microRNA-mediated regulation of microRNA machinery controls cell fate decisions
Abstract
microRNAs associate with Argonaute proteins, forming the microRNA-induced silencing complex (miRISC), to repress target gene expression post-transcriptionally. Although microRNAs are critical regulators in mammalian cell differentiation, our understanding of how microRNA machinery, such as the miRISC, are regulated during development is still limited. We previously showed that repressing the production of one Argonaute protein, Ago2, by Trim71 is important for mouse embryonic stem cells (mESCs) self-renewal (Liu et al., 2021). Here, we show that among the four Argonaute proteins in mammals, Ago2 is the major developmentally regulated Argonaute protein in mESCs. Moreover, in pluripotency, besides the Trim71-mediated regulation of Ago2 (Liu et al., 2021), Mir182/Mir183 also repress Ago2. Specific inhibition of this microRNA-mediated repression results in stemness defects and accelerated differentiation through the let-7 microRNA pathway. These results reveal a microRNA-mediated regulatory circuit on microRNA machinery that is critical to maintaining pluripotency.
Introduction
microRNAs (miRNAs) are endogenous ~22 nucleotide (nt) RNAs with critical roles in modulating gene expression under diverse biological contexts (Bartel, 2009; Bartel, 2018). Most miRNAs are produced from long primary transcripts (pri-miRNAs) through successive processing by two double-stranded RNA (dsRNA) endonucleases named Drosha and Dicer, generating pre-miRNAs and ~22 nt dsRNAs, respectively. One RNA strand in the ~22 nt dsRNA, the mature miRNA, is selectively incorporated into the Argonaute (Ago) protein, forming the miRNA-induced silencing complex (miRISC) (Ha and Kim, 2014). In animals, miRISC recognizes its target mRNAs through partial base pairings mediated by the miRNA (Bartel, 2009). The Ago protein recruits GW182 proteins to down-regulate target mRNA expression through mRNA degradation and/or translational repression (Nilsen, 2007). Although miRNAs play critical regulatory roles in mammalian cell differentiation (Ameres and Zamore, 2013; Ebert and Sharp, 2012), our understanding on how miRNA machinery, particularly the miRISC, are regulated during development is still limited.
We recently found that Ago2, a key component in the miRISC, is repressed at the mRNA translation level by an RNA-binding protein named Trim71 in mouse embryonic stem cells (mESCs) (Liu et al., 2021). This repression of Ago2 inhibits stem cell differentiation mediated by the conserved pro-differentiation let-7 miRNAs (Büssing et al., 2008; Liu et al., 2021). These results suggest that Ago2 is developmentally regulated during stem cell self-renewal and differentiation, and beg for characterization of additional regulators of Ago2. Moreover, besides Ago2, there are three additional Ago proteins (Ago1, Ago3, Ago4) in mammals that function redundantly in the miRNA pathway (Meister, 2013). The relative abundance of these Ago proteins and their contribution to miRNA activities during cell differentiation, however, are still unknown.
Here, using mESC fate decisions between pluripotency and differentiation as a mammalian cell differentiation model, we determined that Ago2 is the predominant Ago protein in mESCs, and Ago2 level increases when mESCs exit pluripotency. In the pluripotent state, Mir182 and Mir183, two conserved miRNAs abundantly expressed in mESCs, repress Ago2 and control the stemness of mESCs. Specific inhibition of Mir182/Mir183-mediated repression of Ago2 results in stemness defects and accelerated differentiation of mESCs through the let-7 miRNA pathway. Collectively, these results reveal an miRNA-mediated regulatory circuit on the miRNA machinery that is critical to maintaining pluripotency.
Results
Ago2 is the predominant developmentally regulated Ago protein in mESCs
Mammals have four Ago proteins (Ago1–4) that function redundantly in miRNA-mediated regulations (Meister, 2013). Transcriptomic profiling on mESCs from different laboratories indicated that mESCs express only Ago1 and Ago2 (Figure 1—figure supplement 1A; Liu et al., 2021; Marks et al., 2012). To examine the relative abundance of Ago1 and Ago2 at the protein level, we generated mESCs with a Flag-tag knocked-in at the N-terminus of the Ago1 and Ago2 loci, respectively, via CRISPR/Cas9-mediated genome editing (Figure 1—figure supplement 1B, C). These mESCs with the Flag-tag knocked-in displayed no stemness defects compared to the wild-type (WT) mESCs (Figure 1—figure supplement 1D) and enabled us to use the same antibody (e.g., anti-Flag) to compare the relative abundance of Ago1 and Ago2. Western blotting via an anti-Flag antibody indicated that Ago2 is the predominant Ago protein in mESCs at the protein level (Figure 1A).
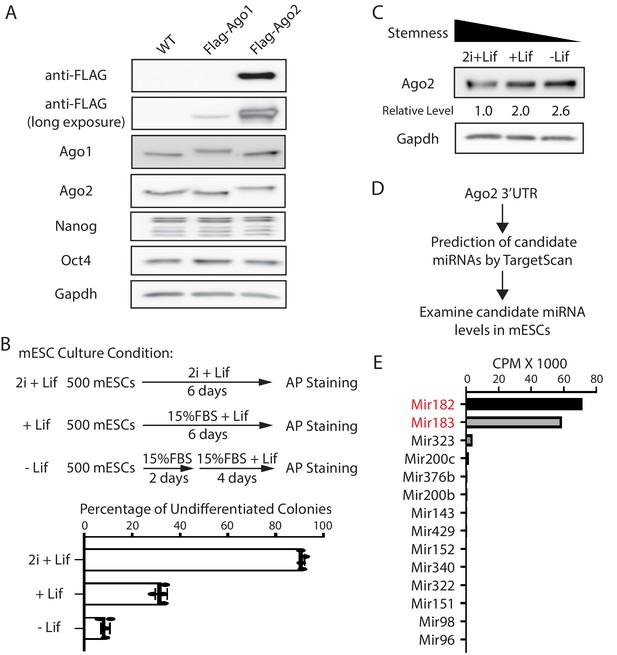
Ago2 is the major developmentally regulated Argonaute protein in mouse embryonic stem cells (mESCs).
(A) Western blotting in the wild-type (WT), Flag-Ago1, and Flag-Ago2 mESCs. (B) Colony formation assay for the mESCs. The WT mESCs were cultured under the indicated conditions, and the resultant colonies were fixed and stained for AP (alkaline phosphatase activity). The results represent the means (± SD) of four independent experiments. (C) Western blotting in the WT mESCs cultured under the indicated conditions. (D) Outline of identifying miRNAs that can potentially regulate Ago2. (E) Expression levels of the identified miRNAs from (D) in mESCs. CPM: counts per million reads.
-
Figure 1—source data 1
Tiff files of raw gel images for Figure 1A and C; Figure 1—figure supplement 1C.
Excel files of numbers for Figure 1B and E; Figure 1—figure supplement 1A, D, Figure 1—figure supplement 2.
- https://cdn.elifesciences.org/articles/72289/elife-72289-fig1-data1-v2.zip
To examine whether Ago2 level is regulated during mESCs differentiation, we cultured mESCs under three different conditions that mimic three different developmental stages: ground/naive state (in 2i + Lif), primed state (in 15% FBS+ Lif), and differentiating state (in 15% FBS without Lif), which resulted in decreasing stemness in mESCs, as determined by the colony formation assay (Figure 1B). Western blotting indicated that Ago2 level increased when mESCs exited pluripotency (Figure 1C). This result indicated that Ago2 is developmentally regulated in mESCs, and Ago2 level is repressed in the pluripotent state.
Mir182/Mir183 regulate Ago2 and maintain stemness in mESCs
To determine how Ago2 is regulated in mESCs, we hypothesized that miRNAs expressed in mESCs might contribute to the repression of Ago2 because miRNAs are important negative regulators of gene expression. We identified the conserved miRNA-binding sites in the 3’UTR of Ago2 mRNA through TargetScan (Agarwal et al., 2015) and then examined the expression level of the corresponding miRNAs in mESCs using existing small-RNA-seq datasets (Liu et al., 2021 Figure 1D). This analysis revealed that among the miRNAs that can potentially regulate Ago2, Mir182, and Mir183, two miRNAs from the same miRNA family that are abundantly expressed in stem cells Dambal et al., 2015, have significantly higher expression levels (Figure 1E). Interestingly, Mir182/Mir183 decrease when mESCs transition from the ground state to the primed and differentiating state (Hadjimichael et al., 2016; Wang et al., 2017), which negatively correlates with the Ago2 expression pattern during this transition (Figure 1C). These observations suggest that Ago2 is repressed by Mir182/Mir183 in mESCs. Consistent with this notion, using RNA antisense purification, we found that Mir182 and Mir183 specifically associated with Ago2 mRNA in mESCs (Figure 1—figure supplement 2).
Two lines of evidence indicated that Mir182/Mir183 regulate Ago2 mRNA. First, Ago2 increased when Mir182, Mir183, or both Mir182 and Mir183 were knocked out in mESCs (Figure 2—figure supplement 1A, Figure 2A and B). Second, when either Mir182 or Mir183 was over-expressed in the WT mESCs (Figure 2—figure supplement 1B), the Ago2 level decreased (Figure 2—figure supplement 1C). The results from these loss-of-function and gain-of-function experiments argue that Mir182/Mir183 repress Ago2 expression in mESCs.
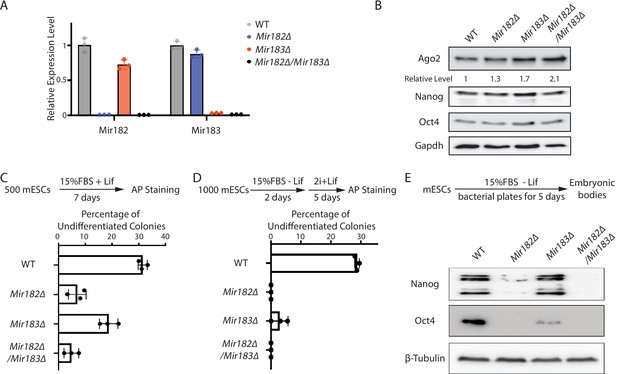
Mir182/Mir183 regulate Ago2 and maintain stemness in mouse embryonic stem cells (mESCs).
(A) qRT-PCR on Mir182 and Mir183. For each miRNA, the expression level in wild-type (WT) cells was set as one for relative comparison. U6 RNA was used for normalization. The results represent the means (± SD) of three independent replicates. (B) Western blotting in the WT, Mir182Δ, Mir183Δ, and Mir182Δ/Mir183Δ mESCs. GAPDH was used for normalization in calculating the relative expression levels. (C) Colony formation assay for mESCs. The mESCs were cultured in 15% FBS+ Lif for 7 days, and the resultant colonies were fixed and stained for alkaline phosphatase (AP). (D) Exit pluripotency assay for mESCs. The mESCs were induced to exit pluripotency in medium without Lif for 2 days and then switched to 2i + Lif medium for 5 days. The resultant colonies were fixed and stained for AP. In (C and D), the colony morphology and AP intensity were evaluated through microscopy; 100–200 colonies were examined each time to determine the percentage of undifferentiated colonies. The results represent the means (± SD) of three independent experiments. (E) Western blotting of pluripotency factors during embryoid body (EB) formation.
-
Figure 2—source data 1
Tiff files of raw gel images for Figure 2B and E; Figure 2—figure supplement 1C.
Excel files of numbers for Figure 2A, C and D; Figure 2—figure supplement 1B.
- https://cdn.elifesciences.org/articles/72289/elife-72289-fig2-data1-v2.zip
Interestingly, Mir182Δ, Mir183Δ, and Mir182Δ/Mir183Δ mESCs displayed defects in self-renewal (Figure 2C), as determined by the colony formation assay in the 15% FBS + Lif medium, where differentiation was not blocked by the two inhibitors in the 2i + Lif medium. Moreover, these miRNA knockout mESCs had accelerated differentiation, as revealed by the exit pluripotency assay (Figure 2D), which evaluates the rate ESCs exit the pluripotent state (Betschinger et al., 2013), and by the measurement of pluripotency factors through Western blotting on differentiating embryonic bodies (Figure 2E). These cellular phenotypes suggest that Mir182/Mir183-mediated regulation of Ago2 is important to mESCs.
Mir182/Mir183-mediated repression of Ago2 is required for maintaining pluripotency
A caveat in interpreting results from miRNA knockout and over-expression experiments is the pleiotropic effects. Because each miRNA can regulate hundreds of mRNAs, when an miRNA is knocked out or over-expressed, hundreds of miRNA:mRNA interactions are altered, making it difficult to determine whether a specific miRNA:mRNA interaction contributes to the phenotypical changes.
To address this issue and specifically examine the functional significance of Mir182/Mir183-mediated regulation of Ago2 in mESCs, we mutated the Mir182/Mir183-binding sites in the 3’UTR of Ago2 mRNA via CRISPR/Cas9-mediated genome editing (Figure 3A and B). Two observations indicated that the mutations disrupted the interaction between Ago2 mRNA and Mir182/Mir183. First, similar to the miRNA knockout mESCs (Figure 2B), Ago2 increased in the 3’UTR mutant mESCs (Figure 3C). Second, in contrast to the results in the WT mESCs (Figure 2—figure supplement 1C), over-expression of either Mir182 or Mir183 in the 3’UTR mutant mESCs did not decrease Ago2 (Figure 3—figure supplement 1A, B). Notably, in the Mir182Δ/Mir183Δ mESCs, these mutations did not increase Ago2 (Figure 3C), indicating the increased Ago2 from these mutations in the WT mESCs is dependent on Mir182/Mir183. Moreover, the 3’UTR mutations did not significantly alter the Mir182/Mir183 levels in mESCs (Figure 3—figure supplement 1C). Altogether, these observations indicated that the functional significance of Mir182/Mir183-mediated repression of Ago2 could be specifically evaluated in the 3’UTR mutant mESCs.
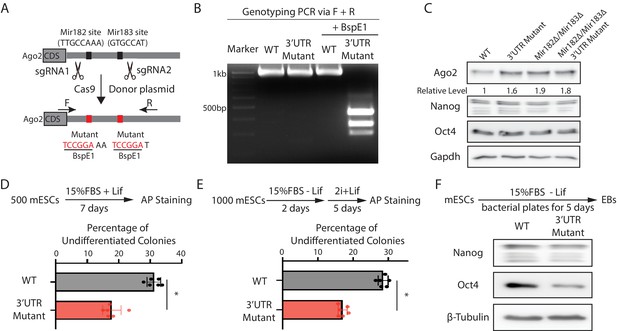
Mir182/Mir183-mediated repression of Ago2 is required for maintaining pluripotency.
(A) Mutating Mir182- and Mir183-binding sites in Ago2 mRNA’s 3’UTR via genome editing. (B) Genotyping of the Ago2 3’UTR mutant. The PCR was performed using the oligos (F and R) indicated in (A). (C) Western blotting in the wild-type (WT), Ago2 3’UTR mutant, Mir182Δ/ Mir183Δ, and Mir182Δ/ Mir183Δ/Ago2 3’UTR mutant. (D) Colony formation assay for mouse embryonic stem cells (mESCs). (E) Exit pluripotency assay for mESCs. In (D and E), the colony morphology and alkaline phosphatase (AP) intensity were evaluated through microscopy. The results represent the means (± SD) of four independent experiments. *p < 0.05 by the Student’s t-test. Western blotting of pluripotency factors in day 5 embryoid bodies (EBs).
-
Figure 3—source data 1
Tiff files of raw gel images for Figure 3C and F; Figure 3—figure supplement 1B.
Excel files of numbers for Figure 3D and E; Figure 3—figure supplement 1A, C.
- https://cdn.elifesciences.org/articles/72289/elife-72289-fig3-data1-v2.zip
When subject to the colony formation assay, the 3’UTR mutant mESCs displayed a defect in maintaining undifferentiated colonies (Figure 3D), indicating compromised self-renewal. When differentiation was evaluated by the exit pluripotency assay, the 3’UTR mutant mESCs had an increased differentiation rate (Figure 3E). Consistent with these findings, differentiating embryonic bodies from the 3’UTR mutant mESCs had a lower amount of pluripotency factors (Figure 3F). Collectively, these results indicate that Mir182/Mir183-mediated repression of Ago2 is important for mESC self-renewal and proper differentiation.
Mir182/Mir183-mediated repression of Ago2 in mESCs inhibits the let-7 miRNA-mediated differentiation pathway
Two observations lead us to the hypothesis that Mir182/Mir183-mediated repression of Ago2 in mESCs counteracts the differentiation pathway controlled by the let-7 miRNAs, a conserved miRNA family that promotes stem cell differentiation (Roush and Slack, 2008). First, in Dgcr8Δ mESCs, where endogenous miRNAs’ biogenesis is blocked, ectopic expression of Mir183 inhibits the stem cell differentiation triggered by exogenous let-7 miRNA (Wang et al., 2017). Second, our recent study indicated that increasing Ago2 levels in mESCs results in stemness defects in a let-7-miRNA-dependent manner. This specificity on let-7 miRNAs is because the pro-differentiation let-7 miRNAs are actively transcribed in mESCs, and the increased Ago2 binds and stabilizes the let-7 miRNAs that are otherwise degraded in mESCs, thereby promoting mESCs differentiation (Liu et al., 2021).
To test this hypothesis, we examined the expression of let-7 miRNAs. The 3’UTR mutant mESCs had significantly higher let-7 miRNAs than the WT mESCs (Figure 4A). This increase is specific to let-7 miRNAs because non-let-7 miRNAs were not elevated (Figure 4A). Moreover, consistent with our previous observation that increased Ago2 stabilizes mature let-7 miRNAs (Liu et al., 2021), the pri-let-7 miRNAs and the pre-let-7 miRNAs were not significantly increased in the 3’UTR mutant mESCs (Figure 4A). To determine whether the increased let-7 miRNAs are responsible for the stemness defects in the 3’UTR mutant mESCs, we inhibited let-7 miRNAs using locked nucleic acid (LNA) antisense oligonucleotides targeting the conserved seed sequence of let-7 miRNAs. When let-7 miRNAs were inhibited, the stemness defects of the 3’UTR mutant mESCs were abolished (Figure 4B), indicating that disruption of Mir182/Mir183-mediated repression of Ago2 in mESCs activates differentiation through the let-7 miRNA pathway.
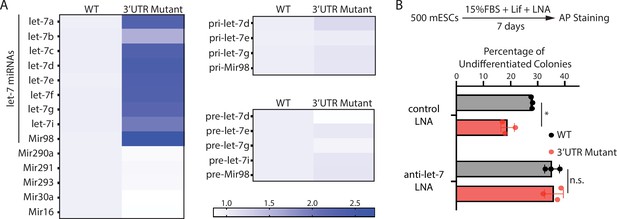
The stemness defects in the 3’UTR mutant mouse embryonic stem cells (mESCs) are caused by elevated let-7 microRNAs (miRNAs).
(A) Relative levels of miRNAs, let-7 pri-miRNAs, and let-7 pre-miRNAs in the wild-type (WT) and the Ago2 3’UTR mutant mESCs. For each miRNA, pri-miRNA, and pre-miRNA, the expression level in WT cells was set as one for relative comparison. U6 RNA was used for normalization in miRNA and pre-miRNA quantification; 18 S rRNA was used for normalization in pri-miRNA quantification. The heatmap was generated from the means of three independent replicates. (B) Colony formation assay for WT and the Ago2 3’UTR mutant mESCs cultured in the presence of 500 nM anti-let-7 locked nucleic acid (LNA) or a control LNA. The results represent three independent experiments. *p < 0.05, and n.s. not significant (p > 0.05) by the Student’s t-test.
-
Figure 4—source data 1
Excel files of numbers for Figure 4A and B.
- https://cdn.elifesciences.org/articles/72289/elife-72289-fig4-data1-v2.zip
Mir182/Mir183 and trim71 function in parallel to repress Ago2 mRNA in mESCs
Our previous study indicated that Ago2 mRNA is also repressed by Trim71 in mESCs (Liu et al., 2021). Interestingly, the Trim71-binding site in the 3’UTR of Ago2 mRNA is different from the Mir182/Mir183-binding sites, suggesting that Mir182/Mir183 and Trim71 function in parallel to repress Ago2 mRNA in mESCs. We performed the following experiments to test this.
At the molecular level, we observed that over-expression of Trim71 still repressed Ago2 in the 3’UTR mutant mESCs (Figure 5A), where Mir182/Mir183-mediated repression is abolished (Figure 3). Moreover, in the 3’UTR mutant mESCs, inhibiting Trim71-mediated repression of Ago2 through deleting the Trim71-binding site in the 3’UTR of Ago2 mRNA (CLIPΔ) (Liu et al., 2021) further increased Ago2 level (Figure 5B, Figure 5—figure supplement 1). These results indicate that Trim71 and Mir182/Mir183 independently repress Ago2 mRNA in mESCs.
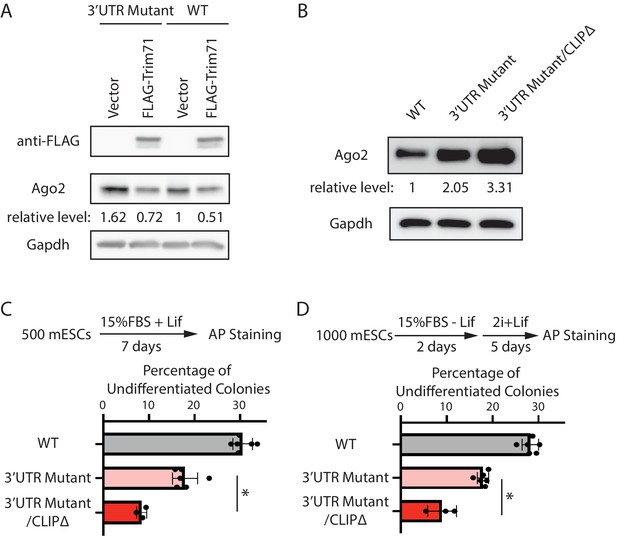
Mir182/Mir183 and Trim71 function in parallel to repress Ago2 mRNA in mouse embryonic stem cells (mESCs).
(A) Western blotting in the wild-type (WT) mESCs expressing either a vector or FLAG-Trim71 and in the 3’UTR mutant mESCs expressing either a vector or FLAG-Trim71. (B) Western blotting in the WT, 3’UTR mutant, and 3’UTR mutant/CLIPΔ mESCs. In (A and B), GAPDH was used for normalization in calculating the relative expression levels. (C) Colony formation assay for mESCs. (D) Exit pluripotency assay for mESCs.
-
Figure 5—source data 1
Tiff files of raw gel images for Figure 5A and B; Figure 5—figure supplement 1.
An Excel file of numbers for Figure 5C and D.
- https://cdn.elifesciences.org/articles/72289/elife-72289-fig5-data1-v2.zip
At the cell function level, we found that introducing the CLIPΔ in the 3’UTR mutant mESCs further decreased stem cell self-renewal, as determined by the colony formation assay (Figure 5C), and accelerated differentiation, as measured by the exit pluripotency assay (Figure 5D). These observations argue that Trim71 and Mir182/Mir183 function independently in regulating stemness in mESCs through modulating Ago2 mRNA.
Collectively, these findings indicate that Mir182/Mir183 and Trim71 function in parallel to repress Ago2 mRNA in mESCs.
Discussion
Our data reveal that the predominant Ago protein in mESCs, Ago2, is developmentally regulated, with gradually increasing levels when mESCs exit pluripotency. Two miRNAs abundantly expressed in mESCs, Mir182/Mir183, contribute to the repression of Ago2 in the pluripotent state. This miRNA-mediated regulation of Ago2 is critical to maintaining stemness. Our findings raise several interesting aspects of miRNAs in stem cell biology.
First, since Ago2 is the predominant Ago protein in mESCs, the Ago2 expression pattern during mESCs’ transition from self-renewal to differentiation argues that although certain individual miRNAs may be required for pluripotency (e.g., Mir182/Mir183), the global miRNA activity is suppressed in the pluripotent state and induced when mESCs initiate differentiation. Consistent with this notion, knocking out key components in global miRNA biogenesis, such as Dgcr8 (Wang et al., 2007), Dicer (Kanellopoulou et al., 2005; Murchison et al., 2005), or Ago2 in the miRISC (Liu et al., 2021), does not negatively affect mESCs self-renewal. However, differentiation in all these mutant mESCs is severely compromised. Thus, at the global level, miRNAs may play more important roles in mESC differentiation.
Second, previous studies indicate that the two components of the miRISC, the Ago protein and its associated miRNA, mutually regulate each other. In the absence of miRNAs, the Ago protein is destabilized (Martinez and Gregory, 2013; Smibert et al., 2013), while miRNAs are also unstable if they are not associated with Ago proteins (Winter and Diederichs, 2011). Thus, the effective miRNA activity depends on the limiting component in the miRISC. Our previous studies indicated that the conserved pro-differentiation let-7 miRNAs are sensitive to Ago2 levels because an increase of Ago2 results in specific stabilization of let-7 miRNAs that are otherwise degraded (Liu et al., 2021). Thus, for let-7 miRISC, Ago2 is possibly the limiting component in mESCs. Repression of Ago2 by either Mir182/Mir183 as we characterized here or Trim71 as we identified previously (Liu et al., 2021) likely limits the effective let-7 miRISCs. Interestingly, the pro-differentiation let-7 miRICSs can positively auto-regulate their own biogenesis through inhibiting Lin28a, a conserved let-7 target, because Lin28a inhibits the biogenesis of let-7 miRNAs through promoting their pre-miRNA degradation (Tsialikas and Romer-Seibert, 2015). Thus, the effective let-7 miRNAs need to be tightly controlled in stem cells. The two repression mechanisms on Ago2 mRNA contribute to limiting the amount of effective let-7 miRISCs and maintaining pluripotency in mESCs. We speculate that similar mechanisms of regulating miRISCs by RNA-binding proteins and miRNAs may exist in other developmental processes. Moreover, Ago2 is dysregulated under many pathological conditions, such as cancer (Adams et al., 2014). Thus, regulating miRISCs through modulating Ago2 levels may also contribute to pathogenesis.
Finally, it is noticed that the Mir182Δ/Mir183Δ mESCs displayed stronger defects in self-renewal and differentiation than the 3’UTR mutant mESCs did (Figure 2C and D versus Figure 3D and E). Thus, besides Ago2 mRNA, Mir182/Mir183 may regulate additional mRNAs that are important for stem cell biology.
Materials and methods
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Antibody | (Mouse monoclonal) anti-FLAG M2 | Sigma-Aldrich | Cat# F1804 | WB (1:5000) |
| Antibody | (Mouse monoclonal) anti-GAPDH (6 C5) | Santa Cruz Biotechnology | Cat# sc-32233 | WB (1:5000) |
| Antibody | (Rabbit monoclonal) anti-beta-Tubulin | Selleckchem | Cat# A5032 | WB (1:5000) |
| Antibody | (Rabbit monoclonal) anti-Ago1 (D84G10) | Cell Signaling Technology | Cat# 5053 | WB (1:1000) |
| Antibody | (Rabbit monoclonal) anti-Ago2 | Bimake | Cat# A5701 | WB (1:3000) |
| Antibody | (Mouse monoclonal) anti-Oct-4 | BD Transduction Laboratories | Cat# 611202 | WB (1:5000) |
| Antibody | (Rabbit monoclonal) anti-Nanog (D2A3) | Cell Signaling Technology | Cat# 8822 | WB (1:3000) |
| Antibody | Goat Anti-Rabbit IgG (H L)-HRP Conjugate | Bio-Rad | Cat# 170-6515 | WB (1:5000) |
| Antibody | Goat Anti-Mouse IgG (H L)-HRP Conjugate | Bio-Rad | Cat# 170-6516 | WB (1:5000) |
| Chemical compound, drug | DMEM/F-12 | Gibco | Cat# 12500096 | |
| Chemical compound, drug | FBS | Millipore | Cat# ES-009-B | |
| Chemical compound, drug | mLIF | Millipore | Cat# ESG1107 | |
| Chemical compound, drug | PD0325901 | APExBio | Cat# A3013 | |
| Chemical compound, drug | CHIR99021 | APExBio | Cat# A3011 | |
| Chemical compound, drug | N2 | Millipore | Cat# SCM012 | |
| Chemical compound, drug | N27 | Millipore | Cat# SCM013 | |
| Chemical compound, drug | MEM NEAA | Gibco | Cat# 11140–50 | |
| Chemical compound, drug | Penicillin-Streptomycin | Gibco | Cat# 11548876 | |
| Chemical compound, drug | L-Glutamin | Sigma-Aldrich | Cat# G7513 | |
| Chemical compound, drug | β-Mercaptoethanol | Sigma-Aldrich | Cat# M3148 | |
| Chemical compound, drug | Accutase | Millipore | Cat# SF006 | |
| Chemical compound, drug | Fugene6 | Promega | Cat# E2691 | |
| Chemical compound, drug | Puromycin | Sigma-Aldrich | Cat# P9620 | |
| Chemical compound, drug | Doxycycline | Sigma-Aldrich | Cat# D9891 | |
| Chemical compound, drug | Protease inhibitors | Bimake | Cat# B14001 | |
| Chemical compound, drug | Gelatin | Sigma-Aldrich | Cat# G1890 | |
| Chemical compound, drug | One Step-RNA Reagent | Bio Basic | Cat# BS410A | |
| Chemical compound, drug | DNaseI | NEB | Cat# M0303L | |
| Chemical compound, drug | SuperScript II Reverse Transcriptase | Invitrogen | Cat# 18064014 | |
| Chemical compound, drug | SsoAdvanced Universal SYBR Green Supermix | Bio-Rad | Cat# 1725270 | |
| Chemical compound, drug | Q5 High-Fidelity DNA Polymerase | NEB | Cat# M0491L | |
| Chemical compound, drug | Control LNA | Qiagen | Cat# 339137 | |
| Chemical compound, drug | anti-let-7 LNA | Qiagen | Cat# YFI0450006 | |
| Commercial assay or kit | Alkaline Phosphatase Assay Kit | System Biosciences | Cat# AP100R-1 | |
| Commercial assay or kit | Gibson Assembly Master Mix | NEB | Cat# E2611L | |
| Commercial assay or kit | Pierce BCA Protein Assay Kit | Thermo Fisher Scientific | Cat# 23225 | |
| Commercial assay or kit | Mir-X miRNA First Strand Synthesis Kit | Takara | Cat# 638313 | |
| Cell line (Mus musculus) | ES-E14TG2a mESC | ATCC | CRL-1821 | |
| Cell line (Mus musculus) | FLAG-Ago1 mESC | This paper | ||
| Cell line (Mus musculus) | FLAG-Ago2 mESC | PMID:33599613 | ||
| Cell line (Mus musculus) | Mir182∆ mESC | This paper | ||
| Cell line (Mus musculus) | Mir183∆ mESC | This paper | ||
| Cell line (Mus musculus) | Mir182∆/Mir183∆ mESC | This paper | ||
| Cell line (Mus musculus) | 3'UTR Mutant mESC | This paper | ||
| Cell line (Mus musculus) | Mir182∆/Mir183∆/3'UTR Mutant mESC | This paper | ||
| Recombinant DNA reagent | PiggyBac-based dox-inducible expression vector | PMID:33599613 | pWH406 | |
| Recombinant DNA reagent | Inducible GFP expressing vector | PMID:33599613 | pWH1055 | |
| Recombinant DNA reagent | Inducible mouse Mir182 expressing vector | This paper | pWH1039 | |
| Recombinant DNA reagent | Inducible mouse Mir183 expressing vector | This paper | pWH1040 | |
| Recombinant DNA reagent | sgRNA and Cas9 expressing vector (pX458) pWH464 | Addgene | Cat# 48138 | |
| Recombinant DNA reagent | Super PiggyBac Transposase expressing vector (pWH252) | System Biosciences | Cat# PB210PA-1 |
All the antibodies, plasmids, and oligonucleotides used in this study are listed in Supplementary file 1.
Cell lines
Request a detailed protocolAll the cell lines from this study are based on ES-E14TG2a mESC (ATCC, CRL-1821). They are listed in Supplementary file 1. The ES-E14TG2a mESCs were authenticated through STR profiling and were negative for mycoplasma contamination determined by a PCR-based kit.
CRISPR/Cas9-mediated genome editing in mESCs
Request a detailed protocolTo generate the FLAG-Ago1, FLAG-Ago2 mESCs, or Ago2 3’UTR mutant mESCs, cells were co-transfected with 2 µg of pWH464 (pSpCas9(BB)-2A-GFP [pX458]) expressing the corresponding targeting sgRNA and 1 µg of the corresponding donor oligo or plasmid using the Fugene6 (Promega). To generate Mir182Δ and Mir183Δ mESCs, cells were transfected with 2 µg of pWH464 expressing a pair of sgRNAs targeting pri-Mir182 or pri-Mir183. The transfected cells were subject to single cell sorting and the resulting clones were subject to genotyping to identify the correct clones.
qRT-PCR
Request a detailed protocolFor pri-miRNA quantification, reverse transcription was performed using random hexamers and Superscript II Reverse Transcriptase. Pre-miRNA and miRNA quantifications were using the Takara’s Mir-X miRNA quantification method. qPCR was performed in triplicate for each sample using the SsoAdvanced Universal SYBR Green Supermix (Bio-Rad) on a CFX96 real-time PCR detection system (Bio-Rad).
Western blotting
Request a detailed protocolProteins were harvested in RIPA buffer (10 mM Tris-HCl pH 8.0, 140 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, 1% Triton X-100, 0.1% sodium deoxycholate, 0.1% SDS, and protease inhibitor cocktail) and quantified with a BCA Protein Assay Kit (Thermo Fisher Scientific). Equal amounts of protein samples were resolved by SDS-PAGE, and then transferred to PVDF membranes. Western blotting was performed using a BlotCycler (Precision Biosystems) with the corresponding primary and secondary antibodies. The membranes were then treated with the Western ECL substrate (Bio-Rad), and the resulting signal was detected using an ImageQuant LAS 500 instrument (GE Healthcare).
Colony formation assay and exit pluripotency assay
Request a detailed protocolFor colony formation assay, 500 cells were plated on a 12-well plate in 2i + Lif media or Lif media (DMEM/F12 supplemented with 15% FBS, 1× penicillin/streptomycin, 0.1 mM non-essential amino acids, 2 mM L-glutamine, 0.1 mM 2-mercaptoethanol, and 1000 U/ml Lif). For exit from pluripotency assay, 1000 cells were plated on a gelatin-coated six-well plate in differentiation media (DMEM/F12 supplemented with 15% FBS, 1× penicillin/streptomycin, 0.1 mM non-essential amino acids, 2 mM L-glutamine, and 0.1 mM 2-mercaptoethanol) for 2 days, then cultured in 2i + Lif media for another 5 days. Colonies were stained using AP staining kit and grouped by differentiation status 6–7 days after plating.
Embryoid body formation
Request a detailed protocolFor differentiation via embryoid body (EB) formation, 3 × 106 cells were plated per 10 cm bacterial grade Petri dish and maintained on a horizontal rotator with a rotating speed of 30 rpm in differentiation media. The resultant EBs were harvested at the indicated time points.
RNA antisense purification mESCs were crosslinked with 0.1% formaldehyde for 5 min at room temperature, and the crosslinking reaction was quenched by adding 1/20 volume of 2.5 M glycine and incubating the mESCs at room temperature for 10 min on a rotating platform. The cells were then harvested and lysed in cell lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 5 mM EDTA, 10% glycerol, 1% Tween-20, with freshly added proteinase inhibitors). The cell lysate was cleared by centrifugation at 20,000 g for 10 min at 4°C. The resulting supernatant was used for RNA antisense purification; 5 mg lysate in 500 µl lysis buffer was used for each purification. Specifically, a set of 5’-end biotinylated anti-sense DNA oligos and 5 µl RNase inhibitor (NEB) were added to the lysate, resulting in a final concentration of 0.1 µM for each oligo. The lysate was incubated at room temperature for 1 hr on a rotating platform. Then, 100 µl Dynabeads MyOne Streptavidin C1 (Invitrogen) was added and the lysate further incubated for 30 min at room temperature on a rotating platform. The magnetic beads were isolated through a magnetic stand and then subject to four washes, with each wash in 500 µl high salt wash buffer (5× PBS, 0.5% sodium deoxycholate, 1% Triton X-100). The washed beads were resuspended in 100 µl DNaseI digestion mix (1× DNaseI digestion buffer with 5 µl DNaseI [NEB]) and incubated at 37°C for 20 min, followed by adding 350 µl LET-SDS buffer (25 mM Tris-HCl pH 8.0, 100 mM LiCl, 20 mM EDTA pH 8.0, 1% SDS) and 50 µl proteinase K (20 mg/ml, Thermo Fisher Scientific). The beads were then incubated on a thermomixer at 55°C 1000 rpm for 2 hr. The RNA was isolated through phenol extraction and isopropanol precipitation with glycoblue (Ambion) as a coprecipitant.
Data availability
All data generated or analyzed during this study are included in the manuscript and supporting files. Source data files have been provided for Figures 1–5.
-
NCBI Gene Expression OmnibusID GSE23943. Epigenome and transcriptome of naive pluripotent mouse embryonic stem (ES) cells cultured in 2i serum-free medium.
-
NCBI Gene Expression OmnibusID GSE138284. Studies on Trim71 in mouse embryonic stem cells.
References
-
Aberrant regulation and function of microRNAs in cancerCurrent Biology 24:762–776.https://doi.org/10.1016/j.cub.2014.06.043
-
Diversifying microRNA sequence and functionNature Reviews. Molecular Cell Biology 14:475–488.https://doi.org/10.1038/nrm3611
-
Let-7 micrornas in development, stem cells and cancerTrends in Molecular Medicine 14:400–409.https://doi.org/10.1016/j.molmed.2008.07.001
-
The Microrna-183 cluster: The family that plays together stays togetherNucleic Acids Research 43:7173–7188.https://doi.org/10.1093/nar/gkv703
-
Regulation of Microrna biogenesisNature Reviews Molecular Cell Biology 15:509–524.https://doi.org/10.1038/nrm3838
-
Argonaute proteins: Functional insights and emerging rolesNature Reviews. Genetics 14:447–459.https://doi.org/10.1038/nrg3462
-
Mechanisms of microrna-mediated gene regulation in animal cellsTrends in Genetics 23:243–249.https://doi.org/10.1016/j.tig.2007.02.011
-
The let-7 family of micrornasTrends in Cell Biology 18:505–516.https://doi.org/10.1016/j.tcb.2008.07.007
-
Homeostatic control of argonaute stability by microrna availabilityNature Structural & Molecular Biology 20:789–795.https://doi.org/10.1038/nsmb.2606
-
Lin28: Roles and regulation in development and beyondDevelopment 142:2397–2404.https://doi.org/10.1242/dev.117580
Article and author information
Author details
Funding
Mayo Foundation for Medical Education and Research
- Qiuying Liu
- Mariah K Novak
- Rachel M Pepin
- Taylor Eich
- Wenqian Hu
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
We thank Dr Xiaoli Chen for his assistance with miRNA prediction. This work is supported by Mayo Foundation for Medical Education and Research.
Copyright
© 2021, Liu et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 1,810
- views
-
- 234
- downloads
-
- 23
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 23
- citations for umbrella DOI https://doi.org/10.7554/eLife.72289






