SASA was calculated to assess structural distortion in the complete (a) TEM-1 ApoEQ, (b) TEM-1 IBEQ, (c) KPC-2 ApoEQ, and (d) KPC-2 IBEQ enzymes, measured over the course of the 250 ns of each replicate. The black line represents the average of 20 replicates.
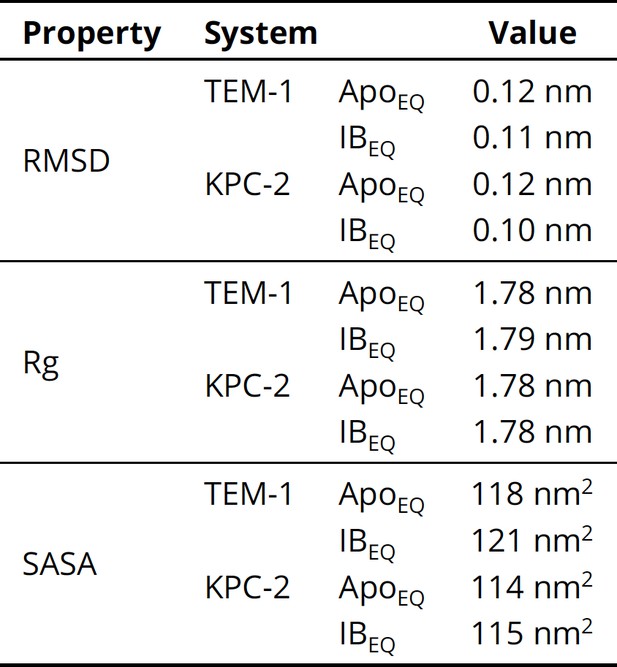
Dynamical properties (root-mean-square deviation [RMSD], radius of gyration [Rg], and solvent accessible surface area [SASA]) used to assess structural stability of the systems over the course of the equilibrium simulation.
The values represent the averages calculated from the equilibrated part (50–250 ns) of all 20 replicate simulations of each system.
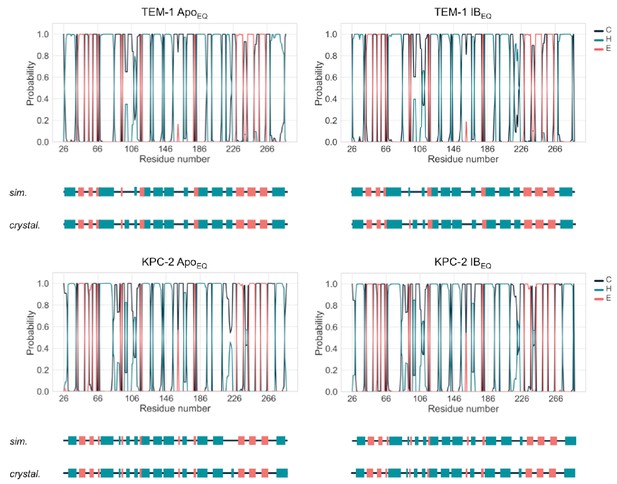

Probability to find each residue in a coil, helix, or strand over the course of the 250 ns of each replicate.
Probability to find each residue in a coil (C), helix (H), or strand (E). The secondary structure element was assigned to each residue in each frame of each of the 20 replicas per system using the DSSP algorithm as implemented in MDTraj python library. The secondary structure of TEM-1 and KPC-2 as assigned from the crystal structure (cryst.), as well as the most probable assignment according to the simulations (sim.) is depicted as a cartoon below each plot.
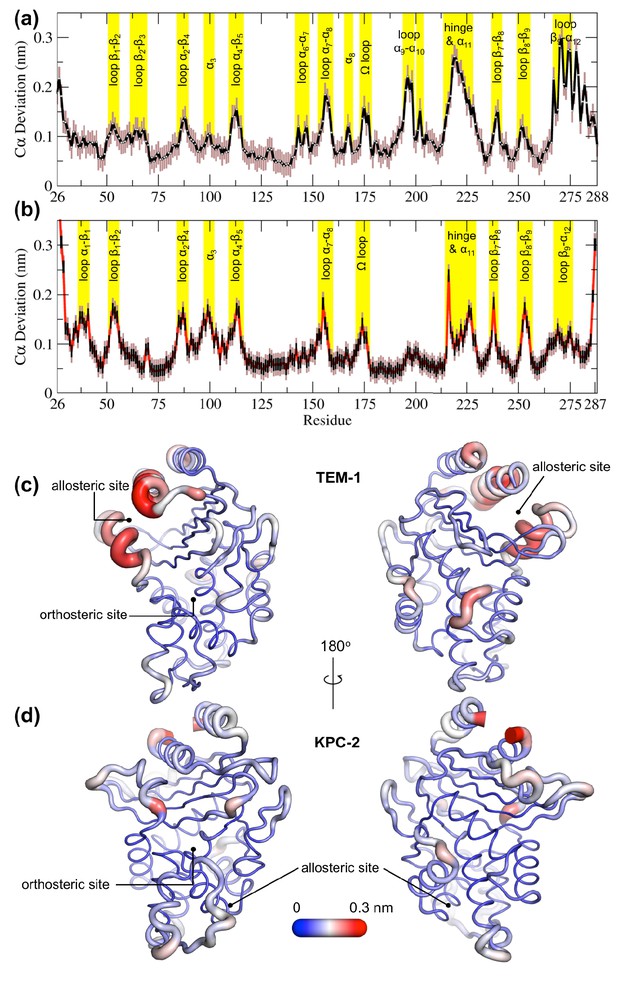
Average positional Cα deviations between the ApoEQ and IBEQ states of (a) TEM-1 and (b) KPC-2.
Important structural motifs are highlighted and labeled on the plots. The brown vertical lines represent the standard deviation of the mean. The averaged Cα positional deviations mapped onto the averaged ApoEQ structures of (c) TEM-1 and (d) KPC-2 to visualize the largest relative displacements. The average deviation was determined from a combination of all 20 ApoEQ and 20 IBEQ trajectories. The thickness of the cartoon corresponds to the Cα deviation.
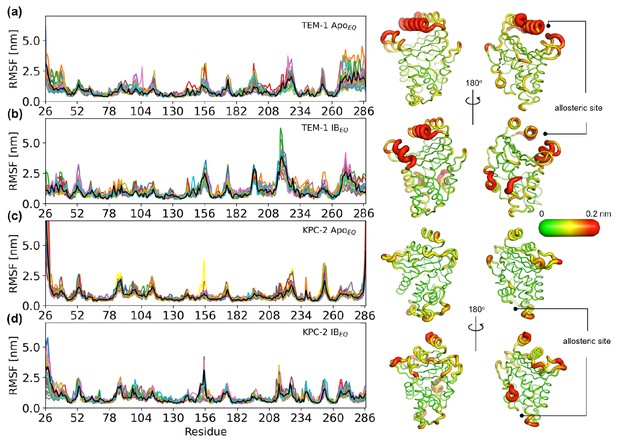
Positional Cα root-mean-square fluctuation (RMSF) of (a) TEM-1 ApoEQ, (b) TEM-1 IBEQ, (c) KPC-2 ApoEQ, and (d) KPC-2 IBEQ systems.
The black line represents the average RMSF calculated from the last 200 ns of all 20 replicate simulations. This average value is mapped onto the structure to highlight regions of high flexibility.
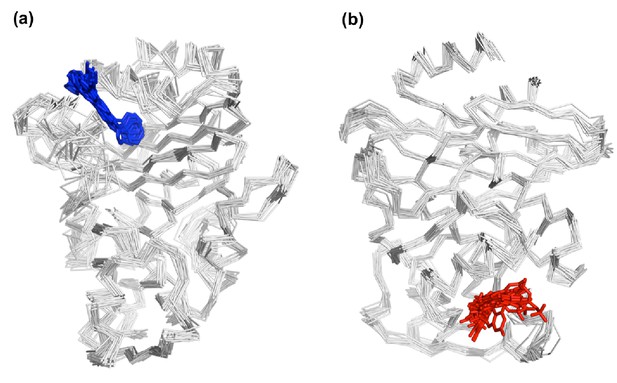
Snapshot of the last frame from TEM-1 IBEQ and KPC-2 IBEQ replicate simulations.
Snapshot of the last frame from (a) TEM-1 IBEQ and (b) KPC-2 IBEQ replicate simulations, highlighting the spatial position of the ligands in the allosteric binding sites. FTA is represented as blue sticks and GTV is colored in red.
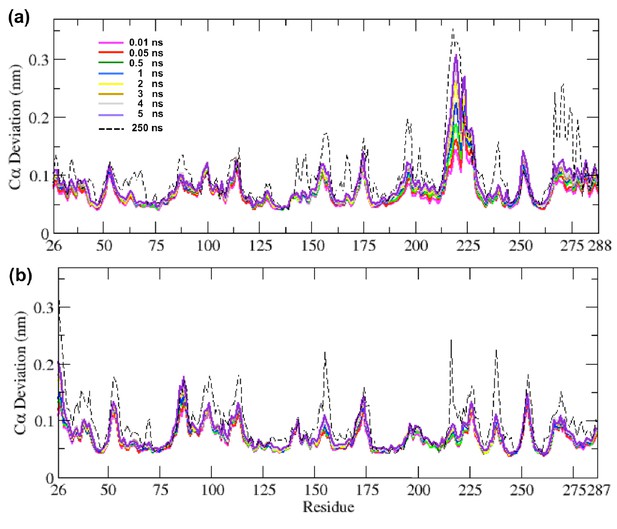
Average Cα deviation between the IBEQ and ApoNE calculated using the subtraction method for (a) TEM-1 and (b) KPC-2.
Average from all 800 simulations at various time points is illustrated. The average Cα deviation between the IBEQ and ApoNE simulations is plotted as a dotted line for comparison.
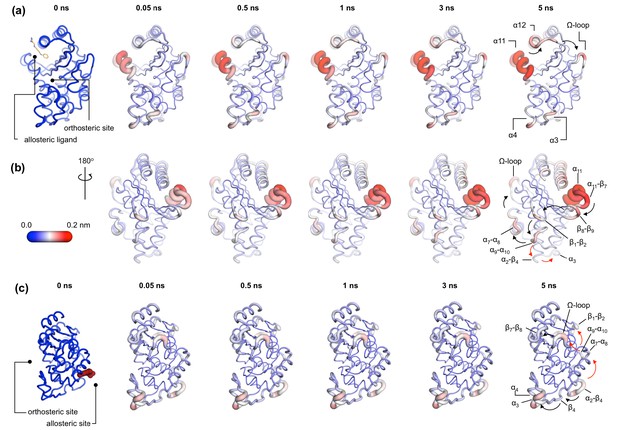
Communication pathways in (a, b) TEM-1 and (c) KPC-2.
The average Cα deviations correspond to the average difference in the position of each Cα atom between all 800 pairs of IBEQ and ApoNE simulations at specific time points. The averaged Cα deviations are mapped onto the average ApoEQ structure. The arrows mark the direction of the propagation of the signal, caused by the perturbation (removal of the ligand). The red and the black arrows highlight different paths taken by the propagating signals (also see movies Figure 4—video 1, Figure 4—video 2, Figure 4—video 3).
Signal propagation in TEM-1 (front view).
Signal propagation in TEM-1 (back view).
Signal propagation in KPC-2.
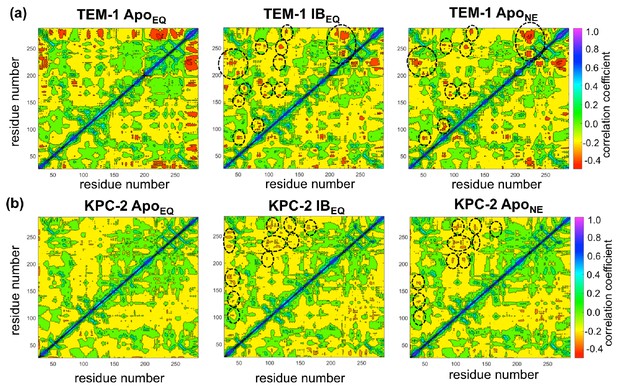
Dynamic cross-correlation maps (DCCMs) computed for (a) TEM-1 and (b) KPC-2 Apo equilibrium (ApoEQ), inhibitor-bound equilibrium (IBEQ), and Apo nonequilibrium (ApoNE) trajectories.
The DCCMs for equilibrium trajectories were calculated as an average of 20 replica simulations, while the ApoNE DCCM indicates an averaged DCCM from an ensemble of 40 short (5 ns) MD trajectories. Green regions indicate no correlation, yellow indicates moderate negative correlation, while orange and red indicate significant negative correlations and blue regions indicate positive correlations. In TEM-1 ApoNE, regions showing significant changes from ApoEQ and IBEQ bound simulations have been marked by black dashed ellipses.
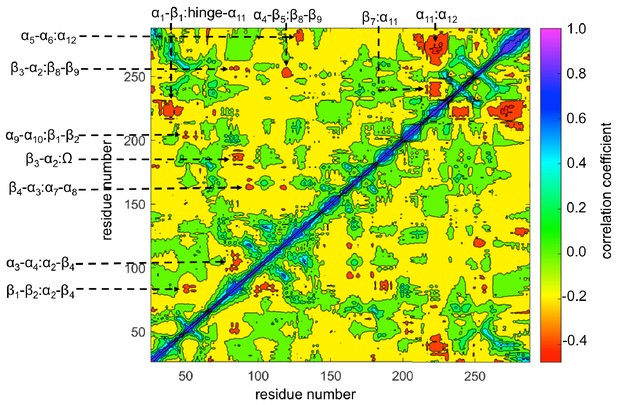
TEM-1 averaged dynamic cross-correlation map (DCCM) computed from all nonequilibrium trajectories.
The regions showing significant correlations are identified: β1-β2:α2-β4, α3-α4:α2-β4, β4-α3:α7-α8, β3-α2:Ω, α9-α10:β1-β2, β3-α2:β8-β9, α5-α6:α12, α1-β1:hinge-α11, α4-β5:β8-β9, β7:α11, and α11:α12.
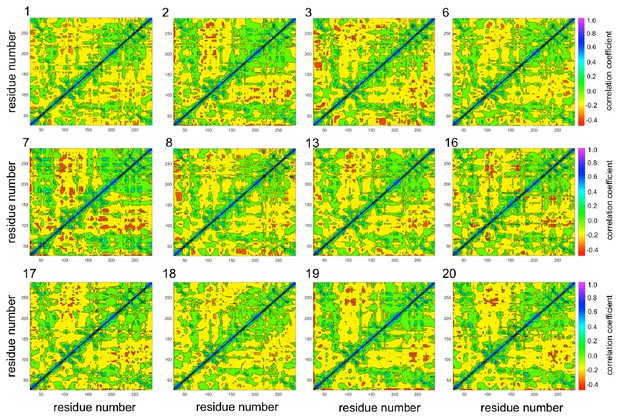
Selected dynamic cross-correlation maps (DCCMs) computed for individual 5 ns nonequilibrium molecular dynamics (MD) trajectories of KPC-2.
These individual trajectories show different behavior from the averaged results (see Figure 5), as these trajectories show several regions of significant correlations similar to the TEM-1 nonequilibrium DCCM.
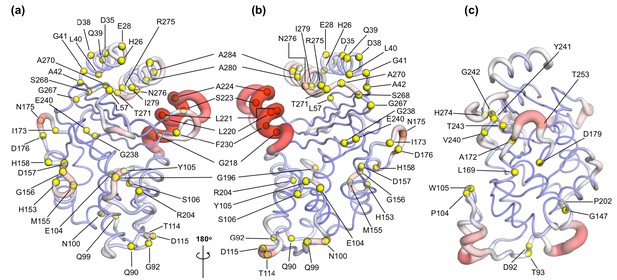
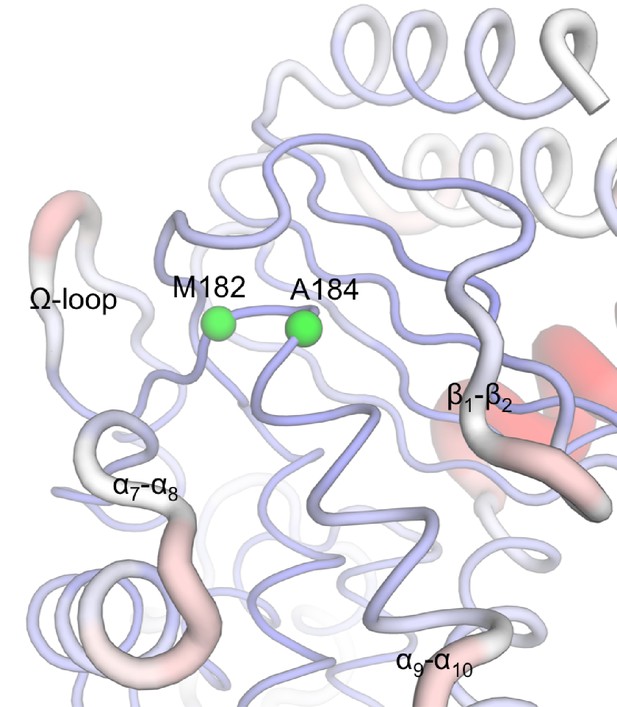
Variant positions in (a, b) TEM-1 and (c) KPC-2 mapped onto the averaged ApoEQ structures, also showing allosteric communication pathways (see Figure 4) identified by nonequilibrium simulations.
The position of the variant is shown as yellow spheres centered at the corresponding Cα. Only the sites of mutations that lie on the allosteric communication pathways have been annotated. The color scheme and cartoon thickness of the rendered structures represents a snapshot of average Cα deviation between IBEQ and ApoNE. Many of these clinically important variant positions lie on the allosteric communication pathway: 45 of the 90 for TEM-1, 15 out of the 25 for KPC-2 single point variants lie on the pathways. This suggests that these variations affect the allosteric behavior of the enzymes.