Gene regulation: A transcriptional switch controls meiosis
To grow and develop properly, cells must control the timing of critical events. This means that the proteins that drive these events need to be active at specific times and, in many cases, this is achieved by precisely regulating when the genes that encode these proteins are transcribed. Now, in two papers in eLife, Elçin Ünal, Folkert van Werven and colleagues at the University of California at Berkeley and the Francis Crick Institute report a transcriptional switch that controls the timing of a critical event in the development of the yeast Saccharomyces cerevisiae (Chen et al., 2017; Chia et al., 2017).
Meiosis is the specialized form of cell division that produces new cells with half as many chromosomes as the parent cell. This process must be precise because too many or too few chromosomes in the new cells can lead to problems. In humans, where meiosis is used to make the egg and sperm cells, these problems include miscarriage or birth defects.
One key player that ensures chromosomes are segregated properly is the kinetochore. This protein complex acts as a molecular bridge between the chromosomes and the microtubule filaments that physically separate pairs of chromosomes during cell division (Duro and Marston, 2015). The kinetochore assembles, disassembles and reassembles during different stages of meiosis (Miller et al., 2012). During the first stage, which is called prophase, homologous chromosomes 'recombine' to exchange genetic information (Figure 1A): the kinetochore must disassemble during this stage to prevent the chromosomes being separated too soon. The expression of a protein called Ndc80, which is a key subunit within the kinetochore, drops when the meiotic kinetochore needs to dismantle, suggesting that the availability of Ndc80 acts as a molecular switch that controls whether kinetochores assemble or disassemble (Meyer et al., 2015; Chen et al., 2017).
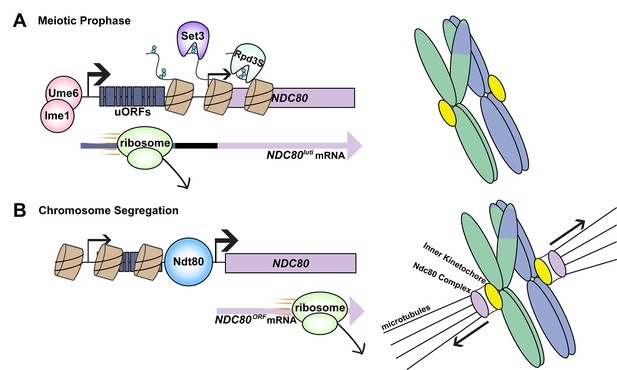
Transcriptional switching between two mRNA transcripts regulates the assembly of the kinetochore during meiosis.
(A) During prophase, homologous chromosomes undergo recombination to exchange genetic information (right). The kinetochore must be dismantled during this time so that chromosomes do not prematurely separate. Cells do this by limiting the availability of a subunit of the kinetochore called Ndc80 (left top). First, Ume6-Ime1 binds to an upstream promoter (thick black arrow) and the DNA is transcribed to produce an extended mRNA transcript named NDC80luti. This process also results in the deposition of methylation marks (small blue circles) on histones (brown cyclinders). Enzymes called histone deacetylases (Set3 and Rpd3S) bind these marks and deacetylate the histones; the end result is to prevent the gene for Ndc80 (lilac rectangle) from being transcribed to produce a short mRNA transcript called NDC80ORF. Reduced transcription from the NDC80ORF promoter is indicated by the thinner black arrow. Additional regulation comes from upstream open reading frames (uORFs) in the extended transcript; the translation of these regions by the ribosome prevents translation reinitiation taking place at the NDC80 start codon (left bottom). (B) Following prophase, the kinetochore reassembles for chromosome segregation (right). Another transcription factor, Ndt80, binds the canonical NDC80 promoter (left top), and the DNA is transcribed to produce the shorter mRNA transcript. Translation of this transcript produces functional Ndc80 protein (left bottom), allowing the kinetochore to assemble and the chromosomes to segregate. Black arrows indicate direction of movement.
In the first paper – which has Jingxun Chen and Amy Tresenrider as joint first authors – the researchers confirmed why the production of Ndc80 must be lowered until just before the stage when the chromosomes segregate (Chen et al., 2017). They saw that, if the gene for Ndc80 in yeast was kept activated throughout meiosis, many cells showed abnormal chromosome segregation.
Earlier studies had revealed that the gene for Ndc80 gives rise to two distinct mRNA transcripts (Brar et al., 2012). Chen et al. now show that the longer of these two transcripts – an extended isoform called NDC80luti – is specific to meiosis, and predominates during prophase. They also show that the gene for Ndc80 has two promoters, one for each transcript, and that the promoter for the extended isoform is recognized by an early meiosis transcription factor complex called Ume6-Ime1 (Figure 1A).
In the second paper, which has Minghao Chia as the first author, the researchers report that switching on the transcription of the extended isoform leads to chemical modifications of chromatin across the downstream promoter. These modifications likely recruit enzymes known as histone deacetylases that modify chromatin in a way that prevents the transcription of the shorter mRNA, which is called NDC80ORF (Jensen et al., 2013; Chia et al., 2017).
Intriguingly, the extended transcript contains nine short upstream open reading frames (uORFs) that are, in fact, translated by the protein synthesis machinery (Brar et al., 2012). The translation of the uORFs in the extended isoform effectively silences production of the Ndc80 protein by preventing translation of the downstream NDC80 coding region. This phenomenon occurs at other uORF-containing genes as well (Hinnebusch et al., 2016). The combined transcriptional and translational interference limits the production of Ndc80 protein in a way that is readily reversible.
Once recombination is complete, the kinetochore must reassemble so that the chromosomes can be segregated. Binding of a later meiotic transcription factor, Ndt80, switches on the expression of the shorter mRNA as the cell exits prophase. Importantly, the repressive chromatin structure established by transcription of the longer mRNA isoform does not block the Ndt80 binding site (Chia et al., 2017). Moreover, the shorter transcript does not contain the uORFs, and so it produces functional Ndc80 protein that can be assembled into the kinetochore (Figure 1B).
Ndc80 is also crucial when chromosomes segregate during mitosis, when cells divide to create daughter cells with the same number of chromosomes as the parent cell. During this kind of cell division in yeast, the expression of the short isoform is ensured by the Ume6 transcription factor repressing the expression of the long isoform. This is because, when Ime1 is absent, Ume6 functions to repress transcription. These observations raise the question of how cells eliminate the existing pools of Ndc80 as they enter prophase in meiosis. Future studies will now need to determine if and how protein degradation contributes to rapid depletion of Ndc80.
These two papers complement previous studies on the importance of proper gene expression in meiotic control. For example, the expression of IME4, an important gene for entry into meiosis, is controlled in yeast by a different type of transcriptional switch in which transcription of the coding strand of DNA is repressed by transcription of the opposite, noncoding strand until the gene’s protein is needed (Hongay et al., 2006). The data used to identify the two NDC80 transcripts also reveal that approximately 190 meiotic genes have extended transcripts like NDC80luti (Brar et al., 2012; Chen et al., 2017; Chia et al., 2017). A goal for the future will be to characterize the expression of these genes, which might confirm that interfering with transcription and translation are general regulatory trends in the tuning of gene expression during meiosis.
References
-
From equator to pole: splitting chromosomes in mitosis and meiosisGenes & Development 29:109–122.https://doi.org/10.1101/gad.255554.114
-
Dealing with pervasive transcriptionMolecular Cell 52:473–484.https://doi.org/10.1016/j.molcel.2013.10.032
-
Ipl1/Aurora-B is necessary for kinetochore restructuring in meiosis I in Saccharomyces cerevisiaeMolecular Biology of the Cell 26:2986–3000.https://doi.org/10.1091/mbc.E15-01-0032
Article and author information
Author details
Publication history
Copyright
© 2017, Hildreth et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 3,492
- views
-
- 300
- downloads
-
- 2
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 2
- citations for umbrella DOI https://doi.org/10.7554/eLife.31911
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Chromosomes and Gene Expression
Cell differentiation programs require dynamic regulation of gene expression. During meiotic prophase in Saccharomyces cerevisiae, expression of the kinetochore complex subunit Ndc80 is downregulated by a 5’ extended long undecoded NDC80 transcript isoform. Here we demonstrate a transcriptional interference mechanism that is responsible for inhibiting expression of the coding NDC80 mRNA isoform. Transcription from a distal NDC80 promoter directs Set1-dependent histone H3K4 dimethylation and Set2-dependent H3K36 trimethylation to establish a repressive chromatin state in the downstream canonical NDC80 promoter. As a consequence, NDC80 expression is repressed during meiotic prophase. The transcriptional mechanism described here is rapidly reversible, adaptable to fine-tune gene expression, and relies on Set2 and the Set3 histone deacetylase complex. Thus, expression of a 5’ extended mRNA isoform causes transcriptional interference at the downstream promoter. We demonstrate that this is an effective mechanism to promote dynamic changes in gene expression during cell differentiation.






