Single-dose BNT162b2 vaccine protects against asymptomatic SARS-CoV-2 infection
Abstract
The BNT162b2 mRNA COVID-19 vaccine (Pfizer-BioNTech) is being utilised internationally for mass COVID-19 vaccination. Evidence of single-dose protection against symptomatic disease has encouraged some countries to opt for delayed booster doses of BNT162b2, but the effect of this strategy on rates of asymptomatic SARS-CoV-2 infection remains unknown. We previously demonstrated frequent pauci- and asymptomatic SARS-CoV-2 infection amongst healthcare workers (HCWs) during the UK’s first wave of the COVID-19 pandemic, using a comprehensive PCR-based HCW screening programme (Rivett et al., 2020; Jones et al., 2020). Here, we evaluate the effect of first-dose BNT162b2 vaccination on test positivity rates and find a fourfold reduction in asymptomatic infection amongst HCWs ≥12 days post-vaccination. These data provide real-world evidence of short-term protection against asymptomatic SARS-CoV-2 infection following a single dose of BNT162b2 vaccine, suggesting that mass first-dose vaccination will reduce SARS-CoV-2 transmission, as well as the burden of COVID-19 disease.
Introduction
The UK has initiated mass COVID-19 immunisation, with healthcare workers (HCWs) given early priority because of the potential for workplace exposure and risk of onward transmission to patients. The UK’s Joint Committee on Vaccination and Immunisation has recommended maximising the number of people vaccinated with first doses at the expense of early booster vaccinations, based on single-dose efficacy against symptomatic COVID-19 disease (Department of Health and Social Care, 2021; Polack et al., 2020; Voysey et al., 2021).
At the time of writing, three COVID-19 vaccines have been granted emergency use authorisation in the UK, including the BNT162b2 mRNA COVID-19 vaccine (Pfizer-BioNTech). A vital outstanding question is whether this vaccine prevents asymptomatic as well as symptomatic SARS-CoV-2 infection or merely converts infections from symptomatic to asymptomatic. Sub-clinical infection following vaccination could continue to drive transmission. This is especially important because many UK HCWs have received this vaccine, and nosocomial COVID-19 infection has been a persistent problem.
Through the implementation of a 24 hour turnaround PCR-based comprehensive HCW screening programme at Cambridge University Hospitals NHS Foundation Trust (CUHNFT), we previously demonstrated the frequent presence of pauci- and asymptomatic infection amongst HCWs during the UK’s first wave of the COVID-19 pandemic (Rivett et al., 2020). Here, we evaluate the effect of first-dose BNT162b2 vaccination on test positivity rates and cycle threshold (Ct) values in the asymptomatic arm of our programme, which now offers weekly screening to all staff.
Results and discussion
Vaccination of HCWs at CUHNFT began on 8 December 2020, with mass vaccination from 8 January 2021. Here, we analyse data from 2 weeks spanning 18–31 January 2021, during which (1) the prevalence of COVID-19 amongst HCWs remained approximately constant and (2) we screened comparable numbers of vaccinated and unvaccinated HCWs. During this period, 4408 (week 1) and 4411 (week 2) PCR tests were performed on individuals reporting well to work, from a weekly on-site HCW population of ~9000. We stratified HCWs <12 days or ≥12 days post-vaccination because this was the point at which protection against symptomatic infection began to appear in the phase III clinical trial (Polack et al., 2020). In the post-vaccination groups, the median number of days between vaccination and testing were 7 (interquartile range [IQR] 4–9; <12 day group) and 16 (14–18; ≥12 day group).
Twenty-six of 3252 (0.8%, Wilson’s interval 0.6–1.2%) tests from unvaccinated HCWs were positive (Ct < 36), compared to 13/3535 (0.4%, Wilson’s interval 0.2–0.6%) tests from HCWs <12 days post-vaccination and 4/1989 (0.2%, Wilson’s interval 0.1–0.5%) tests from HCWs ≥12 days post-vaccination (p=0.023 and p=0.004, respectively; Fisher’s exact test, Figure 1 and Table 1). This suggests a fourfold decrease in the risk of asymptomatic SARS-CoV-2 infection amongst HCWs ≥12 days post-vaccination, compared to unvaccinated HCWs, with an intermediate effect amongst HCWs <12 days post-vaccination.
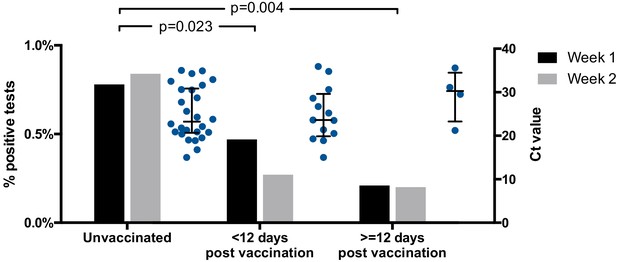
Proportion of positive screening tests for SARS-CoV-2 amongst HCWs from the CUHNHFT asymptomatic screening programme (grey bars; week 1, 18–24 January 2021; week 2, 25–31 January 2021) and Ct values of positive tests (Ct < 36; blue dots; both weeks).
RT-PCR targeting the SARS-CoV-2 ORF1ab genes was conducted at the Cambridge COVID-19 Testing Centre (part of the UK Lighthouse Labs Network). For proportions of positive screening tests, p-values for pair-wise comparisons of unvaccinated HCWs with HCWs <12 days or ≥12 days post-vaccination are shown (Fisher’s exact test; both weeks). For Ct values, medians ± interquartile ranges are shown.
-
Figure 1—source data 1
Proportions of positive asymptomatic SARS-CoV-2 screens and distributions of Ct values.
- https://cdn.elifesciences.org/articles/68808/elife-68808-fig1-data1-v2.xlsx
Weekly numbers and proportions of positive SARS-CoV-2 test results spanning 6 weeks around the main study period (indicated in grey).
| Unvaccinated | <12 Days since vaccination | ≥12 Days since vaccination | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Week start | Total tests | Positive tests | % | Total tests | Positive tests | % | Total tests | Positive tests | % |
| 28 December 2020 | 2097 | 16 | 0.8% | 8 | 0 | 0.0% | 6 | 0 | 0.0% |
| 4 January 2021 | 4762 | 43 | 0.9% | 93 | 0 | 0.0% | 22 | 0 | 0.0% |
| 11 January 2021 | 3273 | 27 | 0.8% | 978 | 6 | 0.6% | 30 | 0 | 0.0% |
| 18 January 2021 | 2183 | 17 | 0.8% | 1716 | 8 | 0.5% | 483 | 1 | 0.2% |
| 25 January 2021 | 1069 | 9 | 0.8% | 1819 | 5 | 0.3% | 1506 | 3 | 0.2% |
| 1 February 2021 | 699 | 1 | 0.1% | 758 | 1 | 0.1% | 2825 | 1 | 0.0% |
A marked reduction in infections was also seen when analyses were repeated with (1) inclusion of HCWs testing positive through both the symptomatic and asymptomatic arms of the programme (56/3370 [1.7%, Wilson’s interval 1.3–2.2%] unvaccinated vs 8/2018 [0.4%, Wilson’s interval 0.2–0.8%] ≥12 days post-vaccination, 4.2-fold reduction, p<0.0001) and (2) inclusion of PCR tests that were positive at the limit of detection (Ct > 36, 42/3268 [1.3%, Wilson’s interval 1.0–1.7%] vs 15/2000 [0.7%, Wilson’s interval 0.5–1.2%], 1.7-fold reduction, p=0.07). In addition, the median Ct value of positive tests showed a non-significant trend towards increase between unvaccinated HCWs and HCWs ≥12 days post-vaccination (23.3 [IQR 13.5–33.0] to 30.3 [IQR 25.5–35.1], Figure 1), raising the possibility that vaccinated individuals who do go on to develop infection may have lower viral loads.
HCWs working in COVID-19 clinical areas were prioritised for vaccination, and a small number of clinically vulnerable HCWs were also given priority. Otherwise, vaccine allocation was arbitrary. Since asymptomatic infection was examined, the date of infection could have been earlier than the test date. These factors would all tend to lead to an underestimate of the vaccine’s effect (bias towards the null). Because of the rapid decline in the incidence of SARS-CoV-2 infection in the Cambridge community, this study could only examine the short-term impact of single-dose BNT162b2 vaccination. The frequency of prior SARS-CoV-2 infection (Cooper et al., 2020) was similar in all groups (seroprevalence 7.1%, unvaccinated; 5.6%, <12 days post-vaccination; 5.7%, ≥12 days post-vaccination), suggesting that this did not confound our observations.
Taken together, our findings provide real-world evidence of short-term protection against asymptomatic SARS-CoV-2 infection after a single dose of BNT162b2 vaccine, at a time when the UK COVID-19 variant of concern 202012/01 (lineage B.1.1.7) accounted for the great majority of infections (24/29 sequenced isolates from asymptomatic HCWs). A fourfold reduction from 0.8% to 0.2% in asymptomatic infection is likely to be crucial in controlling nosocomial SARS-CoV-2 transmission. Nonetheless, protection is incomplete, suggesting that continuing asymptomatic HCW screening, social distancing, mask-wearing, and strict hand hygiene remain vital.
Materials and methods
HCW screening programme
Request a detailed protocolWe previously described protocols for staff screening, sample collection, and results reporting in detail (Rivett et al., 2020; Jones et al., 2020). In general, these methods remained unchanged throughout this study period. Two parallel streams of entry into the testing programme included (1) HCW symptomatic and HCW symptomatic household contact screening arms and (2) an HCW asymptomatic screening arm. Since our prior description of the screening programme, weekly asymptomatic testing is now offered to all CUHNFT staff. Testing was performed (1) at temporary on-site ‘Pods’ and (2) via self-swabbing kits collected by HCWs. Individuals performed a self-swab of the oropharynx and anterior nasal cavity. Samples were subjected to RNA extraction and amplification using real-time RT-PCR, with all sample processing and analysis undertaken at the Cambridge COVID-19 Testing Centre (Lighthouse Laboratory).
Vaccination
Request a detailed protocolHCW vaccination began at CUHNFT on 8 December 2020, with appointments made by invitation only for all high-risk HCWs working on-site. This was followed by self-booked appointments for HCWs working in designated COVID-19 clinical areas, from 8 January 2021 onwards. From 18 January 2021, vaccination was offered to all HCWs, with appointments made via a booking website and latterly using the hospital’s electronic patient record system ‘MyChart’. All vials of Pfizer-BioNTech COVID-19 Vaccine (BNT162b2) were stored at −74°C, before being transferred to storage at 2–8°C. From the moment the vials were removed from the freezer, they were given a 120 hr expiration date, of which 3 hr were dedicated to thawing the vaccines. All vaccine doses were administered intramuscularly by trained vaccinators, in accordance with the manufacturer’s instructions. Vaccination was undertaken exclusively at an on-site vaccination centre, with mandatory mask-wearing and social distancing in place. HCWs remained at the on-site vaccination centre for a minimum observation period of 15 min after vaccination.
Data extraction and analysis
Request a detailed protocolSwab result, vaccination details, and serology data for HCWs were extracted directly from the hospital-laboratory interface software, Epic (Verona, WI). Data were collated using Microsoft Excel and the figure produced with GraphPad Prism (GraphPad Software, La Jolla, CA). Fisher’s exact test was used for the comparison of positive rates between groups, defined in the main text. Additionally, 95% confidence intervals were calculated using Wilson’s method.
Data availability
All data generated or analysed during this study are included in the manuscript and supporting files. Source data file has been provided for Figure 1.
References
-
Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccineNew England Journal of Medicine 383:2603–2615.https://doi.org/10.1056/NEJMoa2034577
Article and author information
Author details
Funding
Wellcome Trust (108070/Z/15/Z)
- Michael P Weekes
Wellcome Trust (207498/Z/17/Z)
- Ian G Goodfellow
Wellcome Trust (210688/Z/18/Z)
- Paul J Lehner
Medical Research Council (MR/P008801/1)
- Nicholas J Matheson
NHS Blood and Transplant (WPA15-02)
- Nicholas J Matheson
EPSRC (EP/P031447/1)
- Richard J Samworth
Medical Research Council (MC_UU_00002/10)
- Shaun Seaman
EPSRC (EP/N031938/1)
- Richard J Samworth
The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Acknowledgements
This work was funded in part by Wellcome Senior Clinical Research Fellowships (Grant numbers 108070/Z/15/Z to MPW, 207498/Z/17/Z to IGG), a Wellcome Principal Research Fellowship to PJL (210688/Z/18/Z), an MRC Clinician Scientist Fellowship (MR/P008801/1) and NHSBT workpackage (WPA15-02) to NJM, EPSRC grants to RJS (EP/P031447/1,EP/N031938/1), and an MRC grant to SS (MC_UU_00002/10). The sequencing costs were funded by the COVID-19 Genomics UK (COG-UK) Consortium, which is supported by funding from the Medical Research Council (MRC) part of UK Research and Innovation (UKRI), the National Institute of Health Research (NIHR), and Genome Research Limited, operating as the Wellcome Sanger Institute. Funding was also received from Addenbrooke’s Charitable Trust and the NIHR Cambridge Biomedical Research Centre. For the purpose of open access, the author has applied a CC BY public copyright licence to any Author Accepted Manuscript version arising from this submission. We also acknowledge contributions from all staff at CUHNFT Occupational Health and Wellbeing, the COVID-19 vaccination programme and the Cambridge COVID-19 Testing Centre.
Ethics
Human subjects: This study was conducted as a service evaluation of the CUHNFT staff testing and vaccination services (CUHNFT clinical project ID ID3682). As a study of healthcare-associated infections, this investigation is exempt from requiring ethical approval under Section 251 of the NHS Act 2006 (see also the NHS Health Research Authority algorithm, available at http://www.hra-decisiontools.org.uk/research/, which concludes that no formal ethical approval is required).
Copyright
© 2021, Jones et al.
This article is distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use and redistribution provided that the original author and source are credited.
Metrics
-
- 5,662
- views
-
- 656
- downloads
-
- 47
- citations
Views, downloads and citations are aggregated across all versions of this paper published by eLife.
Citations by DOI
-
- 47
- citations for umbrella DOI https://doi.org/10.7554/eLife.68808
Download links
Downloads (link to download the article as PDF)
Open citations (links to open the citations from this article in various online reference manager services)
Cite this article (links to download the citations from this article in formats compatible with various reference manager tools)
Further reading
-
- Epidemiology and Global Health
- Medicine
Previously, we showed that 3% (31/1032)of asymptomatic healthcare workers (HCWs) from a large teaching hospital in Cambridge, UK, tested positive for SARS-CoV-2 in April 2020. About 15% (26/169) HCWs with symptoms of coronavirus disease 2019 (COVID-19) also tested positive for SARS-CoV-2 (Rivett et al., 2020). Here, we show that the proportion of both asymptomatic and symptomatic HCWs testing positive for SARS-CoV-2 rapidly declined to near-zero between 25th April and 24th May 2020, corresponding to a decline in patient admissions with COVID-19 during the ongoing UK ‘lockdown’. These data demonstrate how infection prevention and control measures including staff testing may help prevent hospitals from becoming independent ‘hubs’ of SARS-CoV-2 transmission, and illustrate how, with appropriate precautions, organizations in other sectors may be able to resume on-site work safely.
-
- Epidemiology and Global Health
- Medicine
Significant differences exist in the availability of healthcare worker (HCW) SARS-CoV-2 testing between countries, and existing programmes focus on screening symptomatic rather than asymptomatic staff. Over a 3 week period (April 2020), 1032 asymptomatic HCWs were screened for SARS-CoV-2 in a large UK teaching hospital. Symptomatic staff and symptomatic household contacts were additionally tested. Real-time RT-PCR was used to detect viral RNA from a throat+nose self-swab. 3% of HCWs in the asymptomatic screening group tested positive for SARS-CoV-2. 17/30 (57%) were truly asymptomatic/pauci-symptomatic. 12/30 (40%) had experienced symptoms compatible with coronavirus disease 2019 (COVID-19)>7 days prior to testing, most self-isolating, returning well. Clusters of HCW infection were discovered on two independent wards. Viral genome sequencing showed that the majority of HCWs had the dominant lineage B∙1. Our data demonstrates the utility of comprehensive screening of HCWs with minimal or no symptoms. This approach will be critical for protecting patients and hospital staff.






