A host beetle pheromone regulates development and behavior in the nematode Pristionchus pacificus
Abstract
Nematodes and insects are the two most speciose animal phyla and nematode–insect associations encompass widespread biological interactions. To dissect the chemical signals and the genes mediating this association, we investigated the effect of an oriental beetle sex pheromone on the development and behavior of the nematode Pristionchus pacificus. We found that while the beetle pheromone is attractive to P. pacificus adults, the pheromone arrests embryo development, paralyzes J2 larva, and inhibits exit of dauer larvae. To uncover the mechanism that regulates insect pheromone sensitivity, a newly identified mutant, Ppa-obi-1, is used to reveal the molecular links between altered attraction towards the beetle pheromone, as well as hypersensitivity to its paralyzing effects. Ppa-obi-1 encodes lipid-binding domains and reaches its highest expression in various cell types, including the amphid neuron sheath and excretory cells. Our data suggest that the beetle host pheromone may be a species-specific volatile synomone that co-evolved with necromeny.
https://doi.org/10.7554/eLife.03229.001eLife digest
The nematode worm Pristionchus pacificus can live as a parasite inside the oriental beetle, where it waits for the beetle to die so it can feed off the bacteria that live on the beetle's decomposing carcass. This ecologically important interaction is called necromeny. P. pacificus is attracted to a new host by a sex pheromone produced by the beetle, but the genes and biological mechanisms that enable this interaction to occur are not understood in much detail.
To identify the genetic basis of this interaction, Cinkornpumin et al. identified and examined a mutant form of P. pacificus that cannot sense the beetle sex pheromone. This revealed that although this pheromone attracts the adult nematodes, it stops P. pacificus embryos developing and can paralyze larvae. Cinkornpumin et al. suggest that the pheromone has likely evolved this ability in order to counteract the spread of the nematodes. This result implies that being invaded by P. pacificus makes life more difficult for the beetles than was previously thought.
Further investigation of the gene damaged in the P. pacificus mutants revealed that it encodes a protein that may bind to molecules called lipids, which are needed to form cell membranes and are used in cell signaling. As well as helping the nematodes to detect the sex pheromone, the lipid-binding protein also appears to help protect the worms from the pheromone's detrimental effects.
Cinkornpumin et al. observed that the gene for the lipid-binding protein is activated in several tissues, including the cells that form a sheath around some of the nerves that detect chemical signals. Whether this tissue is responsible for the chemical-sensing abilities of the lipid-binding protein, and whether these same tissues are responsible for protecting the nematodes from the damaging effects of the pheromone, remains to be discovered.
https://doi.org/10.7554/eLife.03229.002Introduction
An understanding of ecological interactions and evolutionary history can be beneficial to the understanding of the cellular and developmental processes of an organism. For example, the genes responsible for fruit and bacterial odor preferences in Drosophila melanogaster and Caenorhabditis elegans are most informative when considering the natural ecologies in rotting fruits and vegetation (Bargmann et al., 1993; Bargmann, 2006; Hallem and Carlson, 2006; Kiontke and Sudhaus, 2006). In Pristionchus pacificus, an emerging model organism for the study of development and behavior (Hong and Sommer, 2006b), a systematic effort to identify the natural ecology of Pristionchus nematodes revealed that P. pacificus host preferences include the oriental beetle (Exomala orientalis) found in Japan and northeastern United States (Herrmann et al., 2007). Unlike several fructivorous Caenorhabditis species, Pristionchus species are considered to be necromenic nematodes that have species-specific beetle host preferences and feed on the microorganisms that emerge from the beetle carcass (Herrmann et al., 2006a, 2006b; Brown et al., 2011). Pristionchus species are members of the Diplogasteridae family of nematodes whose common ancestor with the Rhabditidae family (e.g., C. elegans) diverged ∼300 million years ago (Dieterich et al., 2008). Given that beetles (Coleoptera) represent a group with the most described species, the possibility that most beetle species can harbor specific entomophilic nematode associations underscores the vast number of nematode species remaining to be described. Although biologists have been aware of such species-specific insect–nematode associations for some time, the genetic basis for such interactions remains largely unknown.
Previous studies suggest that the beetle host specificity of Pristionchus species is in part due to nematode attraction to various plant volatile compounds released during beetle feeding, as well as to beetle aggregation and sex pheromones during mating (Hong and Sommer, 2006a; Hong et al., 2008b, 2008a). For example, Pristionchus maupasi associates primarily with the European May beetle Melolontha spp. and is attracted to the blend of the green leaf alcohol from plants and the phenol pheromone from beetles (Hong et al., 2008b). Thus, insect and plant odors can synergize to provide nematodes with information on host presence, sex, and maturity. By contrast, C. elegans attractants such as 2,3-butanedione (diacetyl), 2,3-pentanedione, and isoamyl alcohol are significantly less attractive to P. pacificus (Bargmann et al., 1993; Hong and Sommer, 2006a). This piggybacking of intra-species communication for inter-species interactions between Pristionchus and beetles can also evolve rapidly at the population level, as exhibited by the natural variation of P. pacificus odor preference for insect pheromones (Hong et al., 2008a; McGaughran et al., 2013b). In particular, P. pacificus Washington strain (PS1843) is attracted to the lepidopteran sex pheromone E-11-tetradecenyl acetate (ETDA) and the oriental beetle sex pheromone z-7-tetradece-2-one (ZTDO) (Herrmann et al., 2007), but the wild-type reference strain from California (PS312) requires exogenous cGMP treatment to become attracted to these insect pheromones (Hong et al., 2008a). These natural variations in insect pheromone attraction, combined with regional-specific P. pacificus host preferences, may offer support for the geographic mosaic theory of co-evolution (Thompson, 2005).
Although the distinct chemosensory profiles of P. pacificus populations suggest insect pheromone attraction to be an important and rapidly evolving trait, other studies on insect–nematode interactions show that there are other potential roles host chemicals can play in coordinating host–parasite development and nematode dispersal behavior. For example, the pinewood nematode Bursaphelenchus xylophilus is intimately associated with its Monochamus beetle vector, and fatty acid ethyl esters exuded during beetle eclosion promote the formation of the dispersal fourth larval stage B. xylophilus (Zhao et al., 2013). Similarly, the species-specific phoretic association between the burrower bug Parastrachia japonensis and the nematode Caenorhabditis japonica may be mediated by the beetle cuticular compounds that not only attract C. japonica but also assist infestation by inhibiting C. japonica dispersal (Okumura et al., 2013). P. pacificus populations have been found on various beetle hosts worldwide, but the association with the oriental beetle was the first to be identified and the only one whose sex pheromone is a strong attractant (Herrmann et al., 2007; McGaughran et al., 2013b). Therefore, the oriental beetle sex pheromone represents the single most ecologically relevant semiochemical by which we could repeatedly interrogate the P. pacificus genes involved for host sensing and other unexplored developmental roles.
In this study, we show that although the beetle pheromone, ZTDO, is attractive to P. pacificus adults, the beetle host pheromone may moderate nematode infection by arresting embryonic development, inducing larval paralysis, and inhibiting dauer exit. We leveraged the ability to use exogenous cGMP to potentiate the California reference strain for ZTDO attraction to find additional genes important for ZTDO sensitivity by performing an unbiased forward genetic screen for mutants with altered responses to ZTDO. We found that a putative lipid-binding protein Ppa-OBI-1 mediates both beetle pheromone attraction as well as a previously unknown role of the beetle pheromone to regulate P. pacificus development. Our results suggest that the host pheromone may play a complex role in reducing infection or coordinating development for the dispersal cycle of the necromenic lifestyle.
Results
Ppa-obi-1 mutants show altered ZTDO attraction
P. pacificus and C. elegans have diametrically opposed odor preference profiles. Pristionchus species are generally attracted to plant volatiles, insect pheromones, and cuticular hydrocarbons, but each Pristionchus species can have its own chemosensory profile reflective of its preferred beetle hosts (Hong and Sommer, 2006a; McGaughran et al., 2013a). P. pacificus populations found on the oriental beetle E. orientalis in Japan and northeastern United States are attracted to the sex pheromone of the beetle, z-7-tetradecen-2-one (ZTDO) (Zhang et al., 1994; Herrmann et al., 2007). In contrast, the compost-dwelling C. elegans is less attracted to or even repelled by plant and insect odors, including ZTDO, but is strongly attracted to small compounds released by microbes (Bargmann et al., 1993; Hong and Sommer, 2006a).
To identify genes required to mediate ZTDO attraction in a pheromone-insensitive reference strain, California PS312, we performed a behavior-based genetic screen for mutants that are insensitive toward ZTDO following a brief 1-hr treatment with the cell-permeable 8-br-cGMP. We isolated two Oriental Beetle pheromone Insensitive mutants, Ppa-obi-1(tu404) and Ppa-obi-3(tu405). The chemosensory phenotype of Ppa-obi-1 and Ppa-obi-3 is strongly specific for ZTDO and does not affect the chemoattraction towards other insect pheromones ETDA and myristate nor plant volatiles β-caryophyllene and nicotine (Figure 1). Odor avoidance in obi mutants towards 1-octanol, a repellant common between P. pacificus and C. elegans, also resembles wild type. These results indicate that both obi mutants are specifically defective in sensing a beetle pheromone rather than more generally defective in chemosensation. Ppa-obi-1(tu404) and Ppa-obi-3(tu405) are non-allelic recessive mutants that complement each other in their physical and behavioral phenotypes. We proceeded to characterize Ppa-obi-1(tu404) based on its altered response to ZTDO attraction and its unexpected function in protecting larvae.
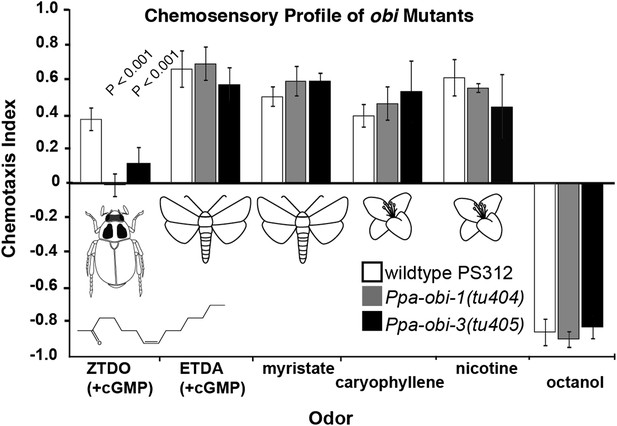
Chemosensory behavior of oriental beetle pheromone insensitive mutants.
Ppa-obi-1 and Ppa-obi-3 mutant animals are specifically defective for sensing the host oriental beetle sex pheromone, Z-7-tetradece-2-one. The ZTDO structure is depicted below the beetle. In contrast, obi mutants' response is wild type towards another cGMP-dependent attractant, E-11-tetradecenyl acetate (ETDA, a lepidopteran pheromone), as well as myristate (methyl tetradecanoate, an insect allomone), plant volatiles beta-caryophyllene and nicotinic acid, and the repellent 1-octanol. N ≥ 15 replicates in ≥3 trials. Significant differences to wild type are indicated by p values (Dunnett's test) and error bars represent SEM.
ZTDO is an unsuspected potent embryocide and anesthetic of P. pacificus larva
Necromeny occupies a midpoint in the continuum of nematode–insect interactions that span from mutualism to pathogenicity. To address possible developmental effects of long-term exposure to ZTDO, we attempted to grow worms in the presence of the ZTDO by placing eggs onto OP50 bacteria in plates containing 10 µl of 0.5% ZTDO suspended under the plate lids, which is approximately a third of the amount used in chemotaxis assays. Surprisingly, very few of the synchronized eggs from wild type and Ppa-obi-1(tu404) hatched after 36 hr, compared to ∼94% of the untreated eggs (Figure 2A). ZTDO therefore may function not just as a volatile attractant but also as an antagonistic host-derived developmental regulator.
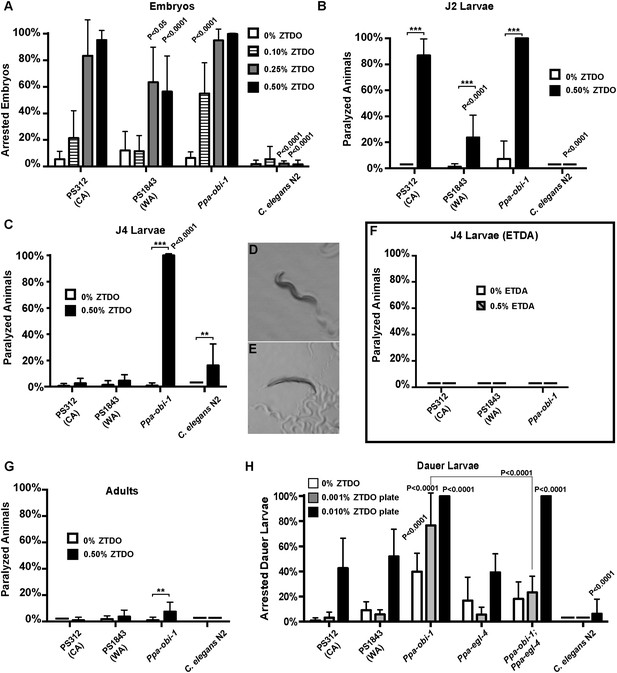
The oriental beetle pheromone ZTDO inhibits P. pacificus development and induces paralysis.
(A) Ppa-obi-1 mutant is hypersensitive to 0.10% ZTDO compared to wild type PS312 (California). However, PS312 is more sensitive to 0.5% ZTDO than another strain, PS1843 (Washington). PS312 (n = 372, 82, 90, 320); PS1843 (n = 309, 88, 92, 237); Ppa-obi-1 (n = 332, 69, 71, 389); C. elegans wild-type N2 (n = 316, 63, 84, 174). (B) Both wild-type PS312 and Ppa-obi-1 mutant J2 larvae are more sensitive to ZTDO than PS1843. PS312 (n = 154, 115); PS1843 (n = 110, 102); Ppa-obi-1 (n = 104,100); C. elegans wild-type N2 (n = 23, 73). (C) Ppa-obi-1 J4 larvae is uniquely sensitive to ZTDO compared to PS312 (p < 0.0001). PS312 (n = 209, 290); PS1843 (n = 102, 99); obi-1 (n = 244, 237); C. elegans wild-type N2 (n = 69, 109). (D) 90 min exposure of a Ppa-obi-1(tu404) J4 larvae to ethanol control compared to (E) 0.50% ZTDO resulted in a rigid fully paralyzed posture. (F) Adults are not paralyzed by ZTDO. PS312 (n = 108, 113); PS1843 (n = 114, 107); Ppa-obi-1 (n = 109, 109); C. elegans wild-type N2 (n = 62, 72). (G) 0.01% ZTDO prevents resumption of post-dauer development in approximately 40% of wild-type animals and nearly all Ppa-obi-1(tu404) mutants. This ZTDO inhibition of dauer exit is dependent on the cGMP-dependent PKG, Ppa-egl-4. PS312 (n = 384, 654, 267); PS1843 (n = 195, 139, 101); Ppa-obi-1 (n = 339, 343, 146); Ppa-egl-4(tu374) (n = 123, 181, 77); Ppa-obi-1;Ppa-egl-4 (n = 132, 144, 63); N2 (n = 131, 109, 115). (H) Exposure to 0.50% ETDA moth pheromone from Spodoptera littoralis does not paralyze Ppa-obi-1 mutant and wild-type J4 larvae. PS312 (n = 40, 103); PS1843 (n = 40, 64); Ppa-obi-1 (n = 40, 193). p values indicate Dunnett's multiple comparisons test to wild-type PS312. ***p < 0.001; **p < 0.01 indicate difference between ethanol control and ZTDO exposed groups (pairwise unpaired t test). Error bars are SEM and n is the number of nematodes tested.
Detailed examination revealed that Ppa-obi-1(tu404) eggs had significantly lower hatching rate in 0.10% ZTDO compared to wild-type PS312. Interestingly, the Washington strain that has natural cGMP-independent attraction to ZTDO was more resistant to 0.25–0.50% ZTDO than the ZTDO-insensitive California strain. Although ZTDO exposure had no noticeable effect on egg laying output by adult hermaphrodites, eggs continuously exposed to ZTDO for 2 days at an early stage remain arrested at the pre-comma globular embryonic stage, before gastrulation. The embryocidal effect of ZTDO was concentration-dependent and species-specific, since wild-type C. elegans eggs were completely unaffected (Figure 2A). Ppa-obi-1(tu404) hypersensitivity to the developmental and paralyzing effects of ZTDO also continued into the subsequent J2 and J4 larval stages (Figure 2B,C). Both wild-type PS312 and Ppa-obi-1(tu404) J2 larvae exhibited ZTDO-induced paralysis, while the Washington strain was significantly more resistant to paralysis. However, by the J4 larval stage, only Ppa-obi-1(tu404) was sensitive to ZTDO compared to both wild-type P. pacificus strains (Figure 2D–E). Ppa-obi-1(tu404) J4 larvae exposed to ZTDO displayed initial agitation, followed by full body paralysis between 90 and 120 min. In contrast, wild-type C. elegans J4 larvae displayed only minute sensitivity to ZTDO. To determine if the Ppa-obi-1(tu404) mutation renders it susceptible to paralysis from other insect pheromones, we also tested the moth pheromone ETDA and found that the J4 larvae of Ppa-obi-1(tu404) and the wild-type strains were not affected (Figure 2H). All strains, including Ppa-obi-1(tu404), were insensitive to ZTDO-induced paralysis when they reach adulthood (Figure 2F), the dominant stage used in our chemotaxis assays. Taken together, our findings show that ZTDO is capable of arresting wild-type P. pacificus embryogenesis and severely paralyzing J2 larvae, indicating that ZTDO is a potential host-specific developmental regulator. We speculate that wild-type Ppa-OBI-1 may serve to protect P. pacificus embryos and reproductive larvae against the volatile anesthetic-like effects of ZTDO from the adult beetle host.
ZTDO inhibits dauer larva exit in P. pacificus
The dauer larva (DL) is an alternative, non-reproductive, long-lasting stage in nematodes capable of withstanding starvation, as well as the infective stage in parasitic nematodes (Lee, 2002). To explore the possibility that ZTDO also acts to restrain the dispersive stage of P. pacificus development after hatching, we asked if ZTDO can also regulate DL exit in the presence of food using culturing plates containing 0.001–0.01% ZTDO and bacteria. Nearly all untreated wild-type P. pacificus DL developed into J4 larvae after 2 days on food, whereas ∼40% of the wild-type worms remained as active DL on 0.01% ZTDO with OP50 food that would have otherwise promoted reproductive development (Figure 2G). However for Ppa-obi-1 DL, this dauer exit defect is exacerbated by ZTDO due to paralysis. Approximately 80% of the developmentally blocked Ppa-obi-1(tu404) DL show immobility on 0.001% ZTDO, and nearly 95% of the Ppa-obi-1(tu404) DL became paralyzed on 0.01% ZTDO. Although DL is the primary infective stage in many entomophilic nematodes found on live beetles (Weller et al., 2010), we were unable to assess DL's chemotaxis response to ZTDO because of its immobilizing effect on DL. The negative regulation of DL exit by ZTDO is specific to P. pacificus and barely affected C. elegans even at 0.01% ZTDO. These results show that 0.01% ZTDO can block active wild-type P. pacificus DL from progressing to the reproductive stage without causing paralysis, but 0.001–0.01% ZTDO significantly reduced dauer exit in Ppa-obi-1 DL by primarily inducing paralysis and death.
Since Ppa-obi-1(tu404) has a defect in DL exit that is further enhanced by exposure to ZTDO, we asked if DL arrest in Ppa-obi-1(tu404) by ZTDO is similarly dependent on Ppa-EGL-4 function. Egl-4 mutants are defective in a cGMP-dependent protein kinase in C. elegans, which affect dauer formation as well as chemosensation (Daniels et al., 2000; L'Etoile et al., 2002). We found that while the Ppa-egl-4(tu374) loss-of-function mutation did not significantly affect the ability to exit the dauer stage, the Ppa-egl-4(tu374) mutation in the Ppa-obi-1(tu404) background did partially suppress the inhibitory effect of ZTDO, such that Ppa-egl-4;Ppa-obi-1 post-dauer development on 0.001% ZTDO is higher than Ppa-obi-1(tu404) (Figure 2G). The Ppa-egl-4(tu374) mutation did not, however, reduce the ZTDO inhibition of DL exit in the presence of a higher amount of 0.01% ZTDO. This outcome suggests that the susceptibility of Ppa-obi-1(tu404) DL to ZTDO is partially dependent on Ppa-egl-4. While the loss of Ppa-obi-1 alone can cause reduced rate of DL exit on food independent of ZTDO, the aggravated DL arrest by ZTDO in Ppa-obi-1(tu404) is partially mediated by Ppa-egl-4.
Although wild-type P. pacificus adult animals are attracted to ZTDO, the host beetle sex pheromone regulates P. pacificus development by inhibiting embryogenesis, restricting DL exit, and inducing paralysis in larval stages. This ZTDO sensitivity is the most acute at the embryonic stage and is likely to be buffered by wild-type Ppa-OBI-1 in later developmental stages as the strongest hypersensitivity to ZTDO was observed in the J4 stage of the Ppa-obi-1(tu404) mutant. By contrast, the PS1843 Washington strain that is naturally attracted to ZTDO is more resistant to ZTDO in non-dauer stages compared to the PS312 California wild type. Further studies in other natural populations are needed to determine the correlation between ZTDO attraction and resistance to ZTDO-induced paralysis in P. pacificus.
Molecular cloning of Ppa-obi-1(tu404)
In addition to the specific chemosensory defect in Ppa-obi-1(tu404) mutants, we also noticed a linked phenotype in its body morphology. Detailed measurements indicate that Ppa-obi-1(tu404) animals are significantly longer and thinner than wild type (Figure 3), resulting in a slim body phenotype with adult Ppa-obi-1(tu404) hermaphrodites ∼20% longer than wild-type animals. The slim body phenotype may also contribute to Ppa-obi-1(tu404)'s moderate egg-laying defect (egl; Table 1) and reduction in hatching rate similar to Ppa-egl-4(tu374) mutants, possibly due to defects in vulva formation and cuticle development. Using molecular markers and the slim phenotype, we were able to delineate the Ppa-obi-1 lesion to an ∼138 kb region on Supercontig 1 of chromosome I between markers SSLP21 and SSLP17 (Figure 4A). We subsequently identified all polymorphic sites between PS312 and Ppa-obi-1(tu404) in the fine mapping interval by Illumina whole genome sequencing and focused on disruptions of predicted open reading frames. We found a single base pair deletion in a predicted coding region orthologous to C06G1.1 in C. elegans and confirmed this tu404 deletion by PCR amplification and Sanger sequencing of both wild type and Ppa-obi-1(tu404) genomic and cDNA. The tu404 allele contains a single base pair deletion in exon 9. The resulting frame shift and nonsense mutation are predicted to stop translation half way into the reading frame to result in a hypomorphic allele. In the two Ppa-obi-1 splice forms, the longer transcript contains a VKDDDPR peptide that may be the result of an alternative splice donor site. Both splice variants were isolated from the California and Washington strains in equal abundance by sub-cloning RT-PCR products (data not shown), and no differences in nucleotide sequence were found in the coding region of these two strains. Therefore, splice form variation in Ppa-OBI-1 is unlikely to contribute to the difference in insect pheromone attraction between these two strains. The region encoding for VKDDDPR is also absent in all the Ppa-obi-1 orthologs we have examined in available databases (Figure 4—figure supplement 1).
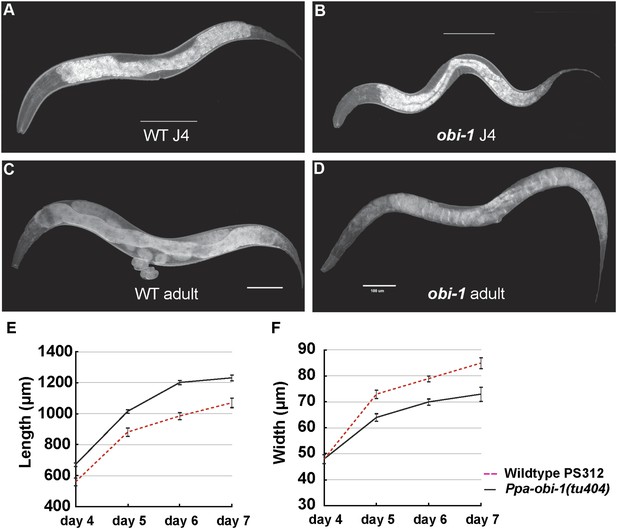
Ppa-obi-1 mutants are long and slim.
(A and B) Wild-type J4 larva and young adult. (C and D) Ppa-obi-1 J4 larva and young adult. (E and F) The body lengths and widths of wild-type (dotted red line) versus Ppa-obi-1 mutants (solid black line) recorded over 4 days post J4 stage (μm); n = 15–30 animals for each time point. Error bars are SEM. Scale bars denote 100 μm.
Ppa-obi-1 affects hatching rate and brood size
| Hatching rate (%) | Brood size (3 days) | n | |
|---|---|---|---|
| Wild-type(PS312) | 95 ± 3 | 156 ± 16 | 18 |
| Ppa-obi-1(tu404) | 85 ± 3 | 103 ± 17 | 15 |
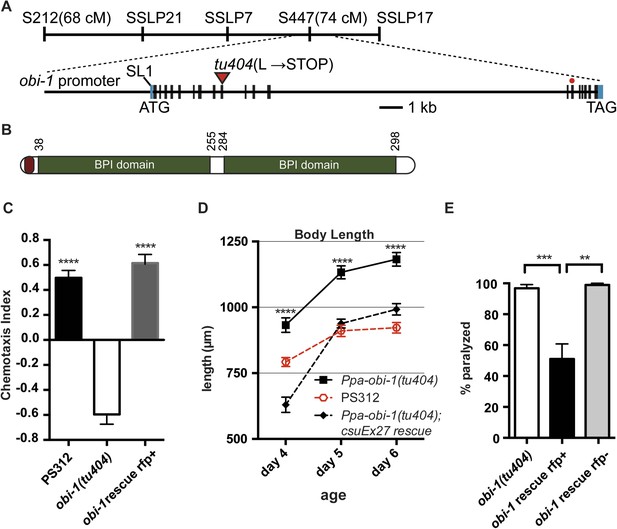
Ppa-obi-1 gene structure and protein comparisons.
(A) The order of mapping markers is depicted on top (not to scale). The Ppa-obi-1 mRNA is SL1 trans-spliced and contains 19 exons spanning ∼19 kb. The 50 bp 5′ UTR and 341 bp 3′ UTR are indicated in light blue. The tu404 allele contains a single base pair deletion in exon 9 that results in a frameshift, converting a codon to a stop codon (inverted triangles). The red dot represents the splice variant. (B) A diagram of the secretion and two BPI lipid-binding domains (PFAM and SMART). (C–E) Transgenic rescue of the Ppa-obi-1(tu404). (C) A 4.3 kb promoter driving Ppa-obi-1 cDNA completely rescued cGMP-dependent ZTDO attraction in Ppa-obi-1(tu404) mutants. PS312 (N = 21); obi-1 (N = 15); obi-1 rescue (N = 12). The Ppa-obi-1 transgenic rescue (D) restored the body length defect in Ppa-obi-1(tu404) mutants, PS312 (n = 15); obi-1 (n = 15); obi-1 rescue (n = 19) as well as (E) significantly reduced ZTDO-induced paralysis in Ppa-obi-1(tu404) J4 larvae. PS312 (N = 11); obi-1 rescue rfp+ (N = 12); obi-1 rescue rfp− (N = 5). **p < 0.01; ***p < 0.001; ****p < 0.0001 (Tukey's multiple comparisons test). Error bars represent SEM. N is the number of assays performed and n is the number of individual worms scored for each time point.
Transgenic rescue and comparison of lipid-binding proteins
To confirm that Ppa-obi-1 is indeed responsible for the lack of ZTDO attraction in adults and for hypersensitivity to ZTDO-induced paralysis in the J4 larvae, we made transgenic lines containing a 4.3 kb Ppa-obi-1 promoter driving the Ppa-obi-1 full-length cDNA (Schlager et al., 2009). The introduction of the wild-type Ppa-obi-1 transgene restored the cGMP-dependent ZTDO attraction in Ppa-obi-1(tu404) adults compared to Ppa-obi-1(tu404) adults and sibling animals that did not segregate for the rescue construct (Figure 4C). Furthermore, the Ppa-obi-1 rescue transgene significantly shortened the body length of Ppa-obi-1(tu404) adults (Figure 4D), confirming that both the ZTDO chemosensory and the long body length phenotype used for subsequent positional mapping were due to the same mutation in Ppa-obi-1(tu404). Lastly, we tested the ability of the wild-type Ppa-obi-1 transgene to reduce the ZTDO hypersensitivity and found that it significantly reduced the percentage of ZTDO-induced paralysis in Ppa-obi-1(tu404) J4 larvae (Figure 4E). Ppa-obi-1 therefore mediates ZTDO chemosensation, body dimensions, and ZTDO susceptibility.
Sequence analyses of coding regions show that Ppa-obi-1 and C06G1.1 encode for proteins that represent a class of highly conserved but functionally diverse group of secreted lipid-transfer/lipid-binding proteins present in almost all metazoans except Drosophila (Beamer et al., 1998), including lipopolysaccharide-binding proteins (LBP) (Tobias et al., 1997), bactericidal permeability-increasing proteins (BPI) (Weiss et al., 1978), phospholipid transfer proteins (PLTP) (Gautier and Lagrost, 2011), and cholesteryl ester transfer proteins in mammals (CETP) (Kawano et al., 2000). The LBP/BPI/CETP/PTLP family in humans is known to play a role in immunological responses to bacterial pathogens and mediate the transfer of cholesteryl ester and triglycerides in the blood plasma (Bruce et al., 1998; Kawano et al., 2000; Schultz and Weiss, 2007). Ppa-OBI-1 and C06G1.1 each contain two predicted lipid-binding domains as well as a predicted secretion peptide sequence in the N-terminal regions (PFAM and SMART; SignalP 4.0 Server) (Figure 4—figure supplement 1). To identify all members of the LBP family in the sequenced genome and transcriptome of P. pacificus, we performed reciprocal BLASTP searches (WormBase WS221), which indicated 12 members predicted to be encoded in the P. pacificus genome. Maximum likelihood phylogenetic analysis of the OBI-1/C06G1.1 LBP protein family indicates 10 closely related orthologous pairs in the two species. Interestingly, the lengths of the branches suggest that all the orthologs between P. pacificus and C. elegans share similar divergence times, with the exception of the OBI-1/C06G1.1 pair that seemed to have undergone faster divergence relative to other paralogs (Figure 4—figure supplement 2). Consistent with their roles in immune response in mammals, a member of the LBP family in C. elegans, F44A2.3, was found to be up-regulated in response to several species of pathogenic bacteria (Wong et al., 2007). Another C. elegans LBP member, NRF-5, is partially necessary for nose contraction in response to the anti-depressant fluoxetine (Prozac) (Choy et al., 2006), and for mediating phosphatidylserine transfer from apoptotic cells to engulfing cells during cell corpse removal (Zhang et al., 2012). We found that nrf-5(sa513) mutants are also defective in attraction to 2,3-butanedione (diacetyl), and isoamyl alcohol, although the C06G1.1(vc20357) allele with a missense mutation did not show chemosensory defects (Figure 4—figure supplement 3). Thus LBP members may have a conserved previously uncharacterized role in chemosensation in diverse nematodes.
To determine if C06G1.1 can also regulate body morphology in C. elegans, we performed C06G1.1 RNAi by feeding. C06G1.1 RNAi was previously observed to generate a long body phenotype in a large-scale RNAi screen in C. elegans (Simmer et al., 2003). We used this fragment (Fragment 1), along with another C06G1.1 fragment more proximal to the 3′ end (Fragment 2), to quantify the effect of C06G1.1 RNAi in the enhanced RNAi rrf-3(pk1426) strain. Although C06G1.1 RNAi did not significantly change the absolute body length of adult worms, we found RNAi against C06G1.1 using both fragments resulted in a significantly slimmer body proportion (length/width ratio) similar to the Ppa-obi-1(tu404) mutant (Figure 4—figure supplement 4). A missense mutation in C06G1.1(VC20357) also resulted in a similar slim body phenotype. Thus both Ppa-OBI-1 and its C. elegans ortholog also share a conserved function in regulating body proportions.
Ppa-obi-1 is expressed in cells with putative roles in secretion
To characterize which tissue types Ppa-OBI-1 functions in, we sought to determine the Ppa-obi-1 expression pattern using a Ppa-obi-1p::gfp transcriptional reporter gene containing a 4.3 kb promoter fragment upstream of the Ppa-obi-1 ATG initiation codon. Our analysis was based on a stably integrated transgenic line csuIs01 and parsimonious inferences to homologous C. elegans cell positions and known cellular functions. Detailed analyses of all developmental stages show that the earliest Ppa-obi-1 expression occurs in at least one cell in the pre-comma embryonic stage but soon disappears until weak expression reappears in the hyp7 and seam cells in J4 larvae (Figure 5). The most robust and diverse Ppa-obi-1 expression was found in mid-J4 hermaphrodites, which is consistent with real-time quantitative PCR results indicating that transcript levels in J2 and J3 larvae are less than in J4 larvae (data not shown). In the mid-J4 stage, Ppa-obi-1 is strongly expressed in the major hypodermal syncytium and the lateral hypodermal cells that cover most of the body (hyp7 and seam cells) as well as the ventral tail (hyp 8 and 9) (Figure 5). In contrast to young adult C. elegans, which have 16 seam cells on each lateral surface (H0–H3, V1–V6, T descendants; The WormAtlas), we counted only 15 seam cells per side in P. pacificus J4 larvae (n = 20). Ppa-obi-1 is also strongly expressed in the P(5–7).p cell descendants that form the vulva lumen (vulC, D, E) during the mid-J4 stage until just prior to vulval eversion as well as in the vulva muscles (likely vm1, 2) (Figure 5B inset; Figure 5—figure supplement 1).
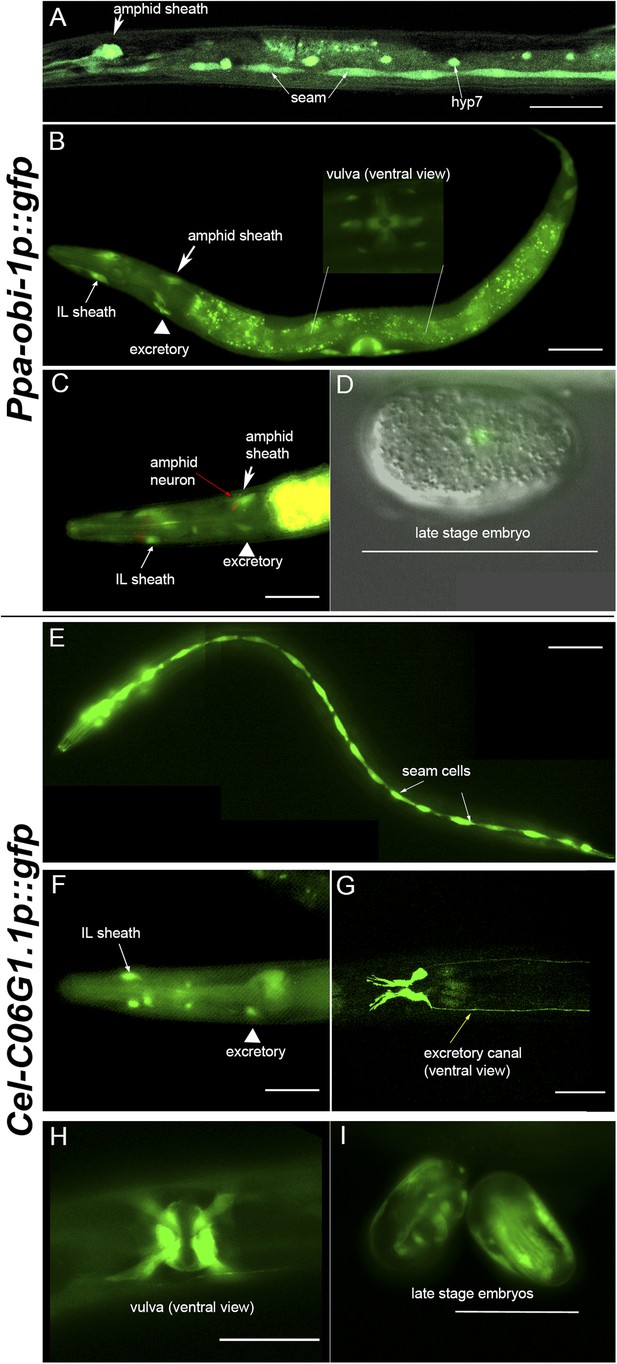
Ppa-obi-1 and C06G1.1 expression in their respective nematode species.
(A–D) An integrated Ppa-obi-1 promoter::gfp transgene (csuIs01) in P. pacificus. (A) Lateral view of a single animal showing prominent expression in an amphid sheath cell, seam cells, and the syncytial hypodermal cell (hyp7) in a J4 hermaphrodite. (B) Medial view of a transgenic animal with expression in the putative dorsal and ventral inner labial (IL) or outer labial (OL) neuronal support cells (socket or sheath), the duct and excretory cells on the ventral side, the amphid sheath cell on the dorsal side (arrow head), and the vulval muscles. The inset shows a ventral view of the young adult vulva in another animal. Autofluorescence is visible in the intestine. (C) A DiI-labeled neuron (red arrow) anterior to a Ppa-obi-1p::gfp expressing amphid sheath cell (white arrow) is dorsal to an excretory cell (arrow head). A putative IL or OL socket or sheath cell (arrow) is also visible. (D) DIC and fluorescent overlay image of expression in early P. pacificus embryo. (E–I) C06G1.1 promoter::gfp transgenes (csuEx02, csuEx03) in C. elegans. (E) Lateral view of a seam cell expression in a L3 hermaphrodite. (F) Medial view of putative inner or outer labial neuron support cells (arrow) and excretory cells (arrowhead). No amphid sheath shows gfp expression. (G) Ventral view of head region shows prominent expression in excretory cells and canal (arrow). (H) Ventral view of young adult vulva. (I) Embryonic expression in C. elegans embryos appears later, stronger, and in more cells than in Ppa-obi-1. Anterior is left and the scale bars represent 50 μm. Each panel is representative of ≥50 animals.
Most notably, Ppa-obi-1 expression was observed in the two bilaterally symmetrical specialized epithelial cells that envelope the chemosensory neurons, known as the amphid sheath cells (Figure 5A–B, arrows). The amphid sheath cells are support cells for the sensory amphid neurons and completely enclose the ciliated endings of the AWA, AWB, and AWC chemosensory neurons in C. elegans. Ppa-obi-1p::gfp expression in the putative amphid sheath cells lie dorsal–lateral and posterior to the DiI-labeled amphid neurons ASK, ADL, and ASI (Figure 5C). DiI staining of amphid neurons in Ppa-obi-1 mutants is superficially wild type, suggesting that Ppa-OBI-1 is unlikely to be involved in the development of the amphid neurons (data not shown). We also noticed expression in four unknown cell types anterior to the metacarpus (anterior pharyngeal bulb), which could be putative neuronal support cells for the inner labial or outer labial quadrant sensilla (IL or OLQ) (Figure 5—figure supplement 1). Ppa-obi-1 is also expressed in putative duct and excretory cells on the ventral anterior side that form part of the excretory system (Figure 5B, arrowhead). Based on conserved DiI labeling of P. pacificus amphid neurons and cell positions, the Ppa-obi-1 expression profile and secretion peptide sequence suggest that Ppa-OBI-1 is likely an extracellular protein involved in sensory neuron support, epithelial cell development, and osmoregulation.
Ppa-obi-1 but not C06G1.1 is expressed in glial-like sheath cells
Although Ppa-OBI-1 and its ortholog C06G1.1 in C. elegans share 61% amino acid identity, P. pacificus and C. elegans have opposite responses to the pheromone ZTDO. While small changes in sequence can lead to large changes in protein function, the absence of expression of the C. elegans obi-1 ortholog in amphid sheath cells suggests that changes in gene expression pattern may also play a role in determining ZTDO sensitivity. We therefore sought to determine if the Ppa-obi-1 ortholog C06G1.1 have shared tissue expression with Ppa-obi-1. We examined endogenous C06G1.1p::gfp expression in C. elegans in five independent transgenic lines with extrachromosomal arrays and found strong mid-L4 larvae expression in the seam cells, two tail hypodermal cells, four cells in the excretory duct system (duct, excretory, gland cells), vulval muscles, and possible IL or OLQ sheath or socket neuronal support cells similar to the expression pattern observed for Ppa-obi-1 (Figure 5E–H). In contrast to the earliest Ppa-obi-1 expression in pre-comma stage, C06G1.1 expression is detectable during late embryogenesis in putative seam cells (Figure 5I). This seam cell expression is earlier than Ppa-obi-1p::gfp in the L1 stage, and is maintained until the young adult stage. A reason for this earlier seam cell expression could be that the equivalent P. pacificus J1 larvae remain inside the egg while C. elegans eggs hatch as L1 larvae (Fürst von Lieven, 2005). Like Ppa-obi-1, C06G1.1 expression is largely absent in all tissue types when the transgenic animals become 1-day old adult hermaphrodites, coinciding with the fusion of seam cells into a syncytium. This observation indicates that C06G1.1 expression in the excretory and epithelial system may be coordinated and involved in seam cell development.
Our expression analysis of obi-1 orthologs in the their respective nematode species revealed that the most striking difference is the lack of expression in the amphid sheath cells in C. elegans, suggesting that unlike Ppa-obi-1, C06G1.1 is not likely to function directly with chemosensory neurons in C. elegans. In addition to differences in gene expression pattern, the possibility that there is a ZTDO receptor that interacts with Ppa-OBI-1 in P. pacificus but not in C. elegans may also help to explain how Ppa-obi-1 orthologues confer different ZTDO sensitivities in the two species during certain developmental stages. To explore yet another possibility that changes in both protein function and expression may lead to differences in ZTDO sensitivity, we expressed the Ppa-obi-1:cDNA under the C. elegans amphid sheath cell-specific promoters T02B11.3 and F16F9.3 in C. elegans (Bacaj et al., 2008; Kramer et al., 2010; Oikonomou et al., 2011; Procko et al., 2011). Both T02B11.3p::Ppa-obi-1:gfp(csuEx29) and F16F9.3p::Ppa-obi-1:gfp(csuEx30) transgenic animals showed strong GFP expression in the amphid sheath cells (Figure 5—figure supplement 2) as well as phasmid sheath cells in the tail region (not shown). While the targeted misexpression of Ppa-obi-1 in C. elegans did not alter the strong avoidance behavior towards either 2% or 10% ZTDO, Ppa-obi-1 expression in C. elegans did reduce the paralytic effects of ZTDO (Figure 5—figure supplement 2). Because wild-type C. elegans N2 is not susceptible to ZTDO-induced paralysis at 0.5% ZTDO that P. pacificus is, we increased the stringency of the assay by using 10% ZTDO, incubating for 16 hr, and exposing the animals in the absence of OP50 food. Under this enhanced ZTDO paralysis assay, we found that F16F9.3p::Ppa-obi-1:gfp(csuEx30) transgenic L4 larvae but not T02B11.3p::Ppa-obi-1:gfp(csuEx29) exhibited a small but statistically significant increase in protection from ZTDO-induced paralysis compared to wild-type N2 (Figure 5—figure supplement 2C,D). Because we could not detect apparent differences in GFP expression patterns between the two amphid promoters, the ability of F16F9.3p::Ppa-obi-1:gfp to reduce ZTDO paralysis may be due to subtle transient expression differences. These results suggest that while the expression of the Ppa-OBI-1 protein in the C. elegans amphid sheath cells is not sufficient to reduce avoidance to ZTDO, it can contribute to the obi-1-dependent protection from ZTDO-induced paralysis.
Discussion
Every organismal interaction is a legacy of its evolutionary arms race. We have identified an important and likely co-evolving trait in the interaction between P. pacifcus nematodes and its beetle hosts. Although not symbiotic, necromenic associations between nematodes and insects have been considered commensalistic or mutualistic, but our results suggest antagonistic interactions between the larval stages of P. pacificus and the adult oriental beetle hosts that produce the ZTDO sex pheromone. Attraction and resistance to ZTDO in adult P. pacificus further points to a refined attenuated antagonism in their co-evolutionary relationship (Thompson, 2005). Our data support the model that Ppa-OBI-1 is required to protect P. pacificus larvae against a host beetle pheromone ZTDO as well as to enable P. pacificus attraction to ZTDO. Furthermore, two different geographical isolates and developmental stages have varied responses to the oriental beetle pheromone. ZTDO may be a species-specific volatile synomone that acts through a lipid-binding protein to moderate nematode infection by arresting embryonic development, inducing larval paralysis, and inhibiting dauer exit. To the best of our knowledge, this is the first genetically defined reception to a species-specific compound mediating the interaction between a nematode and its insect host.
The role of Ppa-OBI-1 in ZTDO sensitivity
Sensitivity to ZTDO-induced paralysis/arrest is Ppa-obi-1-dependent predominantly during the J4 larval stage and, to a lesser degree, in the embryos and dauers. Given the diversity of tissues expressing Ppa-obi-1 in the J4 larvae, it is not apparent which tissues require Ppa-obi-1 expression for proper chemosensation and if the same tissues are involved with protection from ZTDO-induced paralysis in wild-type animals (Figure 6). Our gene expression studies show that despite coding highly similar proteins, P. pacificus Ppa-obi-1 is uniquely expressed in the amphid sheath cells compared to its C. elegans ortholog C06G1.1. Based on this species-specific expression pattern, we are tempted to speculate that these chemosensory glial cells, the IL and amphid sheath cells, play a dominant role in mediating ZTDO sensitivity. However, additional experiments will be needed to test the possibility that epidermal cells such as the vulva, hypodermis, and seam cells, or cells in the excretory system may also mediate ZTDO sensing.
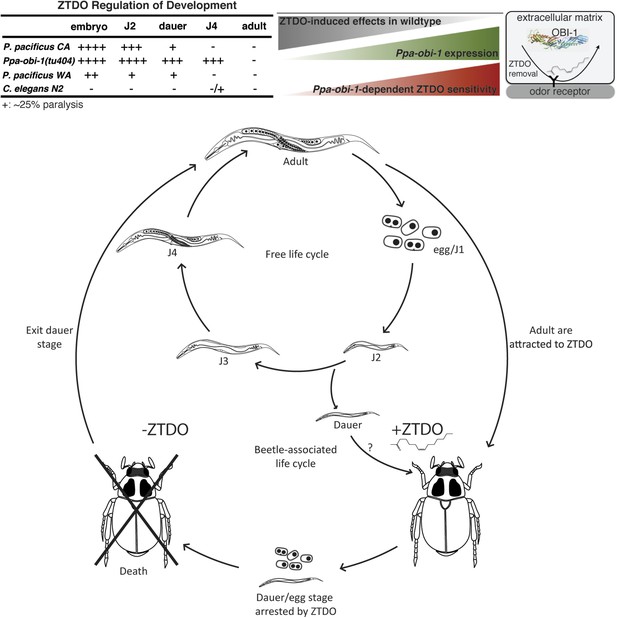
Summary and model of the oriental beetle pheromone ZTDO on P. pacificus development and lifecycle.
(Top) Peak Ppa-obi-1 expression in the wild-type J4 larvae coincides with the least sensitive developmental stage to the paralytic effects of the beetle pheromone ZTDO (at 0.5%). Ppa-obi-1 mutants are most sensitive to ZTDO-induced paralysis in the J4 stage. Our working model on Ppa-OBI-1 functions in the J4 larvae hypothesizes that Ppa-OBI-1 proteins in the extracellular space, such as the lumen of the amphid sheath cell, may be required to remove ZTDO bound to odor receptors on neurons. (Bottom) This working model shows both P. pacificus adults and dauer larvae from non-beetle populations associate with the host oriental beetle through attraction to the beetle's sex pheromone ZTDO, whereby the progeny born on the beetle are arrested as eggs or dauer larvae. Death of the host allows microorganism to grow, which then encourages dauer larvae to exit and resume reproductive development. Several iterations of asynchronous nematode populations likely complete each dispersal cycle.
Given the growing evidence that the amphid sheath cells in C. elegans are critical for proper neuronal development and function of the 12 pairs of chemosensory amphid neurons (Shaham, 2006), we propose the following models on how the Ppa-OBI-1 protein can act in the sheath cells to directly or indirectly modulate ZTDO chemosensation. In C. elegans, many proteins have enriched expression in the amphid sheath, such as DAF-6 and the secreted venom allergen-like VAP-1, and these secretions are corroborated by dense populations of vesicles in amphid sheath cells under electron microscopy (Bacaj et al., 2008; Doroquez et al., 2014). However, it is not yet known which amphid proteins, if any, are required in processing odors using behavioral assays. Interestingly, DAF-6 is a Patched-related protein with expression pattern that overlaps with Ppa-obi-1 expression (amphid lumen, excretory canal, vulva lumen) and is required for chemotaxis and dauer formation (Perens and Shaham, 2005). Similarly, Ppa-obi-1 expression in the putative sheath or socket cells of the inner labial neurons may also be involved in ZTDO sensing, as recent ultrastructural electron micrograph reconstructions indicate that BAG (gas sensing) and FLP (nociceptive) neurons in C. elegans associate closely with the socket cells of the inner labial sensillae (Doroquez et al., 2014). One possible mode of action for ZTDO may be its ability to change cell membrane permeability, thereby indirectly affecting the functions of the glia-associated neurons. Hence, Ppa-OBI-1 may function to counter the cell membrane permeability changes brought about by ZTDO by interacting with cell membrane lipids. Alternatively, Ppa-OBI-1 could mediate pheromone binding directly. In lepidopterans such as the wild silk moth Antheraea polyphemus and the cotton leafworm Spodoptera littoralis, pheromone-degrading and odorant-degrading enzymes (PDEs and ODEs) are esterases specifically expressed in the male antennal sensillar lymph analogous to the amphid lumen (Ishida and Leal, 2005; Durand et al., 2011). The termination of the signal by removing odors that have already triggered a response is an important process for preventing odor adaptation and increasing odor sensitivity. Thus, it is possible that Ppa-OBI-1 acts as a solubilizing factor for the lipid pheromones in the excretion-filled amphid sheath lumen (Figure 6). The weak ability of amphid sheath-specific expression of Ppa-OBI-1 to confer ZTDO protection in C. elegans suggests that both expression pattern and protein function could be important factors in ZTDO sensing in P. pacificus. Future work will be necessary to determine if Ppa-obi-1 expression restricted to the amphid sheath is sufficient to rescue the ZTDO attraction defect in P. pacificus and if the amphid sheath cells are required to mediate ZTDO sensing.
The role of Ppa-OBI-1 in neutralizing a host nematocide
Given that Ppa-obi-1-independent susceptibility to ZTDO occurs during the J2 larval stage when Ppa-obi-1 is expressed in the seam and hypodermal cells rather than in the amphid sheath that is only active in the J4 larvae, ZTDO-induced paralysis likely acts through the extracellular matrix of the nematode cuticle. P. pacificus adults, whose cuticle may be compositionally different from the larval cuticle, are not susceptible to ZTDO. However in P. pacificus embryos, the susceptibility of ZTDO may be due to ZTDO limiting gas or ion exchange with the environment rather than direct interaction between ZTDO and cellular components, given that nematode egg shells are considered to have very limited permeability accessible only to small molecular weight solutes and gas molecules (Chitwood and Chitwood, 1974). Since ZTDO exposure does not lead to embryonic arrest and paralysis in C. elegans, ZTDO must also interact with other non-conserved P. pacificus-specific molecules that have yet to be identified. There are very few known nematocides from natural compounds. One rare example comes from the oyster mushroom Pleurotus ostreatus, which secretes droplets that can immobilize rhabditid nematodes in an hour, though its effects on embryos have not been investigated (Barron and Thorn, 1987; Stadler et al., 1994). Future work on the different susceptibility to ZTDO may help to answer if there is a wider pattern of negative correlation between ZTDO attraction and ZTDO-induced paralysis in other natural isolates, as observed in the California and Washington strains. Such antagonistic traits may reveal possible co-evolution between host finding and host tolerance in necromenic associations.
We further speculate that in the natural ecology of P. pacificus, ZTDO may enable the beetle host to limit the nematode number and to thus synchronize the developmental stage of infective P. pacificus by arresting embryogenesis and lethally paralyzing J2 larvae. The unexpected role of a P. pacificus attractant to act as a developmental regulator against the younger larval stages, particularly the dauer larvae (DL), suggests that the ZTDO-resistant adult stage is a potential dispersal stage. This attenuated antagonistic interaction also revealed that nematode larvae and adults may be susceptible to distinct selective pressures. P. pacificus DL does not disperse readily, likely due to the secretion of the wax ester nematoil that enables P. pacificus to form dauer towers when potential hosts are nearby (Penkov et al., 2014). Therefore under natural conditions, lower concentrations of ZTDO than the concentrations we have tested may serve as a close-range host dwelling strategy by inhibiting DL dispersal to form dauer aggregates. The developmental role of ZTDO may be part of a mechanism to limit nematode dispersal until the beetles mature. In these respects, the promotion of a non-DL dispersal stage and inhibition of DL dispersal by ZTDO resemble other known host-derived compounds found in the entomophilic nematode B. xylophilus and the phoretic nematode C. japonica (Okumura et al., 2013; Zhao et al., 2013). Attraction and paralysis by ZTDO may have co-evolved in the necromenic association to allow successive populations of founder P. pacificus DL and adults to infect beetles, culminating in arrested DL remaining on sexually mature beetles during the dispersal cycle. The pheromone arrested DL state may also help to account the observation that Pristionchus nematodes emerge several days later than rhabditids from the same species of host beetles. Our findings that a lipid-binding protein has a role in coordinating chemosensation, body proportions, and protection from a host pheromone encourages fresh inquiries on additional genes involved in ZTDO sensory signaling and its mechanism of action.
Materials and methods
Strains
All nematodes were raised on NGM media containing OP50 E. coli at ∼23°C as previously described (Hong and Sommer, 2006a; Hong et al., 2008a). The following strains were used: P. pacificus: California (PS312) wild type, Washington (PS1843), Ppa-obi-1(tu404)I; Ppa-egl-4(tu374)IV (null); Ppa-obi-3(tu405); csuIs01 [Ppa-obi-1p::gfp; Ppa-egl-20p::rfp; PS312 gDNA]X; rlh49(csuIs01; tu374) ‘Ppa-obi-1p::gfp; Ppa-egl-4’; rlh50(csuIs01; tu92) ‘Ppa-obi-1p::gfp; roller’; rlh51(csuIs01; tu404) ‘Ppa-obi-1p::gfp; Ppa-obi-1’. C. elegans: Bristol(N2) wild type, nrf-5(sa513), csuEx02 [C06G1.1p::gfp; Cel-myo-2::DsRED]; csuEx03 [C06G1.1p::gfp; pRF4 roller], csuEx29 [T02B11.3p::Ppa-obi-1:gfp; Cel-myo-2::DsRED], and csuEx30 [F16F9.3p::Ppa-obi-1:gfp; Cel-myo-2::DsRED]. The whole-genome sequenced strain vc20357 was outcrossed to him-5(e1467) three times and the C06G1.1 locus was sequenced to confirm that the slim body phenotype is linked to the missense gk306193 mutation (D48N).
Chemotaxis assays and isolation of obi mutants
Request a detailed protocolWe X-ray mutagenized wild-type P. pacificus PS312 (California) at 325 Roentgen for 25 min with radioactive beryllium and picked 40 Po onto individual plates. We subsequently picked 1067 F2s from 400 F1s and screened 713 viable F3 populations for insensitivity to 150 nl of oriental beetle pheromone, Z-7-tetradecen-2-one (99% pure; Bedoukian Research, CT) after 1 hr treatment with 1 mM exogenous 8-Bromo-cGMP (Sigma–Aldrich, MO) (Hong et al., 2008a). 100 mM nicotinic acid (Sigma–Aldrich, MO) was dissolved in water. Population chemotaxis assays were conducted for ∼16 hr at 23°C, and individual candidate F3 from plates showing insensitivity were rescued onto fresh plates by mouth pipetting. We repeated the cGMP-treated chemotaxis assay from the progeny of the candidate mutant lines two more times with at least two duplicate assay plates per line. Only those lines that failed the third round of chemotaxis assay were considered Oriental Beetle pheromone Insensitive obi alleles. Ppa-obi-1 maintained long body morphology and obi-3 displayed more coiled body posture after 3× outcrossing to wild-type PS312 and selecting the ZTDO insensitive phenotype. Because both the chemotaxis and morphological phenotypes co-segregate after 5× outcross, we subsequently used the long body phenotype as a visible marker for Ppa-obi-1 during positional mapping. Population chemotaxis assays were performed as previously described based on modified protocol for C. elegans (Bargmann et al., 1993; Hong and Sommer, 2006a). Well-fed J4 and adult worms were washed in M9 buffer twice before assay. For a more consistent response, C. elegans was washed in spring water rather than M9 buffer before the ZTDO avoidance assay. For the exogenous cGMP treatment, each washed population was split for mock or 1 hr treatment with 1 mM 8-Bromo-cGMP (in water). For each condition, at least 10 worms had to reach the odor sources in each assay and 12 assays (sample size N) were performed over three sessions. Chemotaxis assays with transgenic obi-1 rescue worms were scored by rfp expression in the tail.
ZTDO paralysis assays
Request a detailed protocol20 live worms or eggs from non-starved cultures were placed on NGM agar plates with fresh OP50. For consistency, eggs were synchronized by picking ∼50 adults onto OP50 and allowing them to lay eggs for 3 hr. The eggs were then transferred to different plates and counted. 10 µl of ZTDO or ETDA diluted in ethanol was added to the lid of the plate, immediately sealed with Parafilm, and incubated at room temperature. Larvae were scored 3 hr later or the next day by counting the number of paralyzed and active worms. A worm was scored as paralyzed if it did not move at least one pharynx length after tapping on the outside of the plate. Eggs were scored 2 days later by counting the number of unhatched eggs. For dauer larvae exit assays on NGM plates containing none, 0.001% or 0.01% ZTDO (vol/vol), 25 dauer larvae from the same 2- to 3-week old starved plate were picked onto each type of plate with OP50 and scored for arrested/active dauer larvae, J3 larvae, or J4 larvae after 2 days at ∼23°C.
Whole genome analysis
Request a detailed protocolA single library was paired-end sequenced twice in separate flow cell lanes with 2 × 76 bp that yielded 61 million and 65 million reads after preprocessing (quality control, adapter removal). More than 90% of all reads could be mapped to the P. pacificus genome (www.pristionchus.org) with BWA (version 0.5.9) for each library. We then employed ACCUSA for SNP detection (Froehler and Dieterich, 2010). Briefly, we performed a head-to-head comparison of the mutant Ppa-obi-1 and wild-type PS312 mapped libraries. The algorithm represents read stacks (base calls and qualities thereof) as Dirichlet distributions. For any given genome position, we require a read coverage of at least three in both samples. The smaller read stack serves as reference distribution from which the larger stack is drawn. A probability density for this draw is computed, and a SNP is called if the log-density is lower than −30.
Transgenic reporter gene constructs
Request a detailed protocolStable transgene expression in P. pacificus requires co-injection of endogenous genomic DNA and co-injection markers containing compatible cohesive ends with the target gene as previously described (Schlager et al., 2009; Cinkornpumin and Hong, 2011). A 4.3 kb Ppa-obi-1 promoter fragment was amplified from PS312 gDNA using RHL121/RHL136 (primary) and RHL124/RHL149 (secondary) primers. The Ppa-obi-1 promoter was then fused with the gfp- Ppa-rpl-23-3′UTR by PCR (Hobert, 2002). PS312 animals were injected with a DNA mix containing a final concentration of 10 ng/μl of the SpeI/SalI digested 4.3 kb Ppa-obi-1 promoter::NLS-gfp-SalI fusion PCR product from three independent reactions; 10 ng/μl SpeI digested Ppa-egl-20p::rfp linearized plasmid; and 60 ng/μl SpeI/SalI digested PS312 genomic DNA. Individual F1s expressing Ppa-egl-20p::rfp in the hermaphrodite tail were selected for Ppa-obi-1 expression analysis. Two stable transgenic lines were initially generated with similar expression patterns. We used the stably transmitted reporter line csuIs01[Ppa-obi-1p::gfp; Ppa-egl-20p::rfp; PS312 gDNA]X for detailed expression analyses. csuIs01 is likely to be the result of a spontaneous integration into the X chromosome based on the observation that transgenic males expressing the co-injection rfp marker crossed to Ppa-unc-1(ChrIV) hermaphrodites gave rise to all F1 males lacking rfp expression but all F1 hermaphrodites expressing rfp. More than 50 individuals at all developmental stages were analyzed. Ventral views of the vulva expression were obtained from csuIs01 in the prl-1 roller background. The same 4.3 kb Ppa-obi-1 promoter was used to drive the full length 1.6 kb cDNA fused to GFP by injecting into PS312 and crossing into the Ppa-obi-1(tu404) background to generate csuEx27[Ppa-obi-1p::Ppa-obi-1 cDNA:gfp; Ppa-egl-20::rfp; Ppa-obi-1(tu404) gDNA]. For unknown reasons, the N-terminally fused gfp coding region did not express GFP.
To create the C06G1.1p::gfp reporter gene construct, we PCR amplified a 3.2 kb upstream sequence from the C06G1.1 ATG using RHL169 and RHL187 and fused the promoter with GFP:Ppa-rpl-23 3′ UTR by fusion PCR, using RHL181 and RHL154 primers to obtain a 4183 bp amplicon. Independently amplified C06G1.1p::gfp fragments were co-injected with either pRF4 rol-6(su1006) or myo-2p::DsRED vectors along with the pBluescript vector as carrier DNA (50 ng/μl each). This reporter likely encompasses C06G1.1's maximum regulatory region because the 3.2 kb promoter fragment extends ∼300 bp upstream of the next gene on the opposite strand (C06G1.14), and C06G1.1 does not contain any intron larger than 1 kb that may act as potential intronic enhancers. Selection for DsRED animals in the reporter strain csuEx02 [C06G1.1p::gfp; Cel-myo-2::DsRED] results in 90% GFP+/DsRED+ progeny and equal DsRED+ expression in pharyngeal muscle cells, suggesting stable mitotic transmission of the transgene. All five F1 transgenic lines showed similar expression profile and intensity in post-hatching stages, but strong body epidermal expression was detected only in the csuEx03 [C06G1.1p::gfp; pRF4; pBST] line. Images were obtained using a Leica DM6000 fluorescent microscope and Leica SP5 confocal microscope. To create the T02B11.3p::Ppa-obi-1:gfp amphid sheath expressing translational fusion construct, an ∼2.1 kb T02B11.3 promoter fragment was amplified from C. elegans gDNA using RHL476/RHL490 (primary reaction). The Cel-T02B11.3 promoter is then fused to the same 1.6 kb Ppa-obi-1 cDNA rescue construct by PCR using RHL477/RHL483 (secondary) to generate a 3.7 kb fusion. This construct and the GFP-unc-54-3′UTR fragment, amplified from the pPD95.75 vector using RHL480/RHL481 (primary) was fused using RHL501/RHL482 (tertiary) to create the complete construct. A similar scheme was used to generate F16F9.3p::Ppa-obi-1:gfp using RHL498/RHL500 (promoter). C. elegans N2 were injected with a mixture of purified T02B11.3p::Ppa-obi-1:gfp (25 ng/μl) or F16F9.3p::Ppa-obi-1:gfp, myo-2::DsRED (25 ng/μl) and pBluescript carrier DNA (100 ng/μl). One stable line from each construct was characterized for ZTDO chemotaxis and paralysis.
The GenBank accession for Ppa-obi-1 mRNA is JQ946290. Primer sequences are listed in Table 2.
Primers
| Primer | Primer sequence (5′→3′) | Template target |
|---|---|---|
| RHL101 | TGAACGGGCTTTTAATCTGG | Ppa-obi-1 gDNA |
| RHL102 | GTGTCGAGATAGTCGGGCAT | Ppa-obi-1 gDNA |
| RHL104 | ATGACTGCTCCAAAGAAG | Ppa-obi-1 cDNA |
| RHL107 | ATTTGCCCTTCGTTCCAC | Ppa-obi-1 for RNAi |
| RHL108 | CTGTCTCAGTATGCCGAATG | Ppa-obi-1 for RNAi |
| RHL109 | ATTCAGCCGTGTTACAAATC | Ppa-obi-1 cDNA |
| RHL110 | CTGCATGTGTTCCGGTCTCG | Ppa-obi-1 cDNA |
| RHL113 | AAACCTCTGACACATGCAGC | Ppa-obi-1 cDNA |
| RHL121 | AGTTTCAAAGTTGCATGG | Ppa-obi-1 promoter |
| RHL124 | CTTCTTTGGAGCAGTCATGTAGCGATAATCAGGAGT | gfp; Ppa-obi-1 promoter |
| RHL132 | ATGACTGCTCCAAAGAAG | gfp |
| RHL136 | CAACCGTGTGCAGTAGAACC | Ppa-obi-1 5′ RACE |
| RHL149 | ACTATCCTATCATCGGAAGC | Ppa-obi-1 promoter |
| RHL151 | AGGGCATTGACAAATCATCG | Ppa-obi-1 cDNA |
| RHL154 | CACGACGTTGTAAAACGACG | Ppa-obi-1 cDNA |
| RHL169 | CGCGCTAAACCAATTCCGCC | C06G1.1 promoter |
| RHL181 | GTTGAGAGAAGAGGTGGAGTCC | C06G1.1 promoter |
| RHL187 | CTTCTTTGGAGCAGTCATCAGATCTGAAAATTTGACACATTG | gfp; C06G1.1 promoter |
| RHL267 | GGAACTAACATTAATGTCAC | C06G1.1 RNAi |
| RHL268 | GAGAAGTGCATAGTTGATG | C06G1.1 RNAi |
| RHL477 | GCGCTACAGGGGATTTTGTC | T02B11.3 promoter |
| RHL480 | TGAAAGATGCAAGTAAAGGAGAAGAACTTTTC | p95.75 GFP |
| RHL481 | AAGGGCCCGTACGGCCGACTA | p95.75 GFP |
| RHL482 | GTAGGAAACAGTTATGTTTGG | p95.75 GFP |
| RHL484 | CTTCAGAAAATGATCAGGAGTCTGGTTC | obi-1 cDNA |
| RHL487 | AGTGAAAAGTTCTTCTCCTTTACTTGCATCTTTCACGGAGTC | obi-1 cDNA; gfp |
| RHL490 | AACGAGAACCAGACTCCTGATCATTTTCTGAAGAAAGTTGAAAAAC | T02B11.3 promoter; obi-1 cDNA |
| RHL498 | GCGATAAGATCGGTCAATCTGAG | F16F9.3 promoter |
| RHL499 | GCCAGTAAGGGCTAGTAAGTG | F16F9.3 promoter |
| RHL500 | AACGAGAACCAGACTCCTGATCATATTTTGTTTCTTACTGTCTTG | F16F9.3 promoter |
| RHL501 | ATGGGTACGTTTCTGGGTATAG | T02B11.3 promoter; obi-1 cDNA |
| VR86 | GACTCCTGATTATCGCTACG | Ppa-obi-1 cDNA |
| VR87 | AGATAGTCAAACTATGCATC | Ppa-obi-1 cDNA |
| VR88 | CTAATAATCTACACTAAATG | Ppa-obi-1 cDNA |
| AG11112 | CTCGGAGGAGGAACTGGATC | Ppa-beta-tubulin RT-qPCR |
| AG11113 | GACCGTGTCAGAGACCTTAG | Ppa-beta-tubulin RT-qPCR |
| RH12548 | GCAGGAACATATATTCGCAG | Ppa-egl-4 RT-qPCR |
| RH12549 | TGTCACGGAACGTTTTGTAC | Ppa-egl-4 RT-qPCR |
| SL1 | GGTTTAATTACCCAAGTTTGAG | SL1 for 5′ RACE |
| SSLP21F | ACTGGTGCTCATCGCAAGA | Mapping marker |
| SSLP21R | CCTTCTGTTTTCCTACCCCC | Mapping marker |
| SSLP7F | TCTTGCATAACACCGAACAAA | Mapping marker |
| SSLP7R | AGGGCGTTACGTGAATAAGC | Mapping marker |
| SSLP17F | CAATGCAGTAGCCGAATCAT | Mapping marker |
| SSLP17R | CAATATTGGCCTTCACCCTG | Mapping marker |
| S447F | TGGGGATAGCGAAAATCATC | Mapping marker |
| S447R | CGCATCGTTATTCACGAAATTC | Mapping marker |
-
Letters in bold are based on different templates for fusion PCR.
Egg count and body length measurements
Request a detailed protocolBody measurements were obtained as previously described (Hong et al., 2008a). In brief, we used a Leica DM6000 and LAS application to measure body dimensions at 24-hr intervals starting at mid-J4/L4 stage. To determine hatching rate, we picked single J4-stage PS312 hermaphrodites onto fresh OP50 plates and transferred the nematodes every 24 hr three times at 20°C. The total number of eggs present on the plates following each transfer and the resulting number of 3-day old animals for each worm during the 3-day period were used to calculate the percentage of hatching and brood size.
RNA interference
Request a detailed protocolThe bacterial feeding strain containing the C06G1.1 fragment 1 was previously described in a large-scale C. elegans RNAi screen (Simmer et al., 2003). C06G1.1 fragment 2 was made using RHL267 and RHL268 primers to amplify an ∼400 bp fragment of C06G1.1, digesting this product with BglII and PstI, and ligating the 190 bp fragment into the BglII/PstI sites of pL4440. In vitro synthesis of dsRNA from 1 μg of T7 amplified PCR template from the RNAi vector or the pL4440 negative control was accomplished using the Ambion in vitro synthesis kit (Grand Island, NY). RNAi by bacterial feeding of C06G1.1 was performed by scoring the progeny of L4 stage rrf-3(pk1426) animals raised on RNAi or L4440 control plates containing 1 mM IPTG and 50 μg/ml carbenicillin.
Statistical analysis
Request a detailed protocolMeans, SEM (error bars), and p values for two-tailed t test were performed by Microsoft Excel. One-way ANOVA with multiple comparisons post-hoc testing were performed using Prism statistical software. Sample sizes represent total technical replicates from two or more biological replicates.
Data availability
-
A Lipid-binding Protein in the Necromenic Nematode Pristionchus pacificus is Involved in Insect Pheromone SensingID JQ946290. Publicly available at GenBank (http://www.ncbi.nlm.nih.gov/genbank).
-
WormBase 2012: more genomes, more data, new websiteID C06G1.1. Publicly available at WormBase (http://www.wormbase.org/).
-
Draft genome of the filarial nematode parasite Brugia malayiID XP_001897990.1. Publicly available at GenBank (http://www.ncbi.nlm.nih.gov/genbank).
References
-
Destruction of nematodes by species of PleurotusCanadian Journal of Botany 65:774–778.https://doi.org/10.1139/b87-103
-
Host-finding behaviour in the nematode Pristionchus pacificusProceedings. Biological Sciences/ The Royal Society 278:3260–3269.https://doi.org/10.1098/rspb.2011.0129
-
RNAi mediated gene knockdown and transgenesis by microinjection in the necromenic Nematode Pristionchus pacificusJournal of Visualized Experiments e3270.https://doi.org/10.3791/3270
-
egl-4 acts through a transforming growth factor-beta/SMAD pathway in Caenorhabditis elegans to regulate multiple neuronal circuits in response to sensory cuesGenetics 156:123–141.
-
ACCUSA–accurate SNP calling on draft genomesBioinformatics 26:1364–1365.https://doi.org/10.1093/bioinformatics/btq138
-
Plasma PLTP (phospholipid-transfer protein): an emerging role in “reverse lipopolysaccharide transport” and innate immunityBiochemical Society Transactions 39:984–988.https://doi.org/10.1042/BST0390984
-
PCR fusion-based approach to create reporter gene constructs for expression analysis in transgenic C. elegansBiotechniques 32:728–730.
-
Pristionchus pacificus: a well-rounded nematodeBioessays 28:651–659.https://doi.org/10.1002/bies.20404
-
Natural variation in Pristionchus pacificus insect pheromone attraction involves the protein kinase EGL-4Proceedings of the National Academy of Sciences of USA 105:7779–7784.https://doi.org/10.1073/pnas.0708406105
-
Species-specific recognition of beetle cues by the nematode Pristionchus maupasiEvolution & Development 10:273–279.https://doi.org/10.1111/j.1525-142X.2008.00236.x
-
Rapid inactivation of a moth pheromoneProceedings of the National Academy of Sciences of USA 102:14075–14079.https://doi.org/10.1073/pnas.0505340102
-
Cholesteryl ester transfer protein and phospholipid transfer protein have nonoverlapping functions in vivoThe Journal of Biological Chemistry 275:29477–29481.https://doi.org/10.1074/jbc.M003523200
-
The Caenorhabditis elegans rol-6 gene, which interacts with the sqt-1 Collagen gene to determine organism morphology, Encodes a CollagenMolecular and Cellular Biology 10:2081–2089.
-
Natural variation in chemosensation: lessons from an island nematodeEcology and Evolution 3:5209–5224.https://doi.org/10.1002/ece3.902
-
Species-specific recognition of the carrier insect by dauer larvae of the nematode Caenorhabditis japonicaJournal of Experimental Biology 216:568–572.https://doi.org/10.1242/jeb.073593
-
A wax ester promotes collective host finding in the nematode Pristionchus pacificusNature Chemical Biology 10:281–285.https://doi.org/10.1038/nchembio.1460
-
Glia-neuron interactions in the nervous system of Caenorhabditis elegansCurrent Opinion in Neurobiology 16:522–528.https://doi.org/10.1016/j.conb.2006.08.001
-
BookThe geographical mosaic of coevolutionChicago: The University of Chicago Press.
-
Lipopolysaccharide (LPS)-binding proteins BPI and LBP form different types of complexes with LPSThe Journal of Biological Chemistry 272:18682–18685.https://doi.org/10.1074/jbc.272.30.18682
-
Purification and characterization of a potent bactericidal and membrane active protein from the granules of human polymorphonuclear leukocytesThe Journal of Biological Chemistry 253:2664–2672.
-
Identification and synthesis of female sex pheromone of oriental beetle, Anomala orientalis (Coleoptera: Scarabaeidae)Journal of Chemical Ecology 20:2415–2427.https://doi.org/10.1007/BF02033210
Article and author information
Author details
Funding
National Institute of General Medical Sciences (SC2GM089602)
- Ray L Hong
Center for California Studies, California State University (Research and Scholarship Award)
- Ray L Hong
W.M. Keck Foundation
- James L Go
National Institutes of Health (1SC3GM105579)
- Ray L Hong
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
RLH is supported by the NIH grant SC2GM089602, 1SC3GM105579, and the CSUN Research and Scholarship award. JLG was supported by the Keck Foundation. The Caenorhabditis Genetics Center (CGC) provided the C. elegans strains. We thank J Salandanan, E Bernal, J Escobedo, and G Aguilar-Portillo for technical assistance, the Shaham lab for plasmids, and J Srinivasan and P Sternberg for help with genome sequencing.
Copyright
© 2014, Cinkornpumin et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 3,468
- views
-
- 205
- downloads
-
- 43
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 43
- citations for umbrella DOI https://doi.org/10.7554/eLife.03229
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
A beetle pheromone that lures nematode worms to an insect host can also stop their development or even kill them outright.





