GATA6 mutations in hiPSCs inform mechanisms for maldevelopment of the heart, pancreas, and diaphragm
Abstract
Damaging GATA6 variants cause cardiac outflow tract defects, sometimes with pancreatic and diaphragmic malformations. To define molecular mechanisms for these diverse developmental defects, we studied transcriptional and epigenetic responses to GATA6 loss of function (LoF) and missense variants during cardiomyocyte differentiation of isogenic human induced pluripotent stem cells. We show that GATA6 is a pioneer factor in cardiac development, regulating SMYD1 that activates HAND2, and KDR that with HAND2 orchestrates outflow tract formation. LoF variants perturbed cardiac genes and also endoderm lineage genes that direct PDX1 expression and pancreatic development. Remarkably, an exon 4 GATA6 missense variant, highly associated with extra-cardiac malformations, caused ectopic pioneer activities, profoundly diminishing GATA4, FOXA1/2, and PDX1 expression and increasing normal retinoic acid signaling that promotes diaphragm development. These aberrant epigenetic and transcriptional signatures illuminate the molecular mechanisms for cardiovascular malformations, pancreas and diaphragm dysgenesis that arise in patients with distinct GATA6 variants.
Introduction
Congenital heart disease (CHD), the leading birth defect worldwide that occurs in approximately 1% of newborns (van der Linde et al., 2011), comprises a range of structural malformations arising during embryonic development (Zaidi and Brueckner, 2017). Contemporary treatments for CHD enables survival into adulthood, but many patients have ongoing medical problems related to extra-cardiac malformations and/or neurodevelopmental disabilities (Brickner et al., 2000; Marino et al., 2012). Understanding the developmental mechanisms that cause co-occurrence of these congenital anomalies is expected to provide better patient care, improve predictive risk assessments, and potentially uncover therapeutic targets.
Heart development is an intricately regulated process driven by genetic, epigenetic, and biomechanical events that form a complex, multi-chambered, muscular organ containing cardiomyocytes and other cell lineages (Litviňuková et al., 2020). Early in cardiogenesis, the first heart field creates the left ventricle and portions of the atria, while the second heart field gives rise to the right ventricle and outflow tract, with contributions from neural crest cells (Buckingham et al., 2005). Multiple transcription factors orchestrate particular components of these developmental processes (Srivastava and Olson, 2000), as evidenced by regional defects in CHD patients with damaging mutations in particular genes (Fahed et al., 2013; Jin et al., 2017; Sifrim et al., 2016). For example, dominant human loss of function (LoF) variants in TBX5, NKX2-5, and GATA4 that are highly expressed in the first and second heart fields cause atrial and ventricular septal defects (Basson et al., 1997; Benson et al., 1999; Garg et al., 2003). GATA6, while expressed in the first heart field (Morrisey et al., 1996), plays particularly critical roles in the developing second heart field (Molkentin, 2000) and in recruitment of cardiac neural crest lineages (Lepore et al., 2006) that together shape the cardiac outflow tract. Consistent with these developmental functions, CHD patients with GATA6 mutations have a striking preponderance of outflow tract malformations (Gharibeh et al., 2018; Kodo et al., 2009; Maitra et al., 2010). GATA6 also is critical for endodermal development (Fisher et al., 2017), and some CHD patients with damaging GATA6 variants also have pancreatic agenesis, congenital diaphragmatic hernia (Yu et al., 2014) or other abdominal malformations (Chao et al., 2015; De Franco et al., 2013; Shi et al., 2017).
Developmental transcription factors work in concert to exert network-level effects on organogenesis (Luna-Zurita et al., 2016): physical interactions of NKX2-5, TBX5, and GATA4 proteins modulate the expression of other cardiac genes (Garg et al., 2003; Hiroi et al., 2001; Bruneau et al., 2001; Maitra et al., 2009) while GATA6 and GATA4 proteins together promote pancreatic development (Chao et al., 2015; De Franco et al., 2013; Shi et al., 2017). Notably, members of the GATA family of transcription factors serve as endodermal pioneer factors that participate with FOXA, also a pioneer factor, to engage and open chromatin and recruit additional transcriptional activators (Fisher et al., 2017; Zaret and Carroll, 2011). Thus, we expect that understanding the molecular networks in which GATA6 participates will help to elucidate the mechanisms by which human mutations cause defects in morphogenesis of the heart cardiac other organs.
A traditional approach for studying developmental mechanisms relies on gene disruption in model organisms. While often informative, prior studies of mice with one inactivated Gata6 allele have either subtle dysmorphic aortic valves (Gharibeh et al., 2018) or no CHD (Lepore et al., 2006) and notably lack pancreatic agenesis or congenital diaphragmatic hernia that occur in human CHD patients with damaging GATA6 variants (Chao et al., 2015; De Franco et al., 2013; Shi et al., 2017). Mice with biallelic Gata6 inactivation have deficits in early visceral endoderm formation resulting in nonviable embryos (Morrisey et al., 1996; Zhao et al., 2005), which may explain the absence of homozygous GATA6 null alleles in humans, but provide limited insights into organ morphogenesis. The introduction of human CHD mutations using CRISPR/Cas9 (De Franco et al., 2013) into isogenic human induced pluripotent stem cell-derived cardiomyocytes (hiPSC-CMs) provides an alternative model system suited for analyzing developmental consequences, as differentiation of these cells activate transcriptional networks that regulate early in vivo human cardiogenesis (DeLaughter et al., 2016; Li et al., 2016). Moreover, isogenic hiPSC-CMs containing distinct variants can illuminate variant-specific transcriptional patterns.
We employed this approach to study human de novo GATA6 variants, identified by whole-exome sequencing (WES) of CHD patients. We demonstrate the graded transcriptional effects of GATA6 heterozygous and homozygous LoF variants during hiPSC-CM differentiation. We also examine the transcriptional requirements for a specific arginine residue encoded by GATA6 exon four that is recurrently mutated in unrelated CHD patients. Combining these data with Assay for transposase-accessible chromatin using sequencing (ATAC-seq) (Buenrostro et al., 2015; Corces et al., 2017) and GATA6 ChIP-seq analyses, we demonstrate that GATA6 is a pioneer factor for cardiac development. We define direct and indirect transcriptional responses to GATA6 variants. Integrating these datasets with clinical phenotypes observed in CHD patients with pathogenic GATA6 variants, we demonstrate how disrupted molecular programs cause aberrant development of the cardiac outflow tract, pancreas, and diaphragm.
Results
Identification of CHD patients with GATA6 LoF and missense variants
Among >4000 CHD patients enrolled and studied by WES through National Heart, Lung, and Blood Institute’s Pediatric Cardiac Genomics Consortium (PCGC) (Homsy et al., 2015; Jin et al., 2017), we identified nine heterozygous de novo variants: four LoF (LoF) and five damaging missense variants in GATA6 (Figure 1A). The congenital anomalies in these patients were consistent with previously recognized roles for GATA6 in developing the cardiac outflow tract, arterial-ventricular valves, and posterior brachial arches that form aorta and pulmonary vessels (Gharibeh et al., 2018; Laforest and Nemer, 2011; Losa et al., 2017). Among PCGC CHD patients and 61 previously reported CHD patients with pathogenic GATA6 variants (Figure 1B) there were a preponderance of outflow tract malformations, including persistent truncus arteriosus, double-outlet right ventricle, tetralogy of Fallot, as well as aortic and pulmonary valve and septation defects (Kelly, 2012). Some but not all of these patients also had extra-cardiac phenotypes including pancreatic agenesis, congenital diaphragmatic hernia, and neurodevelopmental deficits.
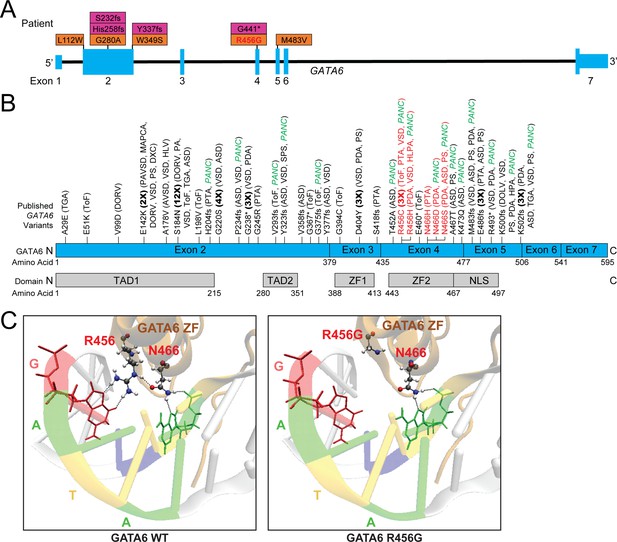
Genetic information and clinical phenotypes of individuals with GATA6 variants.
A) Schematic of GATA6 gene and locations of PCGC GATA6 variants (LoF-purple, missense-orange). (B) Previously described (see Supplementary file 1) GATA6 variants (R456 and N466 variants highlighted in red), and GATA6 protein domains. TAD: Topologically associating domain, ZF: zinc finger, NLS: nuclear localization signal. TGA: Transposition of the Great Arteries, ToF: Tetralogy of Fallot, DORV: Double-Outlet Right Ventricle, DOLV: Double-Outlet Left Ventricle, DXC: Dextrocardia, VSD: Ventricular Septal Defect, HLV: Hypoplastic Left Ventricle, PA: Pulmonary Atresia, ASD: Atrial Septal Defect, PTA: Persistent Truncus Arteriosus, SPS: Supravalvular Pulmonary Stenosis, HLPA: Hypoplastic Left Pulmonary Artery, PS: Pulmonary Stenosis, HPA: Hypoplastic Pulmonary Artery, PANC: Pancreatic Agenesis (C) Model of GATA6 DNA-binding domain bound to major groove of DNA indicating the location of amino acid residue 456. Left panel: GATA6 residues R456 and N466 normally interact with each other via hydrogen bonding (dashed lines) and with target G base and second A base in the GATA motif via hydrogen bonding, respectively (dashed lines). Right panel: Replacing the arginine (R) residue at position 456 with a glycine (G) residue alters normal molecular interactions by disrupting the hydrogen bonds.
We considered whether the distribution of these 70 damaging GATA6 variants (61 published, 9 PCGC) across the 595 encoded GATA6 amino acids correlated with clinical phenotypes (Figure 1; Supplementary file 1A, B). All variants (43 missense, 27 LoF, including eight recurrent variants) caused CHD. Extra-cardiac phenotypes occurred in 29/70 (41%) patients, and more often with LoF (18/27) than missense (11/43) variants (p<0.001). Neurocognitive dysfunction occurred in 13 patients (18.5%). Pancreatic agenesis/hypoplasia or congenital diaphragmatic hernia occurred in 20/70 patients (28.5%) and more frequently with GATA6 LoF (14/27) than missense (9/43) variants (p=0.001).
Among 43 GATA6 missense variants, 11 variants altered residues in the DNA-binding zinc finger (ZF) domain encoded by exon 4 (amino acids 435–477), significantly more than expected by chance (p=0.0004). Nine of these 11 patients with exon four missense variants had pancreatic agenesis or congenital diaphragmatic hernia, but none of 32 patients with missense variants located elsewhere (p=1.1e-6). Within exon 4, recurrent missense mutations altered the basic arginine residue 456. Computational modeling of this domain (PyMOL software) positioned residue 456 alongside a polar residue (asparagine 466) in close proximity to DNA (Bates et al., 2008; Figure 1C). Substitution of a non-polar glycine at residue 456 (R456G) is predicted to disrupt these interactions, and potentially alter GATA6 binding to DNA.
Generation of GATA6 LoF and GATA6R456G/R456G hiPSCs using CRISPR/Cas9
We created GATA6 LoF hiPSCs using two independent guide RNAs (gRNAs) targeting exon 2 (Figure 2—figure supplement 1A) that were transfected with Cas9 endonuclease into an early passage healthy hiPSC line PGP1 (Lee et al., 2009). Targeted hiPSCs were subcloned and GATA6 variants were confirmed by next-generation and Sanger sequencing of PCR-amplified fragments (METHODS). Four independent mutant hiPSC lines were obtained: two carry a heterozygous 1 bp insertion (GATA6+/-, chr18:19,752,124–19,752,124, A:TA) and two have a homozygous 1 bp deletion (GATA6-/-, chr18:19,752,123–19,752,124, TA:T). Using similar strategies, we transfected a gRNA targeting exon four with a single-stranded DNA oligonucleotide to serve as a template for homology-directed repair and generated two hiPSC lines with homozygous missense variant R456G (GATA6R456G/R456G; Figure 2—figure supplement 1B). No lines with a heterozygous R456G variant were obtained. In parallel we produced GATA6+/-, GATA6-/-, and GATA6R456G/R456G variants in a PGP1 hiPSC line that carried expressed green fluorescent protein (GFP) fused to endogenous cardiac troponin T alleles (TNNT2-GFP; Figure 2—figure supplement 1C).
Differentiation of GATA6 mutant hiPSCs into hiPSC-CMs
Wildtype (WT), GATA6+/-, GATA6-/-, and GATA6R456G/R456G hiPSCs were processed for differentiation into cardiomyocytes (hiPSC-CMs) by modulation of the Wnt signaling pathway and subsequent metabolic selection via glucose deprivation (Sharma et al., 2018b). This protocol yields hiPSC-CMs that express first and second heart field genes (Zhang et al., 2019). As previous single-cell RNA- sequencing (RNA-Seq) of cardiomyocytes isolated from developing mouse hearts (DeLaughter et al., 2016) identified peak Gata6 expression in cardiac progenitors and early cardiomyocytes, we studied hiPSC-CMs at differentiation days 4 and 8, which approximate these in vivo developmental stages. GATA6 protein expression and nuclear localization were reduced in GATA6+/- compared to isogenic WT lines and absent in GATA6-/- lines (Figure 2—figure supplement 1D,E). GATA6+/- hiPSCs, unlike WT hiPSCs, exhibited mono-allelic GATA6 expression, suggesting nonsense-mediated decay of RNAs transcribed from targeted alleles.
We assessed sarcomere production as a measure of cardiomyocyte differentiation in the TNNT2-GFP lines. GATA6+/- hiPSC-CMs had weaker fluorescent signal than WT cells, while GATA6-/- lines expressed no fluorescence at baseline or during the differentiation protocol (Figure 2A,B). Consistent with these data, at differentiation day eight when contracting sarcomeres were present in WT cells, fewer independent differentiation rounds of GATA6+/- hiPSC-CMs contained beating sarcomeres and no GATA6-/- cells showed sarcomeres or spontaneous beating (Figure 2C). Analogous studies of GATA6R456G/R456G hiPSC-CMs (Figure 2D–F) showed a fluorescence signal comparable to that of GATA6+/- hiPSC-CMs and spontaneous beating cells in approximately ~60% of differentiation rounds.
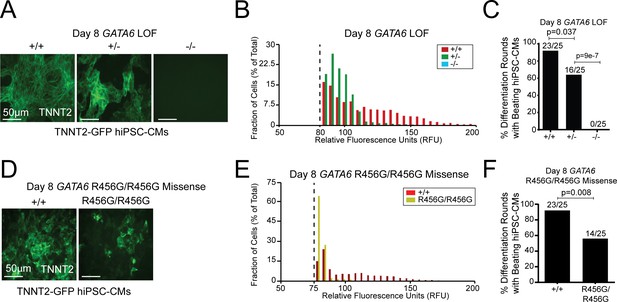
GATA6 mutant hiPSCs exhibit hiPSC-CM differentiation defects.
(A) GATA6 variants in a TNNT2-GFP reporter line showed reduced (GATA6+/-) or absent (GATA6-/-) GFP-tagged sarcomeres in comparison to WT cells. (B) Distribution of dissociated GFP-TNNT2-GATA6 mutant cells assessed using the Countess system with a GFP filter cube (METHODS). (C) Number of GATA6 LoF mutant differentiation cultures (n = 25 per genotype) with beating hiPSC-CMs. (D) The GATA6R456G/R456G variant has reduced expression of GFP-tagged sarcomeres. (E) Fluorescence distribution of differentiated GFP-TNNT2 GATA6R456G/R456G cells assessed using the Countess system with a GFP filter cube (see Materials and methods). (F) Number of GFP-TNNT2 GATA6R456G/R456G differentiation cultures (n = 25) with beating day eight hiPSC-CMs. All lines were studied at differentiation day 8. Significance was assessed using Student’s t-test.
Transcriptional analysis of GATA6 LoF hiPSC-CMs
We assessed transcriptional responses to altered GATA6 levels throughout hiPSC-CM differentiation by bulk RNA-Seq analyses of cultures at days 0, 4, 8, 12, and 30 (Figure 3, Figure 2—figure supplement 1F, Figure 3—figure supplement 1, Supplementary file 2), and by single-cell RNA-Seq on days 4 and 8 (Figure 4) that precedes metabolic selection and enriches cultures for cardiomyocytes. All cells were differentiated, processed, and harvested in parallel. RNA-Seq data was aligned and processed to limit potential batch effects, and clustered using methods implemented in DESEQ2 (bulk) or Seurat (single cell) (METHODS). We observed consistency between bulk and single-cell expression data. In addition, principal component analyses (PCA) (Figure 3A, Supplementary file 3) of independent, genotype-identical lines demonstrated close clustering of RNA expression, indicating that GATA6 genotype and differentiation stage largely accounted for differences in gene expression.
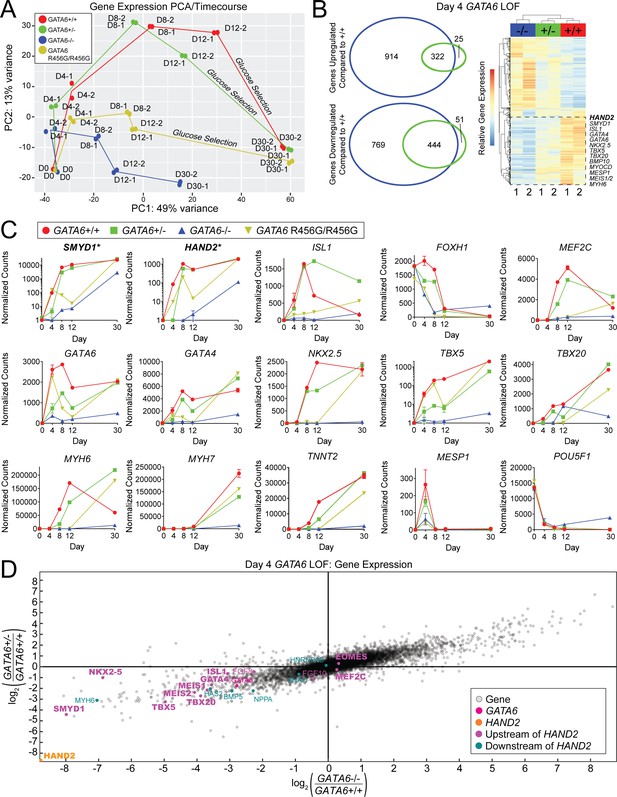
GATA6 mutant cells exhibit downregulation of second heart field-related genes during hiPSC-CM differentiation.
A) Gene expression principal component analysis (PCA) of day 0–30 WT (GATA6+/+), GATA6+/-, GATA6-/-, and GATA6R456G/R456G hiPSC-CMs. RNA-Seq samples were harvested in duplicate for all time points. (B) Venn Diagrams (left) and heat map (right) of day 4 GATA6+/- and GATA6-/- cells. In the heatmap, red indicates upregulated genes whereas blue represents downregulated genes. Samples are in duplicate. Selected second heart field genes are shown. (C) Expression data in normalized counts for second heart field-related genes (top row), cardiac developmental transcription factors (middle row), sarcomere, and other selected genes (bottom row) during differentiation of GATA6 mutant hiPSC-CMs. Data represented as mean ± SD. Note that SMYD1 and HAND2 graphs are plotted with logarithmic scale. (D) Gene expression scatterplot illustrating downregulation of expression of HAND2 upstream and downstream gene network in day 4 GATA6+/- and GATA6-/- cells. X-axis, log2 fold-change of gene expression in GATA6-/- cells relative to WT. Y-axis, log2-fold-change of gene expression in GATA6+/- cells relative to WT. Canonical cardiac development and the second heart field genes are bolded.
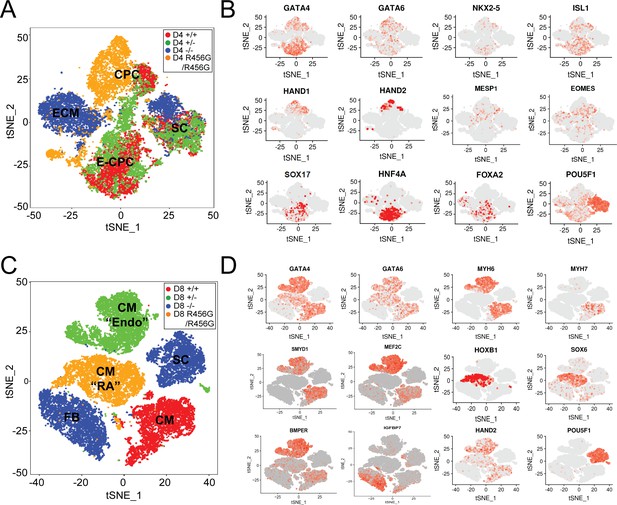
Single-cell transcriptional analysis of GATA6 mutant hiPSC-CMs during differentiation.
(A) tSNE of single-cell RNA-Seq of day 4 GATA6 mutant hiPSC-CMs identified four clusters. Labels reflect marker gene expression: SC, Stem cell; CPC, cardiac progenitor cells; E-CPC, cardiac progenitors enriched with endoderm markers; ECM, endodermal-like cells enriched for extracellular matrix proteins. (B) Examples of marker gene expression in clustered hiPSC-derived cells. CPC cells express mesodermal factors (MESP1, EOMES) as well as cardiac transcription factors (GATA6, GATA4, ISL1, NKX2.5, HAND1, and HAND2). SCs expressed OCT4 (POU5F1). E-CPCs expressed GATA6, SOX17, HNF4A, and FOXA2. (C) tSNE clustering of single-cell RNA-Seq of day 8 GATA6 mutant hiPSC-CMs identified five clusters. Labels reflect marker gene expression: CM, cardiomyocytes; CM (‘RA’), cardiomyocytes with increased RA-signaling pathway genes; SC, Stem Cell; CM (‘Endo’), cardiomyocytes with enrichment in endothelial genes; FB; fibroblast-like cells (D) Examples of marker gene expression in clustered cells. SC expressed OCT4 (POU5F1). CMs expressed sarcomere protein genes (MYH6, MYH7), SMYD1, a CM-specific histone methyl-transferase, and HAND2, a second heart field transcription factor. CMs (RA) also expressed retinoic acid pathway genes (SOX6, HOXB1). CMs (Endo) have upregulated endothelial cell gene expression (MEF2C, BMPER), while FB cells expressed ECM markers (IGFBP7).
At day 4 of differentiation, RNA-Seq of WT cells showed expression of both pluripotent stem cell markers (e.g., POU5F1 encoding OCT4) and transcriptional modulators associated with early cardiomyocyte differentiation (Figures 3B,C and 4A,B). These primordial cardiomyocytes expressed SMYD1 (SET and MYND domain-containing protein-1), a nuclear histone methyl-transferase involved in remodeling chromatin and sarcomere assembly (Li et al., 2011), Tbox transcription factors TBX5 and TBX20, two of the earliest markers of cardiac development (Bruneau et al., 2001; Takeuchi et al., 2005), and MEF2C, an essential transcription factor for sarcomere assembly and function (Lin et al., 1997). From day eight onward, the expression of stem cell marker genes was extinguished while expression of cardiomyocyte differentiation transcripts increased (Figures 3C and 4C,D).
Our protocol for cardiomyocyte differentiation also yielded subpopulations of cells expressing endodermal genes. Day 4 WT cells expressed hepatocyte nuclear factor family members (HNF1, HNF4A, HNF4B) that are depleted in Gata6-null mice (Morrisey et al., 1998), FOXA1 and FOXA2, transcription factors that activate expression of the pancreas/duodenum homeobox-1 gene PDX1, which is essential for pancreas development and β islet cell differentiation (Gao et al., 2008; Gerrish et al., 2001; Lee et al., 2019; Zhou et al., 2008) and other endodermal markers such as SOX17 (Wang et al., 2011; Figure 4A,B and Supplementary file 2). PDX1 was expressed through differentiation day 12 in WT lines, but not after metabolic enrichment for cardiomyocytes.
GATA6+/- cells at day 4 of differentiation had lower transcript levels of primordial cardiomyocyte genes compared to WT cells (Figure 3B,C) and gene ontology (GO) analyses inferred that pathways involved in cardiac muscle contraction, development, and chamber organization were reduced. Notably, the expression of SMYD1, which regulates the expression of HAND2, a critical transcriptional regulator of the second heart field (Gottlieb et al., 2002; Laurent et al., 2017) that gives rise to the outflow tract, was strikingly diminished (Figure 3B–D, Supplementary file 2). Similarly, expression of multiple HAND2 network genes was also lower than in WT cells (Figure 3C,D), including GATA gene family members, ISL1, MEF2C, and prototypic cardiac transcription factors MEIS-1, NKX2-5, TBX5, and TBX20.
Prompted by the transcriptional signatures in GATA6+/- cells, we compared the heart malformations in 54 PCGC patients with damaging variants in 11 HAND2-network and second heart field genes (Supplementary file 1C) with patients who have GATA6 mutations. Forty of these 54 patients also had outflow tract malformations, including all (n = 5) CHD patients with damaging de novo variants in TBX20.
GATA6+/- cells also identified dysregulation of other gene programs involved in forming the outflow tract. The transient peak expression in WT cells of HOXA1, HOXB1, and MSX1, which regulate the induction and expression of critical molecules involved in specifying neural crest cell development (Makki and Capecchi, 2011; Simões-Costa and Bronner, 2015; Tvrdik and Capecchi, 2006), remained low throughout differentiation of GATA6+/- cells. Notably, HOXB1 also participates in specifying endoderm destined for pancreatic and other abdominal cell lineages (Huang et al., 2002). Expression was also diminished of key molecules that couple outflow tract myocardium with developing vascular beds, including KDR (encoding vascular endothelial growth factor (VEGF) receptor-2) and the VEGF co-receptor, NRPI (Supplementary file 2). Human mutations in HOXA1 (Tischfield et al., 2005) and KDR (Reuter et al., 2019) alleles cause cardiac outflow tract malformations. Our data infers a critical role for GATA6 in orchestrating SMYD1-HAND2, VEGF, and neural crest network interactions during outflow tract morphogenesis.
Day 8 GATA6+/- cells expressed many cardiomyocyte transcripts at levels found in WT lines, but single-cell RNA-Seq analyses demonstrated some persistent differences including reduced expression of MYH7, a defining transcript of mature cardiomyocytes (Figure 3C, Figure 4C,D and Figure 2—figure supplement 1F). Genes encoding other sarcomere proteins were reduced through day 12 in GATA6+/- cells and abnormalities remained after metabolic enrichment for cardiomyocytes (Supplementary file 2). Compared to WT, day 30 GATA6+/- cardiomyocytes had 10-fold higher expression levels of ISL1 and 4-fold higher levels of fetal myosin transcripts (MYH6), suggesting developmental immaturity of GATA6 mutant cardiomyocytes.
Endodermal lineage genes were variably misexpressed in differentiating GATA6+/- cells. In comparison to WT cells, transcript levels were lower for GATA4, HNF1, and HNF4A but normal for FOXA1 and FOXA2 (Supplementary file 2). The day 12 peak expression of PDX1 in WT cells was absent in GATA6+/- cells.
GATA6+/- cells also misexpressed transcription factors genes involved in diaphragm development. In day 8 cells transcripts encoding NR2F2 (encoding a retinoic acid (RA) responsive transcription factor) were 2 to 3-fold higher while expression of ZFPM2 (encoding a family member of the Friend of GATA (FOG) transcription factors) was half of WT levels. As damaging variants in these genes cause congenital diaphragmatic hernia (Kardon et al., 2017), these data imply that GATA6 haploinsufficiency causes diaphragmatic dysgenesis by disrupting NR2F2 and ZFPM2 gene programs (Supplementary file 2).
GATA6-/- hiPSC had profound deficits in cardiomyocyte differentiation (Figure 3, Figure 2—figure supplement 1F, Supplementary file 2). Transcripts associated with early mesoderm specification including MESP1 and transcription factors associated with primordial cardiomyocytes were more depressed than in GATA6+/- cells. These findings support prior observations that shRNA knockdown of GATA6 reduces hiPSC-CM differentiation capacity (Yoon et al., 2018). Given the absence of emerging cardiomyocytes, metabolic selection to further enrich for this lineage was not performed.
The expression of endodermal genes in GATA6-/- hiPSCs was also abnormal. Throughout differentiation GATA4 transcripts were 10-fold reduced, FOXA1 and FOXA2 were transiently expressed only at day 4, and HNF4A and PDX1 expression were extinguished (Supplementary file 2). Dysregulated expression of NR2F2 (increased) and ZFPM2 (decreased) was also observed.
Single-cell RNA-Seq (Figure 4C,D) identified two GATA6-/- populations. One population maintained high expression of the stem cell (SC) marker POU5F1 throughout differentiation day 8, while the other (FB) had increased expression of fibroblast markers (COL3A1, IGFBP7), epithelial to mesenchymal transition markers (SNAIL1/2, MMP2, VIM) (Figure 3—figure supplement 1A), and neural differentiation genes (SHH, ZEB1/2, NCAM2) (Figure 3—figure supplement 1A,B). Together these data inferred that GATA6-/- cells, lacking the normal signals involved in specifying cardiomyocyte and endoderm lineages, adopted alternative differentiation programs.
Transcriptional analysis of GATA6R456G/R456G hiPSC-CMs
Parallel analyses of isogenic GATA6R456G/R456G lines showed shared and distinct transcriptional profiles from GATA6+/- or GATA6-/- cells. At days 4–12, primordial cardiomyocyte transcripts (GATA4, HAND2, and SMYD1) in GATA6R456G/R456G lines were expressed at levels midway between GATA6+/- and GATA6-/- cells, but markedly below levels in WT cells (Figures 3C and 4, and Supplementary file 2).
Distinctive transcription profiles in differentiating GATA6R456G/R456G cells (days 4–12) suggested aberrant RA signaling (Figure 4C,D, Supplementary file 2). ALDH1A2 transcripts (encoding the enzyme that converts RA from retinaldehyde) were 50-fold higher than WT cells and the expression of RARA, RARB (RA receptors A and B), and STRA6 (receptor for retinol uptake) were increased 2 to 3-fold. Transcripts encoding targets of RA signaling were also increased. HOXB1, which contains two RA-responsive elements, including one 6.5 kb 3’ of coding sequences that is critical for foregut expression (Huang et al., 2002), was 100-fold increased at differentiation day 4 – a marked contrast to the depressed levels observed in GATA6+/- cells.
RA signaling modulates neural crest cell development (Martínez-Morales et al., 2011) and at day 4 GATA6R456G/R456G cells had increased PAX3 and PAX7 expression (100-fold and 3-fold, respectively; Supplementary file 2, Figure 3—figure supplement 1B). HOXA1 (6-fold increased) and HOXB contribute to activating expression of ZIC1 (8-fold increased) (Jaurena et al., 2015; Makki and Capecchi, 2011; Tvrdik and Capecchi, 2006) that with PAX3/7 induce and specify a neural crest gene program. PAX3 also promotes neural crest migration into the cardiac outflow tract development (Conway et al., 1997) and like STRA6, (Coles and Ackerman, 2013) and contributes to diaphragm development (Stuelsatz et al., 2012) by specifying paraxial mesoderm (Magli et al., 2019). Aberrant RA signaling may contributed to dysregulation of other developmental regulators of diaphragm formation, including NR2F2 (~8-fold increased) and ZFPM2 (5- to 12-fold decreased).
We further assessed activation of RA signaling by comparing RNA-Seq analyses of day 4 differentiating WT and GATA6R456G/R456G cells that were cultured with or without an inhibitor of the ALDH1A2 enzyme (METHODS). Treated WT cells (Figure 4—figure supplement 1A) had significantly altered expression (p<0.05, 1.5-fold), decreasing the expression of transcripts assigned to Gene Ontology terms for heart and muscle cell differentiation and increasing endodermal fate specification and Wnt signaling transcripts. Treated GATA6R456G/R456G cells (Figure 4—figure supplement 1B) had increased expression of transcripts assigned to Gene Ontology terms for myoblast differentiation, circulatory system development and anatomical structures. Notably, the expression of 25% of genes in treated GATA6R456G/R456G cells (Figure 4—figure supplement 1C–F) had levels found in untreated WT cells (HOXB1 and FOXH1) or normalized levels, approaching those in untreated WT cells (HOXA1, HAND2, RARB, TBX20).
GATA6R456G/R456G cells had aberrant expression of other endoderm genes, including markedly lower transcripts of HNF gene family members and striking loss (≥100-fold below WT) of FOXA1 and FOXA2 expression (Supplementary file 2). Biallelic deletion of FOXA2 in human stem cells prevents pancreatic specification (Lee et al., 2019). GATA6R456G/R456G cells also had increased expression of SOX6, which represses PDX1-dependent transcriptional activation of the insulin gene, INS (Iguchi et al., 2005). The cumulative effect of these deficits could account for the extinguished expression of PDX1 and contribute to the high association of pancreatic agenesis with heterozygous GATA6R456G mutations (Figure 1).
Despite multiple transcriptional abnormalities, metabolic enrichment yielded GATA6R456G/R456G cardiomyocytes (day 30) with prototypic gene expression. However, like GATA6+/- hiPSC-CMs, GATA6R456G/R456G hiPSC-CMs exhibited a higher MYH6:MYH7 transcript ratio than WT, indicating immaturity.
GATA6 is a pioneer factor for cardiac development
We studied potential mechanisms by which GATA6 regulated gene transcription by performing Assays for Transposase-Accessible Chromatin sequencing (ATAC-seq), GATA6 chromatin immunoprecipitation sequencing (ChIP-seq) and Hypergeometric Optimization of Motif EnRichment (HOMER) analyses at day 4 (Figure 5, Figure 6) and day 8 (Supplementary file 4). ChIP-seq of GATA6-/- cells yielded very few peaks above background (data not shown), confirming antibody specificity.
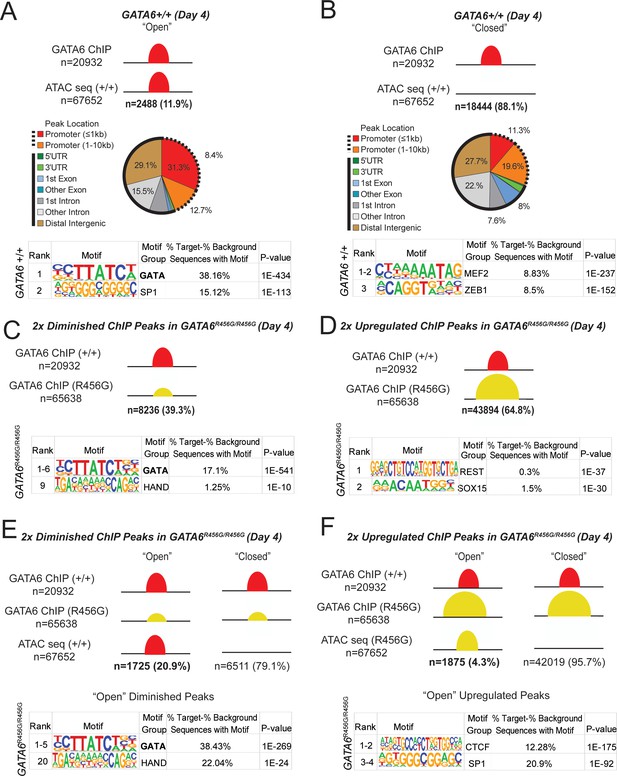
GATA6 is a pioneer factor for cardiac development.
ATAC-seq and GATA6 ChIP-seq were performed in WT and mutant day 4 hiPSCs and overlapped to assess GATA6 direct binding to open vs. closed chromatin. (A) Approximately 12% of GATA6 ChIP-seq peaks overlapped with an ATAC-seq peak and were characterized as ‘Open’. The genomic location of these ChIP-seq peaks in open chromatin were characterized with respect to gene bodies, and DNA-binding motif enrichment was performed using HOMER analysis (METHODS). When GATA6 binds to open regions of chromatin, peaks are enriched for the GATA motif. (B) The remaining 88% of GATA6 ChIP-seq peaks were characterized as ‘Closed’. The genomic location of these ChIP-seq peaks were characterized with respect to gene bodies, and DNA-binding motif enrichment was performed using HOMER analysis. When GATA6 binds to closed regions of chromatin, peaks are enriched for the MEF2 motif. (C) Of the 20932 WT GATA6 ChIP-seq peaks, 39.3% were reduced in GATA6R456G/R456G cells (adjusted p<1e-4, two fold). These peaks were enriched for the GATA and HAND2 binding motifs by HOMER analysis. (D) Of the 67652 GATA6R456G/R456G ChIP-seq peaks, 64.8% were upregulated in GATA6R456G/R456G cells (adjusted p<1e-4, two fold). These peaks were enriched for the REST and SOX binding motifs by HOMER analysis. (E) Peaks diminished in GATA6R456G/R456G cells were overlapped with WT ATAC-seq data to establish chromatin accessibility. Almost 21% of peaks diminished in GATA6R456G/R456G cells were in open chromatin regions; these peaks were enriched for the GATA and HAND motifs. (F) Peaks upregulated in GATA6R456G/R456G cells were overlapped with GATA6R456G/R456G ATAC-seq data to establish chromatin accessibility. Four percent of peaks enriched in GATA6R456G/R456G cells were in open chromatin regions; these peaks were enriched for the CTCF and SP1 motifs.
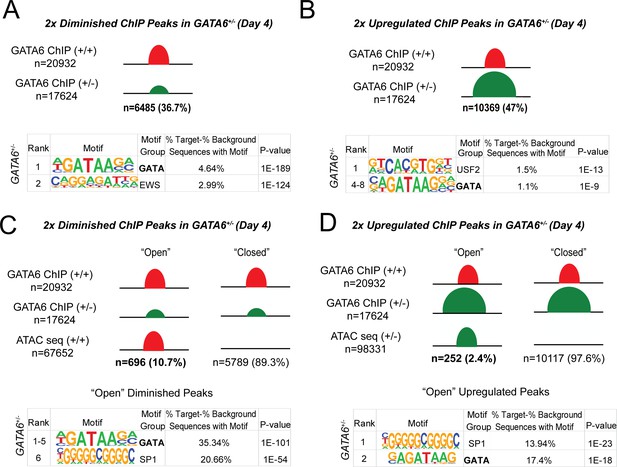
Epigenetic abnormalities in GATA6+/- cells.
(A) Of the 20932 WT GATA6 ChIP-seq peaks, 39.3% were reduced in GATA6+/- cells (adjusted p<1e-4, two fold). These peaks were enriched for the GATA and EWS binding motifs by HOMER analysis. (B) Of the 17624 GATA6+/- ChIP-seq peaks, 47% were upregulated in GATA6+/- cells (adjusted p<1e-4, two fold). These peaks were enriched for the USF2 and GATA-binding motifs by HOMER analysis. (C) Peaks diminished in GATA6+/-cells were overlapped with WT ATAC-seq data to establish chromatin accessibility. Only 10.7% of peaks diminished in GATA6+/- cells were in open chromatin regions; these peaks were enriched for the GATA and SP1 motifs. (F) Peaks upregulated in GATA6R456G/R456G cells were overlapped with GATA6+/- ATAC-seq data to establish chromatin accessibility. Only 2.4% of peaks enriched in GATA6+/- cells were in open chromatin regions; these peaks were enriched for the SP1 and GATA motifs.
At day 4, GATA6 bound ~21,000 open chromatin peaks genome-wide, of which ~ 21% are near promoters. Notably, only 12% of GATA6-bound open chromatin peaks overlapped with ATAC peaks and were markedly enriched for a GATA-binding motif (Figure 5A). ChIP-seq data from day 8 cells were similar (Supplementary file 4). These data inferred that GATA6 directly participates in regulating transcriptional activation of EOMES, GATA4, MEIS1/2, NKX2.5, TBX5, and TBX20 (Supplementary file 4).
By contrast, 88% of GATA-bound peaks occurred in closed chromatin (Figure 5B). Sequences within these peaks were not enriched for the GATA motif but were highly enriched for the MEF2 binding motif. MEF2 proteins contain a DNA-binding domain (MADS) and transcriptional activating domain (TAD) (Backs and Olson, 2006) that promote differentiation gene programs for cardiac, skeletal, and smooth muscle myocytes (Potthoff and Olson, 2007). During cardiomyocyte differentiation of WT hiPSCs MEF2 transcripts increased 120-fold between days 4–8 (Supplementary file 2). In comparison, at day 8–12, MEF2A and MEF2C levels were decreased in GATA6+/- cells. As together, these data implied that GATA6 functioned as a cardiac pioneer factor, which has been proposed for other GATA proteins, we examined the correlation between GATA6 bound to closed chromatin at day 4 and gene expression at day 5 in WT cells. There were 2878 genes differentially expressed between day 4 and day 5 WT cells. Of those, 1049 were associated with GATA6-bound closed chromatin (36.4%). Among these, 583 (56%) genes had increased expression at day 5, including key cardiac developmental transcription factors, GATA4, SMYD1, KDR, and TBX5 (Figure 5—figure supplement 1, Supplementary file 5). Based on these data, we propose that GATA6 is a cardiac development pioneer factor.
Epigenetic abnormalities in GATA6 mutant iPSCs
In comparison to WT, day 4 GATA6+/- cells had~6500 diminished GATA6 ChIP-seq peaks and ~5800 diminished ATAC peaks (Figure 6A and Figure 6—figure supplement 1). Diminished ATAC peaks in proximity to promoters were enriched for GATA, HAND, HNF, and SOX motifs (Figure 6—figure supplement 1, Figure 3, Figure 4 and Supplementary file 2), indicating that attenuated expression of these transcription factor family members could have direct functional consequences. Notably however, few diminished GATA-bound peaks resided within open chromatin (10.7%), of which only 35% were enriched for GATA motifs (Figure 5—figure supplement 1B, Figure 6C). Instead, and consistent with properties of a pioneer factor, GATA6+/- cells had prominent loss (89%) and gain (97%) of GATA-bound peaks within closed chromatin (Figure 6B–D). From these data we infer that GATA interactions with closed chromatin are highly sensitive to protein dosage, while interactions with open chromatin at promoter regions remain relatively intact despite reduced GATA6 protein levels.
By contrast, day 4 GATA6-/- cells contained~29,000 diminished ATAC-seq peaks of which ~ 31% resided in proximity to promoters (Figure 6—figure supplement 1). Profiles at day 8 cells were similar (Figure 6—figure supplement 2). Notably, diminished ATAC peaks showed enrichment in the binding motif for NFY that encodes the ubiquitous promoter element binding factor of CCAAT-boxes. These data implied that extinguishing GATA6-/- expression caused widespread deficits in promoter activation, which likely accounted for failed mesoderm specification and cardiomyocyte differentiation.
ChIP-seq analyses of GATA6R456G/R456G cells (Figure 5C–F, Figure 5—figure supplement 1, Supplementary file 4) identified ~8000 diminished peaks. In comparison to other mutant lines, GATA6R456G/R456G cells had a significantly higher proportion of diminished peaks (21%) in open chromatin that contained a GATA or HAND binding motif and 19% of these diminished peaks were associated with differential gene expression (Figure 5—figure supplement 1C). Additionally, missense cells had a remarkable number (~44,000) of augmented and ectopic peaks. We suggest that these aberrant chromatin interactions reflected direct promiscuous GATA6R456G binding activity or perhaps occurred in response to aberrant RA signaling.
Among the upregulated peaks,~4% of these peaks had GATA-bound to open chromatin, with enrichment of the CTCF binding motif. Furthermore,~12% were associated with differential gene expression, including MSX1, FOXA1, MEIS2, HCN4, and HOXA2 (Supplementary file 4). However, the vast majority of increased or ectopic peaks (96%) resided within closed chromatin and lacked the GATA6 binding motif. Even so, 14% of the differential closed peaks were associated with differential gene expression, including peaks in TBX20, FOXA1, and FOXH1. Our data suggest that the GATA6R456G variant impairs binding to the GATA motif, reducing normal GATA6 function, and also promotes promiscuous binding with either repressive or activating transcriptional effects.
Phenotypes associated with epigenetic abnormalities in GATA6 mutant iPSCs
We examined ATAC-seq and GATA6-ChIP-seq data to identify potential mechanisms for altered gene transcription with relevance to clinical phenotypes in CHD patients with pathogenic GATA6 variants. GATA6-bound peaks were universally decreased in GATA6+/- cells, including those associated many cardiac developmental transcription factors genes (e.g., HAND2, KDR, and TBX5 Figure 7A–C) and these changes were associated with normal or modest reduction in transcript levels (Supplementary file 2). While many of these GATA6-bound peaks were also diminished in GATA6R456G/R456G cells, the missense cells also had many enhanced and augmented peaks which were associated greater differential (decreased or increased) gene expression. For example, only GATA6R456G/R456G cells had diminished GATA6-bound peaks identified in WT cells associated with GATA4 (Figure 7D) and MEF2A and also new GATA6-bound peaks in MEF2C, ZIC1 and ZIC3 (Figure 7E). The distinct epigenetic profiles in GATA6R456G/R456G may contribute to the greater dysregulation of cardiac gene expression and cardiomyocyte maturation in comparison to GATA6+/-cells (Figures 2 and 3 A-C). However regardless of whether epigenetic changes in GATA6R456G/R456G cells decreased normal gene expression or erroneously activated the expression of other genes, the associated cardiac consequence of altered transcription was similar to that of GATA6+/-cells – disruption of cardiac outflow tract development (Figure 1).
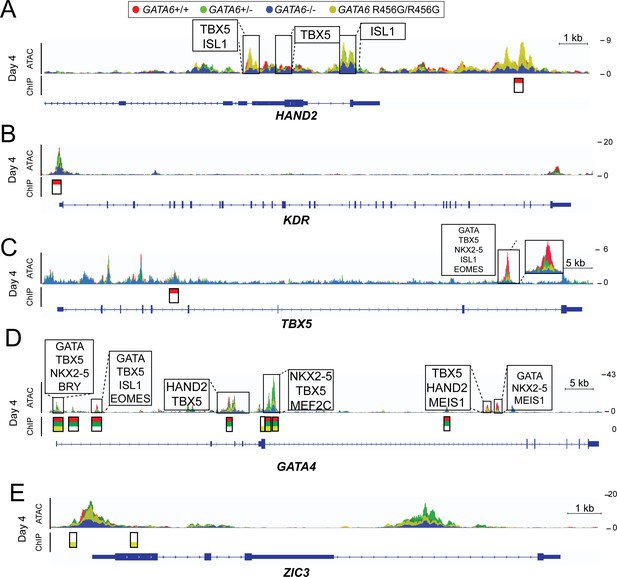
ATAC-seq and GATA6 ChIP-seq analysis of GATA6 variant hiPSC-CMs reveals aberrant binding to congenital heart disease genes.
ATAC peaks (upper), GATA6 ChIP peaks (lower) and DNA-binding motifs (upper, boxed) found in day 4 GATA6 LoF and GATA6R456G/R456G cells, visualized using the Integrative Genomics Viewer (IGV) (A) Lost GATA6 ChIP-seq peak in the HAND2 locus, with differential chromatin accessibility (ATAC-seq). (B) Lost GATA6 ChIP-seq peak at the KDR promoter, leading to reduced chromatin accessibility. (C) Lost GATA6 ChIP-seq peak in TBX5. (D) GATA6R456G does not bind the GATA4 locus in regions of open chromatin. (E) GATA6R456G ectopically binds the ZIC3 promoter.
By contrast, altered GATA6 binding and chromatin accessibility with accompanying dysregulated transcripts likely contributed to the extra-cardiac phenotypes that disproportionately affect patients with heterozygous GATA6 R456G and other exon four missense variants. Pancreatic development requires cooperative GATA4 -GATA6 interactions and compound Gata4-null and Gata6-haploinsufficient mice have reduced Pdx1 expression, pancreatic agenesis or profound hypoplasia (Carrasco et al., 2012). In addition to reduced occupancy of GATA6-motifs in GATA4 with striking attenuated expression (Figure 7D, Supplementary file 2, Supplementary file 4), GATA6 R456G also aberrantly bound chromatin associated with FOXA2, DEANR1 (LINC00261), and PDX1 and diminished transcription of these genes (Figure 7—figure supplement 1). DEANR1 encodes a long noncoding RNA that specifies endoderm and pancreatic lineages by regulating the expression of FOXA2 (Jiang et al., 2015) which in turn regulates PDX1 expression (Gao et al., 2008).
Aberrant RA signaling may have contributed to the abnormal epigenetic profiles in GATA6R456G/R456G cells. In missense cells, prominent ATAC peaks were identified in association with binding motifs for GATA, SOX6, HOX, and RARa in the promoter and exon 1 of ALDH1A2 and transcript levels were increased. SOX6 also had augmented ATAC peaks with RARa and GATA motifs, and GATA6R456G bound an intron one motif (Figure 7—figure supplement 2A–B). GATA6R456G/R456G cells also showed increased ATAC peak heights with RARa binding motifs in close proximity to the reported RA-responsive element in HOXB1 that promotes foregut expression. We also observe GATA6R456G protein aberrantly bound to the STRA6 locus, potentially affecting expression of this gene (Figure 7—figure supplement 2C) These data suggest multiple direct and indirect epigenetic mechanisms by which the GATA6R456G variant distinctly perturbed gene programs and resulted in maldevelopment of the diaphragm, pancreas and other abdominal organs in addition to cause cardiovascular malformations.
Discussion
By combining the strengths of WES, CRISPR/Cas9 genome editing, and hiPSCs, we demonstrate molecular and developmental mechanisms by which damaging variants in GATA6 cause cardiac and extra-cardiac congenital anomalies. Transcriptional and epigenetic analyses of differentiating cardiomyocytes from isogenic WT and mutant hiPSCs provide evidence that GATA6 functions as a pioneer factor that modifies chromatin accessibility and promotes the expression of gene networks involving HAND2, VEGFR and neural crest cells that enable second heart field development and patterning of the outflow tract and atria. Our data also supports critical roles for GATA6 in endodermal and retinoic acid signaling gene networks that gives rise to the pancreas and diaphragm. We find that LoF GATA6 variants repress epigenetic modification and transcriptional activity, while an exon 4 GATA6 missense variant alters normal and also causes ectopic epigenetic effects that enhanced retinoic acid signaling resulting in profound deleterious consequences on gene expression. The selective transcriptional deficits evoked by these variants provides mechanistic insights into the high prevalence of particular heart malformation and associated pancreatic dysgenesis and congenital diaphragmatic hernias that occur in CHD patients with pathogenic GATA6 variants (Figure 8).
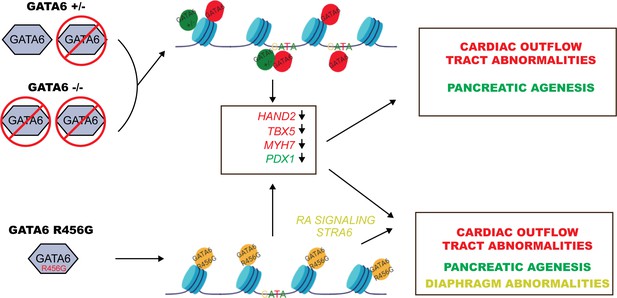
Model for GATA6 transcriptional regulation of cardiac and pancreatic gene expression (see Discussion).
ChIP-seq and ATAC-seq data of WT hiPSCs identified GATA6 bound to closed chromatin in intergenic regions without a GATA-binding motif. Moreover, GATA6 binding was associated with temporal activation of transcription in nearby genes that activate cardiomyocyte and endoderm gene network. These findings indicate that GATA6 engages chromatin and fosters a competent state for transcription factor binding and transcriptional activation, supporting the conclusion that GATA6 is a pioneer factor, as is suggested for other GATA proteins (Fisher et al., 2017). Notably, nonsense-mediated decay of GATA6+/- transcripts reduced GATA6 protein levels, altered chromatin accessibility and decreased gene transcription, implying that intergenic sites are sensitive to GATA6 dosage. In addition, GATA6 functions as a traditional transcription factor, binding GATA motifs in promoters and activating transcription. These functions were relatively insensitive to half normal GATA6 levels.
Our analyses demonstrate that some GATA6 expression is required for lineage specification of human cardiomyocytes. When cultured to promote cardiomyocyte differentiation GATA6-/- cells had markedly reduced promoter activation with NFY transcripts that encode the binding factor of CCAAT-box, a nearly ubiquitous promoter element. GATA6-/- cells were viable but showed very limited transcriptional evidence of mesoderm specification and extinguished cardiomyocyte gene expression. Instead GATA6-/- cells adopted a gene program suggestive of nonspecific cells with expression of neuroectodermal and fibroblast genes.
These observations are consistent with prior analyses of Gata6 ablation in mice that exhibit early embryonic lethality due to defects in extraembryonic endoderm formation (Gottlieb et al., 2002; Morrisey et al., 1998; Zhao et al., 2005). While a tetraploid complementation system reported circumvention of this defect (Zhao et al., 2005), only diminutive Gata6-null embryos were recovered with poor tissue integrity and embryonic resorption after E10.5. Although reverse-transcription-PCR of ‘rescued’ E9.5 Gata6-null embryos identified transcripts associated with early cardiomyocyte lineages, similar transcripts were absent in GATA6-/- hiPSCs from which no cardiomyocytes emerged. Perhaps these differences reflect diffusible factors from other cell lineages not present in hiPSCs cultured in vitro, species-specific dosage requirements, or other mechanisms.
GATA6 haploinsufficiency influenced cardiomyocyte differentiation. ATAC and GATA6 ChIP-seq peaks were depleted in multiple developmental cardiac genes including SMYD1, NKX2-5, and ISL1 in GATA6+/- lines. As SMYD1 regulates HAND2 expression (Gottlieb et al., 2002), depression of SMYD1 transcripts likely resulted in a cascade effect, depleting HAND2-network genes. Notably, transcriptional deficits did not persist at differentiation day 30 GATA6+/- hiPSC-CMs when many cardiac transcription factors (SMYD1, HAND2, NKX2-5, GATA4, and MEIS1) were expressed at levels found in WT cells. We presume that a compensatory transcriptional mechanism restored sufficient expression of many gene programs. However, ISL1 transcripts remained abnormally high, and expression profiles of fetal and adult myosins (high MYH6 and low MYH7, respectively) indicated immaturity of GATA6+/- hiPSC-CMs. Persistence of this developmental deficit in vivo could contribute to longitudinal adverse cardiac outcomes in patients with pathogenic GATA6 variants.
Our cardiomyocyte differentiation protocol also activated endodermal gene programs and uncovered misexpression in mutant lines. Aberrant but variable epigenetic profiles were identified in GATA6+/- cells and in GATA6R456G/R456G cells that led to misexpression of HNF and FOXA gene family-members occurred in both mutant lines. For example, both mutant lines had diminished GATA6 binding of open chromatin binding to HNF1 with reduced transcript levels. By contrast, only GATA6R456G/R456G cells lacked GATA6 binding to open chromatin near DEANR1 a regulator of FOXA2 expression (Gao et al., 2008). GATA6R456G/R456G cell also reduced expression of GATA4. While these data illustrate multiple mechanisms for depleting PDX1 expression and account for the prominent association of CHD and pancreatic agenesis in patients with GATA6 LoF and exon four missense variants, the pathways leading to this shared consequence were strikingly different. Epigenetic changes observed in GATA6+/- cells depleted normal GATA6 binding to chromatin with attenuated gene transcription. Epigenetic changes observed in GATA6R456G/R456G cells were far more sweeping, repressing many more genes and erroneously activating others.
We found that GATA6R456G/R456G cells strikingly increased activation of RA signaling. Indeed, early inhibition of excessive RA signaling normalized some of the many aberrant in GATA6R456G/R456G cells. Transcriptional data showed increased expression of the RA biosynthetic enzyme encoded by ALDH1A2 and of STRA6, which encodes the integral membrane receptor that triggers release and uptake of retinol from circulating retinol-binding protein (Chen et al., 2016). As RARa motifs occur in ATAC peaks associated with HOXA2, HOXB1, and SOX6 genes and others, we deduced that GATA6R456G/R456G likely activates transcription of these genes by increasing RA signaling. Moreover, as ATAC peaks in the ALDH2A2 contain SOX6-binding motifs, enhanced SOX6 expression would increase RA biosynthesis that in turn would further increase STRA6 expression (Bouillet et al., 1997) to further amplify RA signaling.
Human congenital anomalies arise from inadequate and excessive expression of key developmental signaling genes. Our analyses support the conclusion that CHD, pancreatic malformations and congenital diaphragmatic hernia that occur in GATA6 exon four missense mutants reflect both loss of physiologic levels of GATA6 and excessively activated RA signaling. Development of the second heart field and neural crest cell migration into the nascent outflow tract is regulated by RA signaling (Zaffran and Kelly, 2012), and when inappropriate, cardiac morphogenesis is perturbed (Perl and Waxman, 2019). RA signaling also regulates transcriptional signals to develop the diaphragm (Kardon et al., 2017), particularly formation of the central tendon (Coles and Ackerman, 2013) around which diaphragmatic myocytes are patterned. As RA signals inhibit the expression of myogenic specification genes, including PAX3 (El Haddad et al., 2017; Magli et al., 2019), excessive RA signaling could impair diaphragm formation by impairing tendon development in addition to altering differentiation and maturation of diaphragmatic myocytes. Increased RA signaling also augments SOX6 expression, which attenuates PDX1 activation of insulin signaling. When combined with deficient expression in critical endodermal genes (FOXa1 FOXa2), specification and differentiation of pancreatic progenitors are likely to be impaired. Consistent with these RA-mediated mechanisms, we note that human mutations in STRA6 cause syndromic congenital diaphragmatic hernia with both CHD and pancreatic anomalies (Golzio et al., 2007; Pasutto et al., 2007).
Based on the heightened risk for congenital diaphragmatic hernia and pancreatic agenesis only among patients with missense variants within exon 4, we suggest that the encoded ZF domain in GATA6 has critical interactions with chromatin and DNA sequences that evoke the epigenetic and transcriptional consequences. Exon four missense residues perturb these functions- culminating in augmented RA signaling and extinguished PDX1 expression. We suggest that missense variants residing outside of exon four do participate in these molecular processes, nor convey similar risks for developmental anomalies. Conformation of this hypothesis would improve clinical interpretation of GATA6 genotypes.
Our studies also demonstrate that in vitro differentiation of hiPSCs evoke developmental gene expression profiles, independent of the many three-dimensional morphological cues that occur in vivo. This system has the potential to illuminate the consequences of lethal mutations that difficult to study in model organisms. While we recognize the limitations of cell models, by interpreting our molecular data with prior mouse studies and clinical studies of human patients, we believe that these iPSC studies provide valid molecular mechanism by which human GATA6 mutations cause cardiac outflow tract defects, pancreatic agenesis, and congenital diaphragmatic hernias.
In conclusion, we show that genetic perturbation of a pioneer factor, GATA6, altered network-level transcriptional pathways that are critically involved in development of the heart (GATA4, HAND2), endodermal lineages (HNF, FOXA1, FOXA2), and pancreas (PDX1, SOX6) and diaphragm (NR2F2, STRA6, ZFPM2). Development of these organs is highly sensitive to changes in GATA6 gene dosage from LoF variants and from missense variants in the second zinc-finger domain that alter RA signaling.
The combination of WES, genome sequencing data, CRISPR/Cas9 genome editing, and molecular analyses of developmentally immature human iPSC-derived lineages represents a valuable paradigm for mechanistic studies of human development. We expect that this platform will continue to illuminate molecular understandings of organogenesis that may improve clinical use of genetic data for personalized medicine.
Materials and methods
| Reagent type (species) or resource | Designation | Source or reference | Identifiers | Additional information |
|---|---|---|---|---|
| Cell line (human) | PGP1 | Lee et al., 2009 | GM23338 | Male; mycoplasma-free |
| Cell line (human) | TNNT2-GFP | This paper | Derived from PGP1 cell line GM23338, mycoplasma-free | |
| Commercial assay or kit | Zero Blunt TOPO PCR Cloning Kit | ThermoFisher | K280002 | |
| Commercial assay or kit | Human Stem Cell Nucleofector Kit | Lonza | VPH-5022 | |
| Commercial assay or kit | Nextera XT Sample Preparation Kit | Illumina | FC-131–1096 | |
| Commercial assay or kit | Nextera DNA Sample Kit (ATAC-seq) | Illumina | FC-121–1030 | |
| Commercial assay or kit | Tru ChIP Chromatin Shearing Hit | Covaris | 520154 | |
| Commercial assay or kit | Chromium i7 Multiplex Kit | 10X Genomics | 1000073 | |
| Commercial assay or kit | Chromium Chip B Single-Cell Kit | 10X Genomics | 1000075 | |
| Sequence-based reagent | Guide RNAs | This paper | GATA6 Exon 2 Guide 1: GAGCCCCTACTCGCCCTACG GATA6 Exon 2 Guide 2: GCCCCTACTCGCCCTACGTG GATA6 Exon 4 Guide 1: GGCGTTTCTGCGCCATAAGG | |
| Sequence-based reagent | GATA6 sequencing primers | This paper | PCR Primers | GATA6 Exon 2 Sequencing Primer Left/Forward GACGTACCACCACCACCA GATA6 Exon 2 Sequencing Primer Right/Reverse CTTACCTGCACTGGGACCC GATA6 Exon 4 Sequencing Primer Left/Forward TGAATTCACGGAGACAGGCT GATA6 Exon 4 Sequencing Primer Right/Reverse TACAAGTGAGCAGAATACATGGCA |
| Sequence-based reagent | ATAC-seq amplification oligos | Buenrostro et al., 2015 | ||
| Recombinant DNA reagent | Cas9 plasmid | Addgene | PX459v2 | |
| Chemical compound, drug | WIN 18446 | Tocris | ||
| Antibody | rabbit mono-clonal Gata6 | Cell Signaling Technology | 5851S | 10 ug/ChIP |
| Antibody | rabbit mono-clonal Gata6 | Abcam | Ab175927 | 1:1000 dilution |
| Software, algorithm | RNA-seq pipeline: bcbio-nextgen | Chapman et al., 2020 | v.1.2.3 | Hg19 |
| Software, algorithm | R Package: DESEQ2 | Love et al., 2014 | v. 2.1.18.1 | |
| Software, algorithm | R Package: Seurat | Stuart et al., 2019 | v.3.0.0 | |
| Software, algorithm | R Package: ChIP-seeker | Yu et al., 2015 | v.1.14.1 | |
| Software, algorithm | HOMER | Heinz et al., 2010 | v4.10.3 |
Human subjects
Request a detailed protocolCHD subjects were recruited to the Congenital Heart Disease Network Study of the Pediatric Cardiac Genomics Consortium (CHD GENES: ClinicalTrials.gov identifier NCT01196182) after approval from Institutional Review Boards as previously described (Jin et al., 2017; Gelb et al., 2013). Written informed consent was received from subjects or their parents prior to inclusion in the study. Any subject with CHD, regardless of age, race, or ethnicity was eligible for enrollment. Subjects with variants in GATA6 and 2nd heart field genes including CHD diagnoses, and extra-cardiac phenotypes are provided in Supplementary file 1.
Exome sequencing and analyses
Request a detailed protocolDNA was extracted peripheral blood samples and captured using the Nimblegen v.2 exome capture reagent (Roche) or Nimblegen SeqxCap EZ MedExome Target Enrichment Kit (Roche) followed by Illumina DNA sequencing as previously described (Jin et al., 2017). Reads were mapped to the reference genome (hg19), and further processed using the GATK Best Practices workflows as previously described (Jin et al., 2017; McKenna et al., 2010). Single nucleotide variants (SNVs) and small indels were called with GATK HaplotypeCaller and filtered for rarity (ExAC allele frequency ≤10–5) and quality as previously described (Jin et al., 2017). The MetaSVM algorithm was used to predict deleteriousness of de novo missense mutations (annotated as ‘D-Mis’) using software defaults (Dong et al., 2015; Sulahian et al., 2014).
TNNT2-GFP, GATA6+/-, GATA6-/-, and GATA6R456G/R456G hiPSCs and iPSC-CMs
Request a detailed protocolAll hiPSCs and hiPSC-CMs used here are derived from the male parent iPSC line PGP1 (Akerman et al., 2017; RRID-GM23338) and are mycoplasma-free. The TNNT2-GFP iPSC line was generated by homology-directed repair, using a plasmid with the endogenous TNNT2 sequence connected to a GSSSS linker region, attached to the GFP gene, and the entire construct was flanked by 500 bp homology arms. PGP1 iPSCs were transfected as described (Sharma et al., 2018b) to obtain homozygous integration of the GFP tag in both TNNT2 alleles. Edited clones were selected using puromycin and expanded and genotyped as previously described (Sharma et al., 2018b) and differentiated into cardiomyocytes as described (Sharma et al., 2018a).
GATA6+/- and GATA6-/- hiPSCs were generated from TNNT2-GFP and PGP1 iPSCs by non-homologous end joining using a 2 µg plasmid expressing Cas9 (PX459v2 from Addgene) that was co-transfected with 2 µg plasmid expressing guide RNA (provided in Key Resource Table) using a stem cell nucleofector kit (Amaxa). Edited clones were selected using puromycin and expanded and genotyped as previously described (Sharma et al., 2018b) and differentiated into cardiomyocytes as described (Sharma et al., 2018a).
GATA6R456G/R456G hiPSCs were generated by homology-directed repair using a 2 µg plasmid expressing Cas9 (PX459v2 from Addgene) that was co-transfected with 2 µg plasmid expressing guide RNA (provided in Key Resource Table) and 2 µg single-stranded oligonucleotide (HDR template) using a stem cell nucleofector kit (Amaxa). Edited clones were selected using puromycin and expanded and genotyped as previously described (Sharma et al., 2018b) and differentiated into cardiomyocytes as described (Sharma et al., 2018a).
Confirmation of editing in subcloned iPSC lines
Request a detailed protocolAll edited lines sub-cloned, by dissociating 1000 genome-edited hiPSCs cells, filtering through a 60 µm strainer, and evenly distributing them onto a 15 cm dish containing Matrigel and mTeSR+ rho kinase inhibitor. Individual monoclonal hiPSC colonies were picked when colony size reached approximately 200 cells and placed into individual separate wells of a 96 well plate. Clones were allowed to grow to 80% confluency, at which time a sample was obtained for PCR amplification to verify the GATA6 variant or GFP-tagged TNNT2 and to assess zygosity. PCR-amplified fragments (primers provided in Key Resource Table) containing putative variants were submitted for Sanger sequencing and next-generation sequencing. Sanger sequencing traces were deconvoluted using TIDE software to confirm zygosity. Clones carrying GATA6 mutations were further expanded for subsequent culture and differentiation.
hiPSC-CM differentiation
Request a detailed protocolThe hiPSC-CMs were generated from GATA6 mutant hiPSCs using a small-molecule mediated differentiation approach that modulates Wnt signaling (Sharma et al., 2018a). Cells began beating at approximately day 7 post-differentiation. Cardiomyocytes were metabolically selected from other differentiated cells by using glucose deprivation as previously described (Sharma et al., 2018a). hiPSC-CM Western blots: GATA6 protein expression and nuclear localization were performed using ab175927.
RNA-sequencing and analysis
Request a detailed protocolRNA-seq experiments were performed on at least two independent cultured cell samples per time point, with the exception of GATA6R256G/R256G DMSO control in the RA-signaling experiment. Trizol (Thermo Fisher) was used to harvest differentiating GATA6 mutant hiPSC-CMs (days 0, 4, 8, 12, 30 of differentiation) designated for RNA-sequencing analysis. Samples harvested in Trizol were stored at −80°C until RNA was extracted. All RNA samples had an RNA integrity number of ≥8. Library preparation was conducted using the Nextera library preparation method. RNA-Seq library samples were pooled and run on four lanes (one flow cell) using the Illumina NextSeq500 platform. All data was combined into a single fastq file. Samples typically had 30–50M reads each. The raw reads were aligned by HISAT2 (v.2.1.0) to human genome (hg38). The aligned reads were quantified by FeatureCounts, counts were normalized, and differentially expressed genes were identified using DESeq2 (v1.24.0). DESeq2 data was analyzed and visualized in R using the ggplot2 (v3.1.0), pheatmap (v1.0.12), gProfiler (v0.6.7), and VennDiagram (v1.6.20) packages. Single-cell RNA-seq analysis for hiPSC-CMs was conducted using a 10x Genomics Chromium platform and analyzed in R using the Seurat (v. 3.5.1) pipeline. Seurat was used to regress out cell-cell variation in cell complexity, gene expression driven by batch, cell alignment rate, the number of detected molecules, and mitochondrial gene expression. Starting with Seurat v2.0, Seurat implements this regression as part of the data scaling process.
ATAC-seq and Hi-C chromatin analysis
Request a detailed protocolATAC-seq was performed as previously described (Buenrostro et al., 2015; Corces et al., 2017). Briefly, 50,000 cells were harvested and lysed to isolate nuclei. Nuclei were treated with Tn5 transposase (Nextera DNA Sample Prep Kit, Illumina) and DNA was isolated. Fragmented DNA was amplified using bar-coded PCR primers (defined in Buenrostro et al., 2015) and libraries were pooled. Pooled libraries were sequenced (Illumina Next-Seq) to a depth of 100 million reads per sample. Reads were aligned to the hg19 reference genome using BWA-MEM and peaks were called using HOMER (v4.10.3) (http://homer.ucsd.edu/homer/index.html). Functional analysis of ATAC-seq peaks was performed using ChIP-Seeker (v.1.14.1). Motif enrichment was performed using HOMER (v4.10.3). Differential peaks were called using HOMER (v4.10.3).
ChIP-seq
Request a detailed protocolCells (6-well plates) were grown to approximately 80% confluency and then fixed with 1% fresh formaldehyde diluted in media. Cells were quenched with glycine, washed, and harvested. Cell pellets were stored at −80°C. Cells were thawed and nuclei were prepared using the Covaris Tru ChIP Chromatin Shearing Kit (Covaris, MA, USA). Chromatin was sheared using the Covaris E210 to an average fragment size of 200–700 bp and an input sample was purified. Chromatin (5 µg) was incubated with GATA6 antibody (RRID:AB_5851S) at 4°C O/N and then added to Protein G beads for 2 hr at 4°C. Beads were washed and bound chromatin was eluted at 65°C for 30 min on a heated vortex. DNA was then purified and quantified. ChIP-seq libraries were prepared using Nextera XT DNA sample prep (Illumina). Sequences were aligned to hg19 using BWA-MEM. Peaks were called using MACS2 using default parameters with a q value of 0.001. Differential peaks were called using HOMERv4.10.3.
Retinoic acid inhibitor treatment assay
Request a detailed protocolhiPSCs were seeded and differentiated to day 4, and then treated for 24 hr with either DMSO or 1 µM WIN 18446 (Tocris), an inhibitor of the ALDH1A2 enzyme. Cells were harvested and processed for RNA-seq.
Statistical analyses
Request a detailed protocolThe distribution of damaging human variants across the GATA6 gene was analyzed using the binominal test, implemented in R. The association of GATA6+/- and GATA6R456G/+ variants and pancreatic agenesis or congenital diaphragmatic hernia was analyzed using the Fisher Exact test, implemented in R. Transcriptional responses of iPSC-CMs with or without a GATA6 variant was compared using the Student’s t-test was used for comparison. All error bars refer to standard deviation, unless otherwise specified. A p value of < 0.05 was considered significant.
Data availability
All data generated or analyzed during this study are included in the manuscript.
-
NCBI Gene Expression OmnibusID GSM575227. GATA6 ChIP-seq in differentiated cells.
-
NCBI Gene Expression OmnibusID GSM1151694. GATA6 ChIP-seq.
-
NCBI Gene Expression OmnibusID GSE51936. Genome-wide analyses of GATA6 occupancy and functions provide insights into its oncogenic mechanisms in human gastric cancer.
-
NCBI Gene Expression OmnibusID GSM575226. GATA6 ChIP-seq in proliferating cells.
References
-
Control of cardiac growth by histone acetylation/deacetylationCirculation Research 98:15–24.https://doi.org/10.1161/01.RES.0000197782.21444.8f
-
Mutations in the cardiac transcription factor NKX2.5 affect diverse cardiac developmental pathwaysJournal of Clinical Investigation 104:1567–1573.https://doi.org/10.1172/JCI8154
-
Congenital heart disease in adults first of two partsThe New England Journal of Medicine 342:256–263.https://doi.org/10.1056/NEJM200001273420407
-
Building the mammalian heart from two sources of myocardial cellsNature Reviews Genetics 6:826–835.https://doi.org/10.1038/nrg1710
-
Atac‐seq: a method for assaying chromatin accessibility genome‐wideCurrent Protocols in Molecular Biology 109:1–9.https://doi.org/10.1002/0471142727.mb2129s109
-
GATA4 and GATA6 control mouse pancreas organogenesisJournal of Clinical Investigation 122:3504–3515.https://doi.org/10.1172/JCI63240
-
Development of a lethal congenital heart defect in the splotch (Pax3) mutant mouseCardiovascular Research 36:163–173.https://doi.org/10.1016/S0008-6363(97)00172-7
-
Retinoic acid maintains human skeletal muscle progenitor cells in an immature stateCellular and Molecular Life Sciences 74:1923–1936.https://doi.org/10.1007/s00018-016-2445-1
-
Genetics of congenital heart disease: the glass half emptyCirculation Research 112:707–720.https://doi.org/10.1161/CIRCRESAHA.112.300853
-
Dynamic regulation of Pdx1 enhancers by Foxa1 and Foxa2 is essential for pancreas developmentGenes & Development 22:3435–3448.https://doi.org/10.1101/gad.1752608
-
The role of hepatic nuclear factor 1 alpha and PDX-1 in transcriptional regulation of the pdx-1 geneThe Journal of Biological Chemistry 276:47775–47784.https://doi.org/10.1074/jbc.M109244200
-
Matthew-Wood syndrome is caused by truncating mutations in the retinol-binding protein receptor gene STRA6The American Journal of Human Genetics 80:1179–1187.https://doi.org/10.1086/518177
-
Analysis of two distinct retinoic acid response elements in the homeobox gene Hoxb1 in transgenic miceDevelopmental Dynamics 223:353–370.https://doi.org/10.1002/dvdy.10057
-
SOX6 attenuates glucose-stimulated insulin secretion by repressing PDX1 transcriptional activity and is down-regulated in hyperinsulinemic obese miceJournal of Biological Chemistry 280:37669–37680.https://doi.org/10.1074/jbc.M505392200
-
Congenital diaphragmatic hernias: from genes to mechanisms to therapiesDisease Models & Mechanisms 10:955–970.https://doi.org/10.1242/dmm.028365
-
The second heart fieldCurrent Topics in Developmental Biology 100:33–65.https://doi.org/10.1016/B978-0-12-387786-4.00002-6
-
GATA5 interacts with GATA4 and GATA6 in outflow tract developmentDevelopmental Biology 358:368–378.https://doi.org/10.1016/j.ydbio.2011.07.037
-
GATA-6 regulates semaphorin 3C and is required in cardiac neural crest for cardiovascular morphogenesisJournal of Clinical Investigation 116:929–939.https://doi.org/10.1172/JCI27363
-
ConferenceA transcriptional pathway for cardiac developmentCold Spring Harbor Symposia on Quantitative Biology. pp. 405–411.
-
Interaction of Gata4 and Gata6 with Tbx5 is critical for normal cardiac developmentDevelopmental Biology 326:368–377.https://doi.org/10.1016/j.ydbio.2008.11.004
-
The zinc finger-containing transcription factors GATA-4, -5, and -6. ubiquitously expressed regulators of tissue-specific gene expressionThe Journal of Biological Chemistry 275:38949–38952.https://doi.org/10.1074/jbc.R000029200
-
Reiterative mechanisms of retinoic acid signaling during vertebrate heart developmentJournal of Developmental Biology 7:11.https://doi.org/10.3390/jdb7020011
-
MEF2: a central regulator of diverse developmental programsDevelopment 134:4131–4140.https://doi.org/10.1242/dev.008367
-
Differentiation and contractile analysis of GFP-Sarcomere reporter hiPSC-CardiomyocytesCurrent Protocols in Human Genetics 96:1–12.https://doi.org/10.1002/cphg.53
-
CRISPR/Cas9-Mediated fluorescent tagging of endogenous proteins in human pluripotent stem cellsCurrent Protocols in Human Genetics 96:1–20.https://doi.org/10.1002/cphg.52
-
A contemporary atlas of the mouse diaphragm: myogenicity, Vascularity, and the Pax3 connectionThe Journal of Histochemistry and Cytochemistry 60:638–657.https://doi.org/10.1369/0022155412452417
-
Reversal of Hox1 gene subfunctionalization in the mouseDevelopmental Cell 11:239–250.https://doi.org/10.1016/j.devcel.2006.06.016
-
Birth prevalence of congenital heart disease worldwide: a systematic review and meta-analysisJournal of the American College of Cardiology 58:2241–2247.https://doi.org/10.1016/j.jacc.2011.08.025
-
New developments in the second heart fieldDifferentiation 84:17–24.https://doi.org/10.1016/j.diff.2012.03.003
-
Genetics and genomics of congenital heart diseaseCirculation Research 120:923–940.https://doi.org/10.1161/CIRCRESAHA.116.309140
-
Pioneer transcription factors: establishing competence for gene expressionGenes & Development 25:2227–2241.https://doi.org/10.1101/gad.176826.111
-
GATA6 is essential for embryonic development of the liver but dispensable for early heart formationMolecular and Cellular Biology 25:2622–2631.https://doi.org/10.1128/MCB.25.7.2622-2631.2005
Article and author information
Author details
Funding
National Institutes of Health (UM1HL128711)
- George Porter
- Martin Tristani-Firouzi
- Deepak Srivastava
- Jonathan G Seidman
- Christine E Seidman
National Institutes of Health (UM1HL128761)
- Christine E Seidman
National Institutes of Health (UM1HL098147)
- Daniel M DeLaughter
National Institutes of Health (U01-HL098153)
- Jonathan G Seidman
- Christine E Seidman
National Institutes of Health (U01-HL098163)
- Jonathan G Seidman
- Christine E Seidman
National Institutes of Health (U01-HL098123)
- Jonathan G Seidman
- Christine E Seidman
National Institutes of Health (U01-HL098162)
- Jonathan G Seidman
- Christine E Seidman
National Science Foundation (EEC-1647837)
- Jonathan G Seidman
- Christine E Seidman
National Institutes of Health (T32HL116273)
- Arun Sharma
Howard Hughes Medical Institute
- Lauren K Wasson
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
We gratefully acknowledge the NHLBI Pediatric Cardiac Genomics Consortium (PCGC) and Cardiovascular Development Consortium (CVDC) investigators for their support and expertise regarding the mechanisms driving congenital heart disease. Funding support for this study was provided in part by the NHLBI CVDC (U01HL098166) PCGC grants (U01-HL098188, U01-HL098147, U01-HL098153, U01-HL098163, U01-HL098123 and U01-HL098162), Fondation Leducq, the Engineering Research Centers Program of the National Science Foundation (NSF Cooperative Agreement No. EEC-1647837), NIH T32 HL116273, the Wellcome Trust (Sir Henry Wellcome fellowship), the German Academic Scholarship Foundation, and the Howard Hughes Medical Institute.
Ethics
Human subjects: CHD subjects were recruited to the Congenital Heart Disease Network Study of the Pediatric Cardiac Genomics Consortium (CHD GENES: ClinicalTrials.gov identifier NCT01196182) after approval from Institutional Review Boards as previously described (Pediatric Cardiac Genomics et al., 2013; Jin et al., 2017). Written informed consent was received from subjects or their parents prior to inclusion in the study. Clinical diagnoses were standardized based on review of medical data and family interviews.
Copyright
© 2020, Sharma et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 4,150
- views
-
- 578
- downloads
-
- 56
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 56
- citations for umbrella DOI https://doi.org/10.7554/eLife.53278






