PHAROH lncRNA regulates Myc translation in hepatocellular carcinoma via sequestering TIAR
Figures
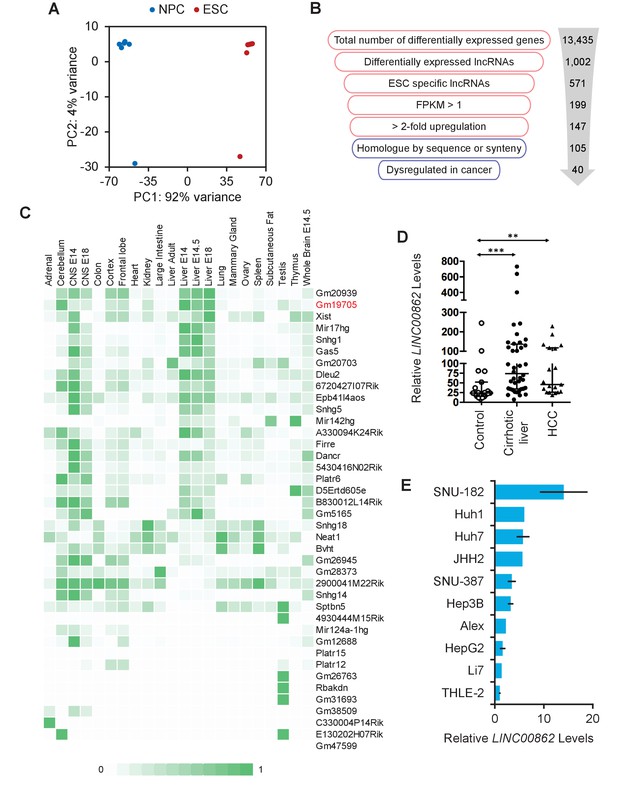
Long non-coding RNA (lncRNA) screen to identify transcripts enriched in embryonic stem cells (ESCs) and dysregulated in cancer.
(A) Principal component analysis (PCA) plot of 10 RNA-seq libraries from mouse-derived ESCs, and two from cell lines. Differentiation from ESCs to neural progenitor cells (NPCs) created the largest difference in variance, while there was minimal difference between isolated clones vs. cell lines. (B) Workflow of the filtering process performed to obtain ESC-enriched lncRNAs that are also dysregulated in cancer. Red indicates analysis performed in mouse and blue indicates human. (C) lncRNA candidate expression across ENCODE tissue datasets show that lncRNAs are mostly not pan-expressed, but are rather tissue specific. Counts are scaled per row. (D) LINC00862 is upregulated in both human cirrhotic liver and hepatocellular carcinoma (HCC) tumor samples when compared to control patient liver tissue samples. **p<0.01; ***p<0.005; Student’s t-test. (E) LINC00862 is upregulated in various human HCC cell lines.
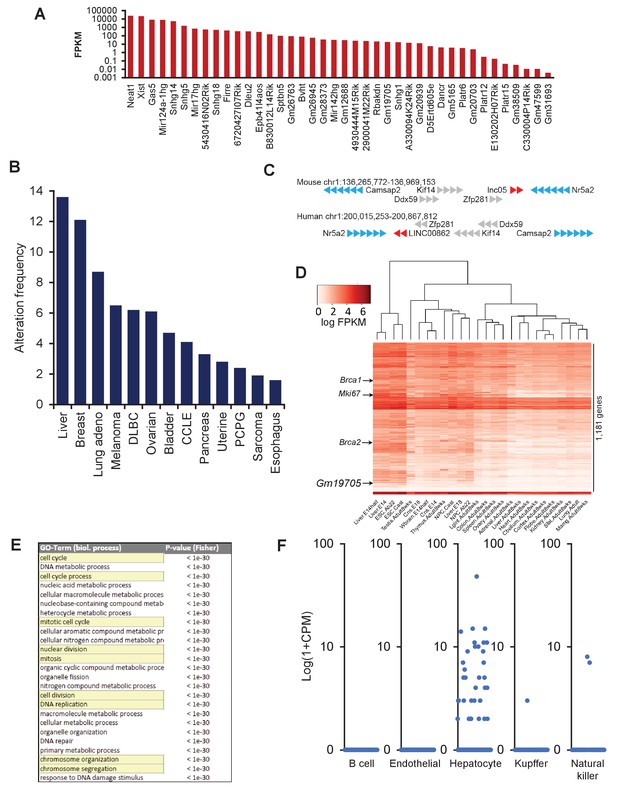
LncRNA screen to identify transcripts enriched in ESCs and dysregulated in cancer, related to Figure 1.
(A) Long non-coding RNA (lncRNA) screen identifies candidates with varying levels of expression in embryonic stem cells (ESCs). (B) LINC00862 is altered in 13% of all hepatocellular carcinoma (HCC) patient cases according to TCGA data. (C) Gm19705 gene locus on chromosome 1 shows that the order of the genes is conserved between mouse and human, but the order is reversed, suggesting a reversal event occurrence. (D) Weighted gene correlation network analysis of Gm19705 places it in a module with cell cycle genes and proliferation genes, such as Brca1/2, and Mki67. (E) GO term analysis of the module containing Gm19705 shows enrichment of genes related to cell cycle, mitosis, and DNA replication. (F) Reanalysis of single-cell data of adult liver (Tabula Muris Consortium et al., 2018) reveals expression of Gm19705 is highly enriched in hepatocytes, but only a subset of the cells.
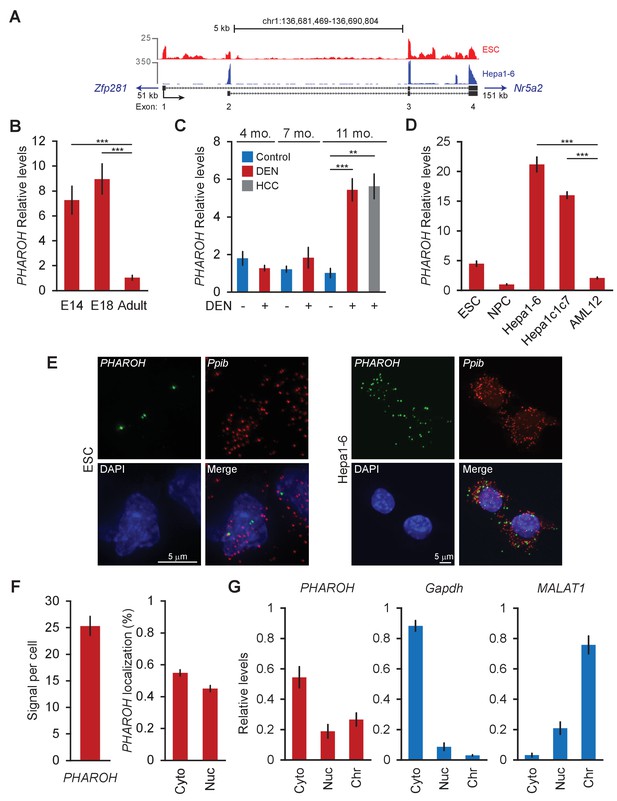
PHAROH long non-coding RNA (lncRNA) is highly expressed in embryonic stem cells (ESCs), embryonic liver, models of hepatocarcinogenesis, and hepatocellular carcinoma (HCC) cell lines.
(A) 5′ 3′ rapid extension of cDNA ends (RACE) reveals two isoforms for PHAROH, which have exons 3 and 4 in common. PHAROH is an intergenic lncRNA where the nearest upstream gene is Zfp218 (51 kb away), and downstream is Nr5a2 (151 kb away). RNA-seq tracks of ESC (red) and Hepa1-6 (blue) cells show cell-type-specific isoform expression of PHAROH. (B) PHAROH is highly expressed in embryonic liver in E14 and E18 mice, but not adult liver (**p<0.01; ***p<0.005; Student’s t-test). (C) A diethylnitrosamine (DEN) model of hepatocarcinogenesis shows high upregulation of PHAROH in the liver and HCC tumor nodules (gray bar) in DEN-treated mice (**p<0.01; ***p<0.005; Student’s t-test). (D) PHAROH is upregulated in HCC cell lines (Hepa1-6 and Hepa1c1c7) compared to normal mouse hepatocytes (AML12) (***p<0.005; Student’s t-test). (E) Single-molecule RNA-FISH of PHAROH in ESCs shows nuclear localization and an average of 3–5 foci per cell. In Hepa1-6 cells, PHAROH shows 25 foci per cell, distributed evenly between the nucleus and cytoplasm (n = 75 cells for each sample). Ppib is used as a housekeeping protein coding gene control. (F) Quantitation of panel PHAROH foci in panel (E) in HepA1-6 cells. (G) Cellular fractionation of Hepa1-6 cells shows equal distribution of PHAROH in the cytoplasm and nucleus, where it also binds to chromatin. Gapdh is predominantly cytoplasmic, and MALAT1 is bound to chromatin.
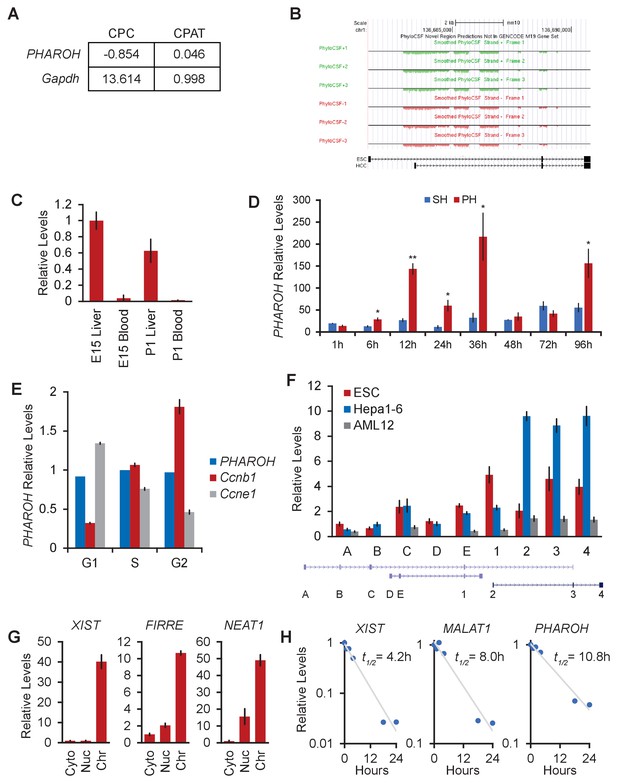
PHAROHlncRNA is highly expressed in ESCs, embryonic liver, models of hepatocarcinogenesis, and HCC cell lines, related to Figure 2.
(A) CPC and CPAT coding potential analysis for PHAROH and Gapdh. (B) PhyloCSF tracks showing low coding potential for the PHAROH locus. (C) PHAROH is expressed in fetal liver, but not in the blood. (D) Sham hepatectomy (SH) or partial hepatectomy (PH) of the liver, a model of liver regeneration, shows upregulation of PHAROH during time points of concerted cell division. *p<0.05; **p<0.01; ***p<0.005; Student’s t-test. (E) HepA1-6 cells were stained with Hoechst 33258 and sorted according to their cell cycle phase. qRT-PCR analysis shows that PHAROH does not cycle with the cell cycle, unlike Ccnb1 and Ccne1. (F) qRT-PCR of each annotated exon spanning the current Gencode M20 annotation. Exons 1–4, which are numbered similarly as Figure 2A, are confirmed rapid extension of cDNA ends (RACE) exons. Isoform with exons 1, 3, and 4 is embryonic stem cell (ESC) specific, and the isoform with exons 2–4 is hepatocellular carcinoma (HCC) specific. Exons A, B, C, D, and E are currently annotated exons, but not identifiable via RACE. (G) XIST, FIRRE, and NEAT1 serve as additional controls for the cellular fractionation. (H) Calculated RNA half-life based upon α-amanitin-treated cells. PHAROH has a half-life of 10.8 hr, longer than that of XIST and MALAT1.
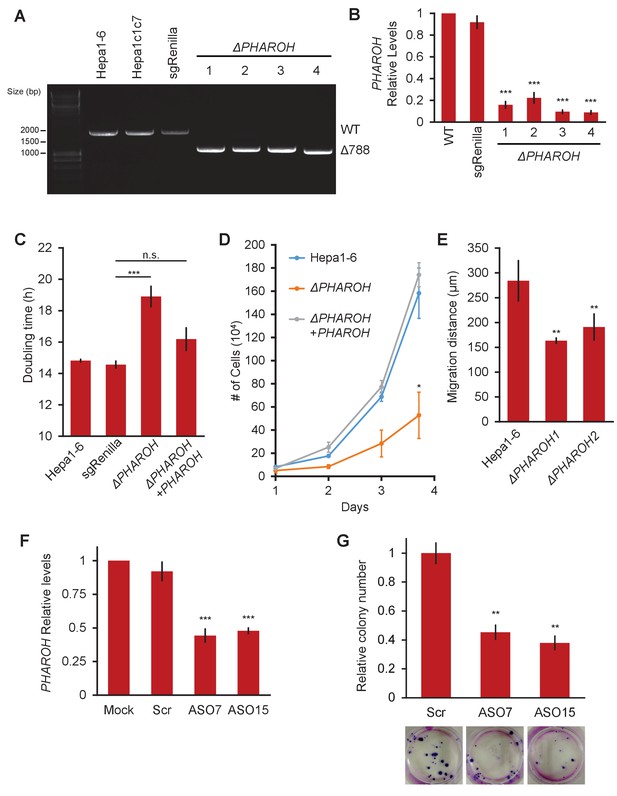
Depletion of PHAROH results in a proliferation defect.
(A) Four isolated clones all have a comparable deletion of 788 bp. The wildtype band is ~1.8 kb. (B) qRT-PCR of PHAROH knockout clones show a >80% reduction in PHAROH levels (***p<0.005; Student’s t-test). (C) Aggregated doubling time of clones shows knockout of PHAROH increases doubling time from 14.8 hr to 18.6 hr. Addition of PHAROH back into knockouts rescues this defect (***p<0.005; Student’s t-test). (D) Manual cell counting shows proliferation defect in PHAROH knockout cells that is rescued by ectopic expression of PHAROH (*p<0.05; Student’s t-test). (E) Migration distance for PHAROH knockout clones is decreased by 50% (**p<0.01; Student’s t-test). (F) 50% knockdown of PHAROH can be achieved using both antisense oligonucleotide (ASO)7 and ASO15 at 24 hr (***p<0.005; Student’s t-test). (G) Colony formation assay of Hepa1-6 cells that are treated with scrambled or PHAROH targeting ASOs. After seeding 200 cells and 2 weeks of growth, a 50% reduction in relative colony number is observed (**p<0.01; Student’s t-test).
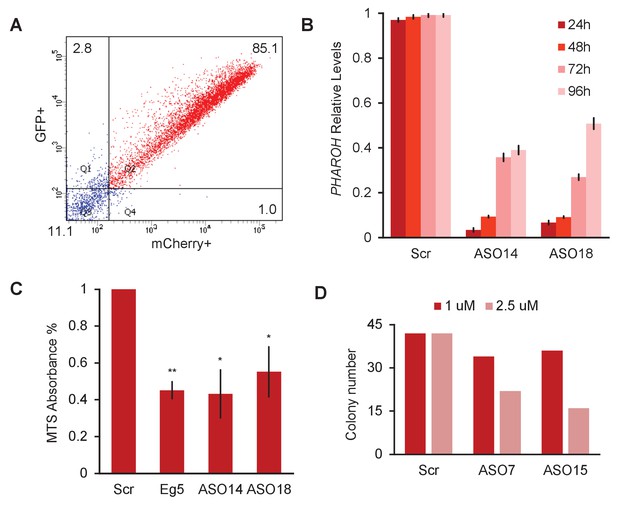
Depletion ofPHAROHresults in a proliferation defect, related to Figure 3.
(A) FACS for double GFP+/mCherry+ cells shows an 85.1% nucleofection efficiency for both plasmids. (B) Knockdown of PHAROH using nucleofection of 2 μM antisense oligonucleotide (ASO) is effective over 96 hr. (C) MTS assay for proliferation 96 hr after nucleofection. MTS absorbance is reduced by 50% in ASO-treated samples targeting PHAROH and Eg5. (D) Reduction of colony formation number is dose dependent.
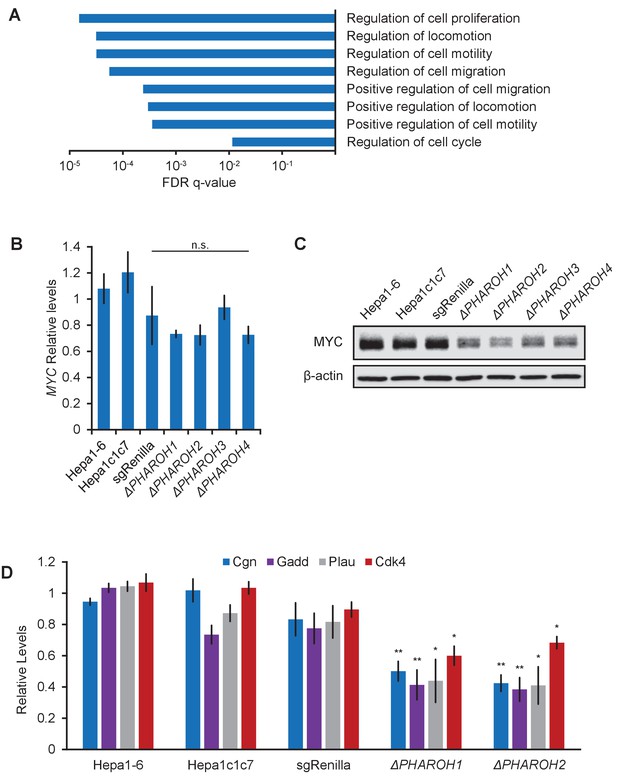
Gene expression analysis of PHAROH knockout cells reveals a link to MYC.
(A) GO term analysis of differentially expressed genes shows enrichment of cell proliferation and migration genes. (B) qRT-PCR of Myc mRNA levels indicates that Myc transcript does not appreciably change when PHAROH is knocked out. (C) Western blot analysis of MYC protein shows downregulation of protein levels in PHAROH knockout cells. β-Actin is used as a loading control. (D) qRT-PCR of genes downstream of Myc shows a statistically significant decrease in expression (*p<0.05; **p<0.01; Student’s t-test).
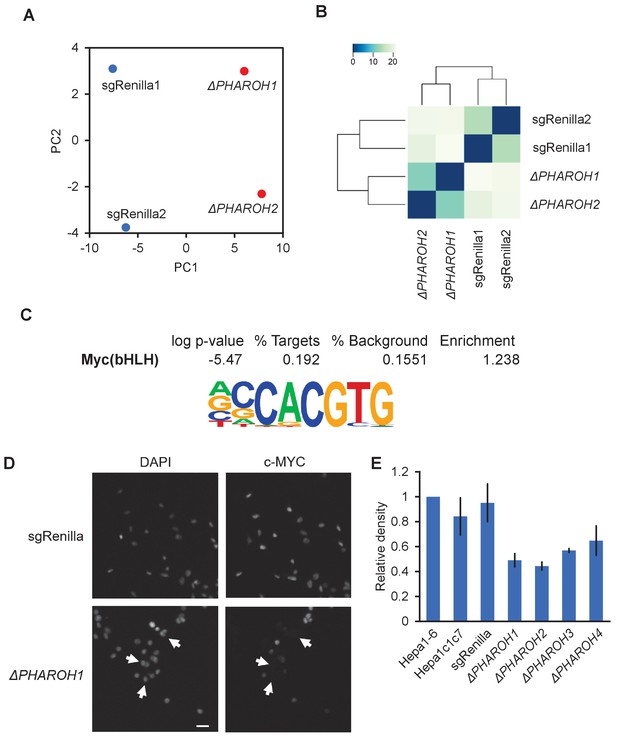
Gene expression analysis ofPHAROHknockout cells reveals a link to MYC, related to Figure 4.
(A) Principal component analysis of two sgRenilla-negative control clones and two PHAROH knockout clones. Deletion of PHAROH is well separated by PC1. (B) Euclidean distance plot indicating that the negative control clones and PHAROH knockout clones cluster independently. (C) Motif analysis of promoter region of differentially expressed genes. MYC motif is enriched 1.24-fold over background sequences. (D) Immunofluorescence of MYC in PHAROH knockout clones shows absence of MYC signal in a majority of cells. Scale bar = 50 μm. (E) Quantification of western blot in Figure 4C.
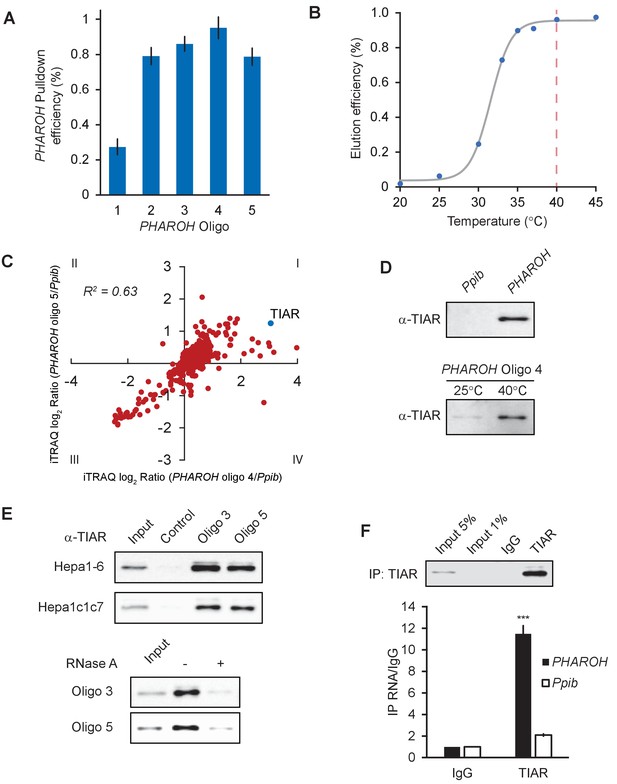
RNA antisense purification-mass spectrometry (RAP-MS) identifies TIAR as a major interactor of PHAROH.
(A) Five different biotinylated oligos antisense to PHAROH were screened for pulldown efficiency. Oligos 2–5 can pull down PHAROH at ~80% efficiency or greater. (B) PHAROH can be eluted at a specific temperature. Maximum elution is reached at 40°C. (C) iTRAQ results using two different oligos targeting PHAROH compared to PPIB reveal nucleolysin TIAR as the top hit. (D) TIAR is pulled down by PHAROH oligos and is specifically eluted at 40°C, but not by PPIB oligos. (E) TIAR can be pulled down using additional oligos and in two different cell lines. RNase A treatment of the protein lysate diminishes TIAR binding to PHAROH, indicating that the interaction is RNA-dependent. (F) Immunoprecipitation of TIAR enriches for PHAROH transcript when compared to IgG and PPIB control (***p<0.005; Student’s t-test).
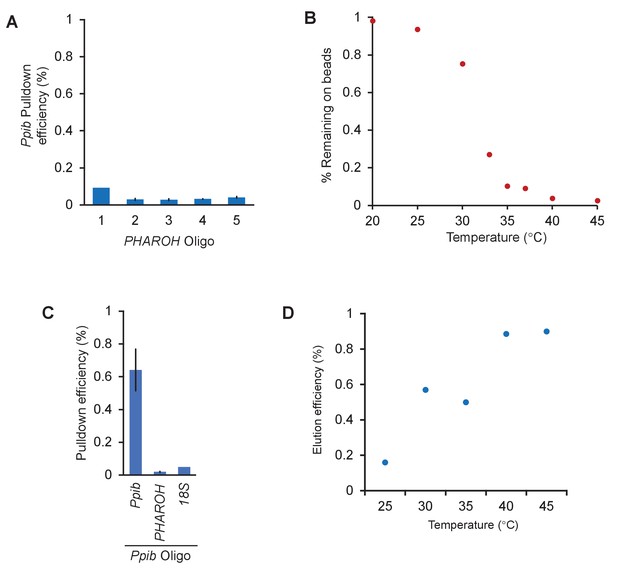
RNA antisense purification-mass spectrometry (RAP-MS) identifies TIAR as a major interactor of PHAROH, related to Figure 5.
(A) The amount of PHAROH RNA remaining on the beads after thermal elution is inverse to that of the eluate. (B) Off-target pulldown of Ppib using PHAROH oligos is low. (C) An oligo designed against Ppib can pull the RNA down at ~65% efficiency and does not pull down PHAROH or 18S. (D) Ppib can also be eluted via a temperature gradient and is optimally released at 40°C.
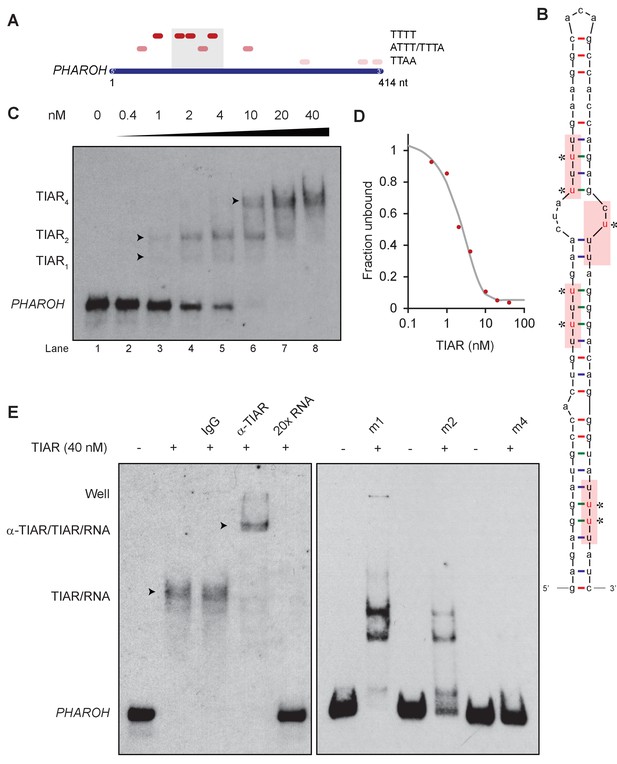
TIAR binds to the 5′ end of PHAROH.
(A) Sequence analysis of PHAROH with published TIAR binding motifs shows a preference for the 5′ end of PHAROH. (B) Schematic of the conserved hairpin of PHAROH that contains four potential TIAR binding sites indicated in the red boxes. Mutations created within the PHAROH hairpin are indicated in red asterisks. (C) RNA electromobility shift assay (EMSA) of the 71-nt PHAROH hairpin with human recombinant TIAR shows three sequential shifts as TIAR concentration increases. (D) Densitometry analysis of the free unbound probe estimates the dissociation constant of TIAR as ~2 nM. (E) TIAR/PHAROH binding is specific as a supershift is created when adding antibody against TIAR, and the interaction can be competed out using 20× unlabeled RNA. RNA EMSA of the mutant hairpins reveals decreasing affinity for TIAR. Mutants were made in a cumulative 5′ to 3′ fashion. M1 shows high signal of single and double occupancy forms, and m2 has reduced signal overall. When all four sites are mutated, binding is nearly abolished.
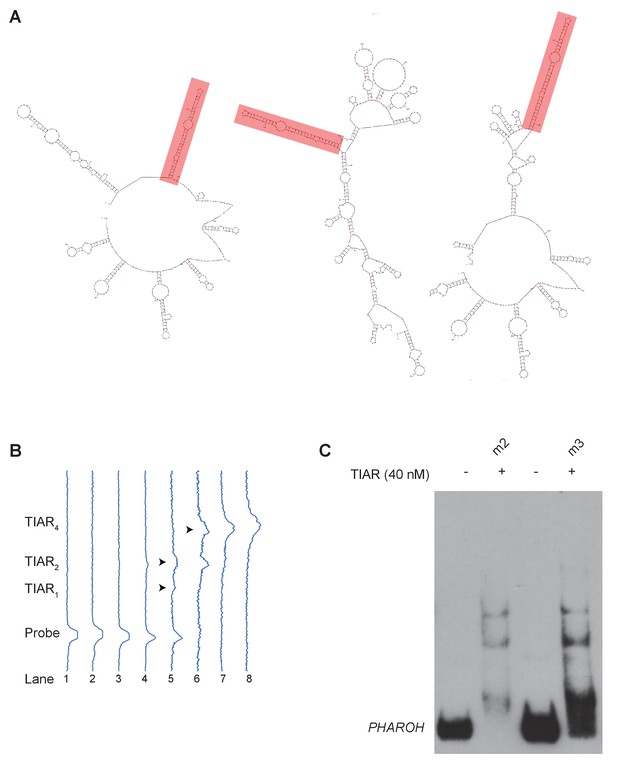
TIAR binds to the 5’ end of PHAROH, related to Figure 6.
(A) Mapping the top seven binding sites to predicted structures (top three shown here) reveals a conserved hairpin on the majority of predicted structures. (B) Profile analysis of the RNA electromobility shift assay gel in Figure 6C, showing the shift in intensity. (C) Binding of TIAR to m2 and m3 is similar, possibly due to the mutation of a weaker binding site does not greatly impact overall binding.
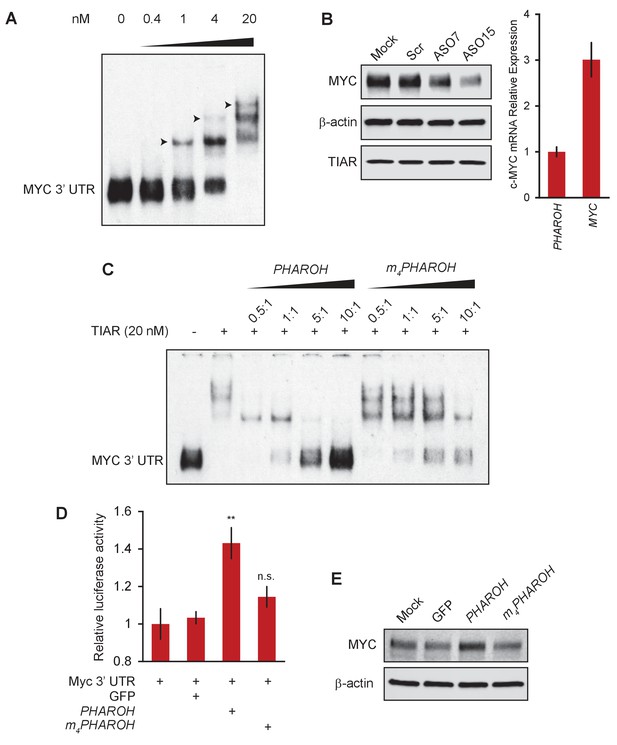
Loss of PHAROH releases TIAR, which inhibits Myc translation.
(A) RNA electromobility shift assay of the 53-nt Myc 3′ UTR fragment shows that TIAR has three potential binding sites, but prefers a single binding event (note arrows). (B) Knockdown of PHAROH reduces MYC protein levels, but not TIAR levels, even though MYC is expressed threefold higher than PHAROH. (C) Wildtype PHAROH hairpin is able to compete out the MYC-TIAR interaction, but the mutated hairpin is not as effective in competing with the Myc-TIAR interaction. (D) Luciferase activity is increased with the addition of PHAROH but not with m4PHAROH (**p<0.01; Student’s t-test). (E) Overexpression of PHAROH increases MYC protein expression, but overexpression of m4PHAROH does not change MYC levels appreciably.
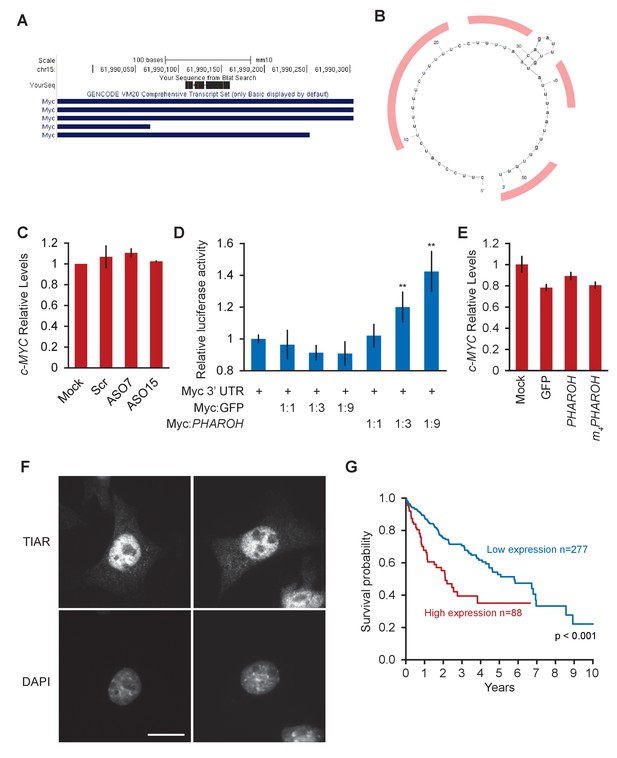
Loss ofPHAROHreleases TIAR, which inhibits Myc translation, related to Figure 7.
(A) Of the two TIAR binding sites on MYC’s 3′ UTR, only one maps to the mouse genome. (B) Potential TIAR binding sites on the mouse Myc 3′ UTR highlighted in red. (C) Knockdown of PHAROH does not change Myc mRNA levels, suggesting that PHAROH acts at a post-transcriptional level. (D) Addition of PHAROH to a luciferase construct with a Myc 3′ UTR increases luciferase activity in a dose-dependent manner. (E) MYC RNA levels do not change when PHAROH or TIAR are overexpressed. (F) IF microscopy of TIAR showing predominantly nuclear localization. Scale bar = 25 μm. (G) Kaplan–Meier survival plot of patients with low and high TIAR expression.
Tables
40 LncRNAs that are enriched in ESCs and dysregulated in cancer.
| Gene name | Sequence homology | Synteny | Human homologue |
|---|---|---|---|
| Platr15 | - | + | LOC284798 |
| 4930444M15Rik | 64.4% of bases, 99.9% of span | + | In TUSC8 region |
| 5430416N02Rik | 16.6% of bases, 100.0% of span | + | Thap9-AS1 |
| Platr6 | 45.2% of bases, 85.5% of span | + | LINC01010 |
| 6720427I07Rik | 94.3% of bases, 100.0% of span | + | LINC02603 |
| B830012L14Rik | 57.4% of bases, 83.8% of span | + | Meg8 (GM26945) |
| C330004P14Rik | - | + | LINC01625 |
| Gm38509 | 22.9% of bases, 84.4% of span | + | LINC01206 |
| A330094K24Rik | 54.7% of bases, 100.0% of span | + | C18orf25 (PCG) |
| Bvht | 53.2% of bases, 100.0% of span | + | Carmn |
| Dancr | 48.2% of bases, 49.0% of span | + | Dancr |
| 2900041M22Rik | 50.2% of bases, 60.5% of span | + | LINC01973 |
| Dleu2 | 72.8% of bases, 100.0% of span | + | Dleu2 |
| E130202H07Rik | 61.7% of bases, 65.2% of span | Tusc8 | |
| Epb41l4aos | 69.0% of bases, 100.0% of span | + | Epb41l4a-AS1 |
| Firre | 7.0% of bases, 14.5% of span | + | Firre |
| Gm20939 | - | + | LINC00470 |
| Gas5 | 71.3% of bases, 97.7% of span | + | Gas5 |
| Gm12688 | 92.6% of bases, 100.0% of span | + | FOXD3-AS1 |
| Gm47599 | 21.6% of bases, 85.0% of span | + | Socs2-AS1 |
| Gm19705 | 27.6% of bases, 47.8% of span | + | LINC00862 |
| Gm20703 | 79.2% of bases, 100.0% of span | + | GAPLINC |
| Gm26763 | 3.6% of bases, 3.8% of span | + | Smarca5-AS1 |
| Gm26945 | 65.4% of bases, 67.8% of span | + | Meg8 |
| AC129328.1 | - | + | LINC01340, |
| Gm28373 | 44.6% of bases, 83.5% of span | + | Itpk1-AS1 |
| Gm31693 | 12.7% of bases, 24.9% of span | + | LINC00578 |
| Mir124a-1hg | 91.7% of bases, 100.0% of span | + | LINC00599 |
| Mir142hg | 74.5% of bases, 100.0% of span | + | TSPOAP1-AS1 |
| Mir17hg | 74.7% of bases, 100.0% of span | + | Mir17Hg |
| Neat1 | 37.5% of bases, 100.0% of span | + | NEAT1 |
| Platr12 | 16.2% of bases, 33.7% of span | + | GPR1-AS |
| Rbakdn | 96.4% of bases, 99.1% of span | + | Rbakdn |
| Snhg1 | 73.3% of bases, 89.2% of span | + | Snhg1 |
| Snhg14 | 4.5% of bases, 5.4% of span | + | Snhg14 |
| D5Ertd605e | - | + | Pan3-AS1 |
| Snhg18 | 83.3% of bases, 100.0% of span | + | Snhg18 |
| Snhg5 | 67.8% of bases, 81.6% of span | + | Snhg5 |
| Sptbn5 | 78.8% of bases, 100.0% of span | + | Sptbn5 |
| Xist | 70.1% of bases, 100.0% of span | + | Xist |
Top protein candidates that interact with PHAROH.
| Protein hit | Ratio |
|---|---|
| Tial1 | 2.15559 |
| Hnrnpab | 1.80692 |
| Rbm3 | 1.77037 |
| Hnrnpd | 1.62883 |
| Hnrnpa1 | 1.6283 |
| Ptbp2 | 1.57804 |
| Hnrnpa3 | 1.53035 |
| Caprin1 | 1.50299 |
| Lmna | 1.37542 |
| Fubp3 | 1.34941 |
| Banf1 | 1.34137 |
| Hnrnpa2b1 | 1.33969 |
| H2afj | 1.3213 |
| Lima1 | 1.20909 |
| Nolc1 | 1.20733 |
| Abcb5 | 1.19592 |
| Nup62 | 1.18297 |
| Elavl1 | 1.09477 |
| Ssbp1 | 1.08439 |
| Hist1h2bc | 1.07366 |
| Itgax | 1.00222 |
| Rbm8a | 0.98396 |
| Dhx9 | 0.95827 |
| Smu1 | 0.94938 |
| Cnbp | 0.9225 |
| Nup93 | 0.82199 |
| Lsm3 | 0.79027 |
| Xrcc5 | 0.78242 |
| Med25 | 0.76892 |
| Actc1 | 0.76507 |
| Khsrp | 0.75921 |
| Actb | 0.75109 |
| Nipsnap1 | 0.75014 |
| Pnn | 0.74713 |
| Hba-a1 | 0.74299 |
| Snrpe | 0.74052 |
| Nol11 | 0.73772 |
| Erh | 0.73354 |
| Psmb1 | 0.72391 |
| Efhd2 | 0.71468 |
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Strain, strain background (C57BL/6J) (Mus musculus) Female | C57BL/6J | The Jackson Laboratory | Stock No: 000664 RRID:IMSR_JAX:000664 | |
| Gene (Homo sapiens) | Tial1 (NM_009383) Mouse Tagged ORF Clone | Origene | Cat# MG226372 | |
| Gene (Mus musculus) | Myc | GenBank | NC_000081.7 | |
| Recombinant Protein (Homo sapiens) | Recombinant Human TIAL1 Protein | Novus Biologicals | Cat# NBP2-51914-0.1mg | |
| Cell line (Mus musculus) | AB2.2 (ESCs) | Bergmann et al., 2015 | Cell line maintained in D. L. Spector Lab | |
| Cell line (Mus musculus) | NPC | Bergmann et al., 2015 | Cell line maintained in D. L. Spector Lab | |
| Cell line (Mus musculus) | Hepa1-6 | ATCC | Cat# CRL-1830 | Cell line maintained in D. L. Spector Lab |
| Cell line (Mus musculus) | Hepa1c1c7 | ATCC | Cat# CRL-2026 | Cell line maintained in D. L. Spector Lab |
| Cell line (Mus musculus) | AML12 | ATCC | Cat# CRL-2254 | Cell line maintained in D. L. Spector Lab |
| Cell line (Mus musculus) | MEF | MTI-Global Stem | Cat# GSC-6601G | Irradiated feeder MEFs |
| Cell line (Homo sapiens) | SNU-182 | ATCC | Cat# CRL-2235 | Cell line maintained in D. L. Spector Lab |
| Cell line (Homo sapiens) | Huh1 | N/A | Generous gift from Scott Lowe (MSKCC) | |
| Cell line (Homo sapiens) | Huh7 | N/A | Generous gift from Scott Lowe (MSKCC) | |
| Cell line (Homo sapiens) | JHH2 | N/A | RNA gifted from Scott Lowe (MSKCC) | |
| Cell line (Homo sapiens) | SNU-387 | ATCC | Cat# CRL-2237 | Generous gift from Scott Lowe (MSKCC) |
| Cell line (Homo sapiens) | Hep3B | ATCC | Cat# HB-8064 | Generous gift from Scott Lowe (MSKCC) |
| Cell line (Homo sapiens) | Alex | ATCC | Cat# CRL-8024 | RNA gifted from Scott Lowe (MSKCC) |
| Cell line (Homo sapiens) | HepG2 | ATCC | Cat# HB-8065 | Generous gift from Scott Lowe (MSKCC) |
| Cell line (Homo sapiens) | Li7 | N/A | RNA gifted from Scott Lowe (MSKCC) | |
| Cell line (Homo sapiens) | THLE-2 | ATCC | Cat# CRL-2706 | Cell line maintained in D. L. Spector Lab |
| Antibody | c-Myc (rabbit monoclonal) | Cell Signaling | Cat# 5605 RRID:AB_1903938 | (IB: 1:1000) |
| Antibody | TIAR (rabbit monoclonal) | Cell Signaling | Cat# 8509 RRID:AB_10839263 | (IB: 1:1000) (IF: 1:2000) (IP: 1:100) |
| Antibody | β-Actin (mouse monoclonal) | Cell Signaling | Cat# 3700 RRID:AB_2242334 | (IB: 1:10,000) |
| Antibody | IRDye 800CW (Goat anti-Rabbit IgG) | LI-COR Biosciences | Cat# 925-32211 RRID:AB_2651127 | (IB: 1:10,000) |
| Antibody | IRDye 680RD (Goat anti-Mouse IgG) | LI-COR Biosciences | Cat# 925-68070 RRID:AB_2651128 | (IB: 1:10,000) |
| Antibody | Goat anti-Rabbit IgG (H + L) Cross-Adsorbed Secondary Antibody Alexa Fluor 488 | Thermo Fisher | Cat# A-11008 RRID:AB_143165 | (IF: 1:1000) |
| Antibody | Rabbit IgG Isotype Control | Thermo Fisher | Cat# 10500C RRID:AB_2532981 | |
| Recombinant DNA reagent | eSpCas9-1.1 | Addgene | RRID:Addgene_71814 | Backbone for constructing GFP and mCherry variants |
| Recombinant DNA reagent | eSpCas9-1.1-GFP (plasmid) | This study | N/A | |
| Recombinant DNA reagent | eSpCas9-1.1-mCherry (plasmid) | This study | N/A | |
| Recombinant DNA reagent | pmirGLO | Promega | Cat# E1330 | Dual-Luciferase miRNA Target Expression Vector |
| Recombinant DNA reagent | pCMV6-A-Puro | Origene | Cat# PS100025 | pCMV6 backbone |
| Transfected construct (Mus musculus) | sgPHAROH_F- eSpCas9-1.1-GFP (plasmid) | This study | N/A | Upstream PHAROH sgRNA |
| Transfected construct (Mus musculus) | sgPHAROH_ R-eSpCas9- 1.1-mCherry (plasmid) | This study | N/A | Downstream PHAROH sgRNA |
| Transfected construct (Mus musculus) | sgRenilla- eSpCas9-1.1-GFP (plasmid) | Chang et al., 2020 | N/A | Negative control sgRNA |
| Transfected construct (Mus musculus) | pmirGLO-MYC (plasmid) | This study | N/A | Construct for luciferase assay readout |
| Transfected construct (Mus musculus) | pCMV6-pharoh (plasmid) | This study | N/A | Construct for rescue and luciferase assay readout |
| Transfected construct (Mus musculus) | pCMV6-m4pharoh (plasmid) | This study | N/A | Construct for luciferase assay readout |
| Transfected construct (Mus musculus) | pCMV6-GFP (plasmid) | Chang et al., 2020 | N/A | Construct for luciferase assay readout |
| Sequence-based reagent | ASO 7 | This study | PHAROH Gapmer ASO | CGTGTCATCTTCTTGGCCCC |
| Sequence-based reagent | ASO 15 | This study | PHAROH Gapmer ASO | TCGTGTCATCTTCTTGGCCC |
| Sequence-based reagent | ASO 14 | This study | PHAROH cEt ASO | GTTACAGGACGCATGT |
| Sequence-based reagent | ASO 18 | This study | PHAROH cEt ASO | CACATAGTTATTCCCG |
| Sequence-based reagent | Forward | This study | PHAROH genomic PCR | TGCTTAGCACGT CCTCAGTGC |
| Sequence-based reagent | Reverse | This study | PHAROH genomic PCR | AGTTCCCCAGC AACCCTGTT |
| Sequence-based reagent | Upstream | This study | PHAROH sgRNA | GCAGGTAGTGT GGTAACTCC |
| Sequence-based reagent | Downstream | This study | PHAROH sgRNA | CGGGTCCTCCC AGCGCACAC |
| Sequence-based reagent | Exon 4 Fwd | This study | PHAROH qRT-PCR | GGGGCCAAGAA GATGACACG |
| Sequence-based reagent | Exon 4 Ref | This study | PHAROH qRT-PCR | GGACGCATGT GGAGGTCAGA |
| Sequence-based reagent | Exon A Fwd | This study | PHAROH qRT-PCR | TGCCTCACAA GGGACAACACTC |
| Sequence-based reagent | Exon A Rev | This study | PHAROH qRT-PCR | GAATTTGCTCA GGGGCTCCA |
| Sequence-based reagent | Exon B Fwd | This study | PHAROH qRT-PCR | GGACTTGAACT GGCACTGTTGC |
| Sequence-based reagent | Exon B Rev | This study | PHAROH qRT-PCR | CAGAAGGACC ATCATCACGA |
| Sequence-based reagent | Exon C Fwd | This study | PHAROH qRT-PCR | TGAACCCGAGC TTTGCCATT |
| Sequence-based reagent | Exon C Rev | This study | PHAROH qRT-PCR | CGGTGCTCTG CAGGACGTTT |
| Sequence-based reagent | Exon D Fwd | This study | PHAROH qRT-PCR | AGGCTGCCGC CACACTTAAA |
| Sequence-based reagent | Exon D Rev | This study | PHAROH qRT-PCR | TTCAGCTGCTGG CATTCTTCC |
| Sequence-based reagent | Exon E Fwd | This study | PHAROH qRT-PCR | GGAGAGAACAA GGGCCTTCC |
| Sequence-based reagent | Exon E Rev | This study | PHAROH qRT-PCR | GCCCTGCTGCA TTCTGGGTA |
| Sequence-based reagent | Exon 1 Fwd | This study | PHAROH qRT-PCR | GGTGTGAACCAA GTGCACGTCT |
| Sequence-based reagent | Exon 1 Rev | This study | PHAROH qRT-PCR | GGGATCTGACA CCGCCTTCTT |
| Sequence-based reagent | Exon 2 Fwd | This study | PHAROH qRT-PCR | CTTCTGAGTCTG ACGGGCTGGT |
| Sequence-based reagent | Exon 2 Rev | This study | PHAROH qRT-PCR | TCAGTCCTACCC AAGAAATTTAGGA |
| Sequence-based reagent | Exon 3 Fwd | This study | PHAROH qRT-PCR | TGTGGAAACTCA GAGAGGATGC |
| Sequence-based reagent | Exon 3 Rev | This study | PHAROH qRT-PCR | CTCTGGTGGCTG TGCCTTCAAA |
| Sequence-based reagent | MycF | This study | Myc qRT-PCR | CAACGTCTTGG AACGTCAGA |
| Sequence-based reagent | MycR | This study | Myc qRT-PCR | TCGTCTGCTT GAATGGACAG |
| Sequence-based reagent | Outer 1 | This study | 5' RACE | TTCCTGCGTG AAAGTGTCTG |
| Sequence-based reagent | Outer 2 | This study | 5' RACE | TGACCTTCTCA GGAAGTGGAA |
| Sequence-based reagent | Inner 1 | This study | 5' RACE | CCTGAGAGGAC GAGGTGACT |
| Sequence-based reagent | Inner 2 | This study | 5' RACE | TTTGCAGGTTA GGATCAGAGC |
| Sequence-based reagent | Outer | This study | 3' RACE | CACTTCCATT CCTCCCCATA |
| Sequence-based reagent | Inner | This study | 3' RACE | GGGGACTCAGA CACTCACCA |
| Sequence-based reagent | PHAROH hairpin | This study | T7 Transcription Primer | TAATACGAC TCACTATA gagaggatgccactgttttg aactattttgaaggcacag ccaccagagctttaggg acagggtattttatc |
| Sequence-based reagent | Myc 3' UTR | This study | T7 Transcription Primer | TAATACGACTCACTATAG cttcccatcttttttctttttcc ttttaacagatttg tatttaattgttttt |
| Sequence-based reagent | m1 | This study | T7 Transcription Primer | TAATACGACTCACTATA gagaggatgccactgtCt Cgaactattttgaaggca cagccaccagagcttta gggacagggtattttatc |
| Sequence-based reagent | m2 | This study | T7 Transcription Primer | TAATACGACTCACTATA gagaggatgccactgtCtC gaactaCtCtgaaggcac agccaccagagctttaggg acagggtattttatc |
| Sequence-based reagent | m3 | This study | T7 Transcription Primer | TAATACGACTCACTATA gagaggatgccactgtCtC gaactaCtCtgaaggcac agccaccagagcCtta gggacagggtattttatc |
| Sequence-based reagent | m4 | This study | T7 Transcription Primer | TAATACGACTCACTATA gagaggatgccactgtCtC gaactaCtCtgaaggcaca gccaccagagcCttaggg acagggtatCCtatc |
| Sequence-based reagent | PHAROH 1 | This study | Biotin antisense pulldown oligo | AGAAATTTAGGAG CCACGCT |
| Sequence-based reagent | PHAROH 2 | This study | Biotin antisense pulldown oligo | GCTGTGCCTTC AAAATAGTT |
| Sequence-based reagent | PHAROH 3 | This study | Biotin antisense pulldown oligo | GCCCCAAGAAA CTCAAGAAT |
| Sequence-based reagent | PHAROH 4 | This study | Biotin antisense pulldown oligo | TTAATTTTCT CCTTTATGCA |
| Sequence-based reagent | PHAROH 5 | This study | Biotin antisense pulldown oligo | ACAACGTGTGG ATGTGTGTT |
| Sequence-based reagent | PPIB 1 | This study | Biotin antisense pulldown oligo | CCTACAGATT CATCTCCAAT |
| Sequence-based reagent | PPIB 2 | This study | Biotin antisense pulldown oligo | GTTATGAAGAA CTGTGAGCC |
| Commercial assay or kit | DNase I, Amplification Grade | Life Technologies | Cat# 18068 | |
| Commercial assay or kit | TaqMan Reverse Transcription Reagents | Thermo Fisher | Cat# 4304134 | |
| Commercial assay or kit | SF Cell Line 4D-Nucleofector X Kit L | Lonza | Cat# V4XC-2024 | |
| Commercial assay or kit | View ISH Cell Assay Kit | Affymetrix | Cat# QVC0001 | |
| Commercial assay or kit | MEGAscript T7 Transcription Kit | Thermo Fisher | AM1333 | |
| Commercial assay or kit | Pierce RNA 3' End Biotinylation Kit | Thermo Fisher | Cat# 20160 | |
| Commercial assay or kit | LightShift Chemiluminescent RNA EMSA Kit | Thermo Fisher | Cat# 20158 | |
| Commercial assay or kit | Pierce BCA Protein Assay Kit | Life Technologies | Cat# 23227 | |
| Commercial assay or kit | CellTiter 96 AQueous One Solution Cell Proliferation Assay | Promega | Cat# G3582 | |
| Commercial assay or kit | SMARTer RACE 5′/3′ Kit | Takara | Cat# 634858 | |
| Commercial assay or kit | Promega Dual-Luciferase Reporter Assay System | Promega | Cat# E1960 | |
| Commercial assay or kit | DNeasy Blood and Tissue kit | Qiagen | Cat# 69504 | |
| Software, algorithm | Benchling | https://www.benchling.com/ | Used for sgRNA design and cloning | |
| Software, algorithm | CPAT | doi: 10.1093/nar/gkt006 | ||
| Software, algorithm | CPC | doi: 10.1093/nar/gkm391 | ||
| Software, algorithm | PhyloCSF | doi: 10.1093/bioinformatics/btr209 | ||
| Software, algorithm | FastQC | https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ | RRID:SCR_014583 | |
| Software, algorithm | STAR | doi: 10.1002/0471250953.bi1114s51 | RRID:SCR_004463 | |
| Software, algorithm | DESeq2 | doi: 10.1186/s13059-014-0550-8 | RRID:SCR_015687 | |
| Software, algorithm | Volocity 3D Image Analysis Software | Perkin Elmer | RRID:SCR_002668 | |
| Software, algorithm | SoftWoRx | SoftWoRx Software | RRID:SCR_019157 | |
| Software, algorithm | Sequest HT | doi: 10.1016/1044-0305(94)80016-2 | ||
| Software, algorithm | Mascot 2.5 | doi: 10.1002/(SICI)1522-2683(19991201)20:18<3551::AID-ELPS3551>3.0.CO;2–2 | RRID:SCR_014322 | |
| Software, algorithm | HOMER Suite | doi: 10.1016/j.molcel.2010.05.004 | RRID:SCR_010881 | |
| Software, algorithm | Image Studio Software | LI-COR | RRID:SCR_015795 | |
| Software, algorithm | RNAfold | doi: 10.1093/nar/gkg599 | RRID:SCR_008550 | |
| Software, algorithm | mFold | doi: 10.1093/nar/gkg595 | RRID:SCR_008543 | |
| Software, algorithm | ImageJ | NIH, Bethesda, MD | RRID:SCR_003070 | |
| Chemical compound, drug | Hoechst dye | Thermo Fisher | Cat# 62249 | 1 μg/ml |
| Chemical compound, drug | DAPI | Life Technologies | Cat# D1306 | 1 μg/ml |
| Chemical compound, drug | α-Amanitin | Sigma-Aldrich | Cat# A2263 | 5 μg/ml |
| Chemical compound, drug | Diethylnitrosamine | Sigma-Aldrich | Cat# 73861 | 25 mg/kg |





